
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 9,395,156 – 9,395,270 |
| Length | 114 |
| Max. P | 0.835717 |

| Location | 9,395,156 – 9,395,270 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.93 |
| Mean single sequence MFE | -24.88 |
| Consensus MFE | -21.60 |
| Energy contribution | -21.52 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.835717 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9395156 114 + 27905053 UUCCUUUUUCAUUUCUCGUCCGUUACGUUUGAGUGAUUCAAACUUGACGUUUGCCUUUGUAUAAAUUU--UUUCUGCCUCAUUUGAUGCAGAGAAUGAAAAGUUUCCAU----GCACCCC ....((((((((((..((..(((((.(((((((...))))))).)))))..))...............--..(((((.((....)).))))))))))))))).......----....... ( -25.40) >DroVir_CAF1 76266 116 + 1 UUCCGUUUUCAUUUCUCGUCCGUUACGUUUGAGUGAUUCAAACUUGACGGUUGCCUUUGUAUACAUUUUUUUUCCGCCUCAUUUGAUGCAGAGAAUGAAAAGUUUUCAU----GCACCGC .....((((((((.(((..((((((.(((((((...))))))).)))))).........................((.((....)).)).)))))))))))........----....... ( -25.70) >DroPse_CAF1 27196 114 + 1 UUCCUUUUUCAUUUCUCGUCCGUUACGUUUGAGUGAUUCAAACUUGACGUUUGCCUUUGUAUAAAUUU--UUUCUGCCUCAUUUGGUGCAGAGAAUGAAAAGUUUCCAC----ACACCGC ....((((((((((..((..(((((.(((((((...))))))).)))))..))...............--..(((((.((....)).))))))))))))))).......----....... ( -24.50) >DroGri_CAF1 77037 115 + 1 UUCCGUUUUCAUUUCUCGUCCGUUACGUUUGAGUGAUUCAAACUUGACGGUUGCCUUUGUAUACAUUUU-UUUCCGCCUCAUUUGAUGCAGAGAAUGAAAAGUUUUCUU----GCAUCGC ....((...((.....((.((((((.(((((((...))))))).)))))).))....))...)).....-..............((((((((((((.....)))))).)----))))).. ( -27.30) >DroWil_CAF1 21959 118 + 1 UUCCGUUUUCAUUUCUCGUCCGUUACGUUUGAGUGAUUCAAACUUGACGUUUGCCUUUGUAUAAAUUU--UUUCUGCCUCAUUUGAUGCAGAGAAUGAAAAGUUUCCCUAGCUGCACCUU .....(((((((((..((..(((((.(((((((...))))))).)))))..))...............--..(((((.((....)).))))))))))))))................... ( -25.10) >DroMoj_CAF1 85485 115 + 1 UUCCGCUUUCAUUUCUCGUCCGUUACGUUUGAGUGAUUCAAACUUGACGGUUGCCUUUGUAUACAUUUU-UUUCCGGCUCAUUUGAUCCACAGAAUGAAAAGUUUUCAU----GCACCGC ....((((((((((..((.((((((.(((((((...))))))).)))))).))................-.....((.((....)).))...)))))))).((......----))...)) ( -21.30) >consensus UUCCGUUUUCAUUUCUCGUCCGUUACGUUUGAGUGAUUCAAACUUGACGGUUGCCUUUGUAUAAAUUU__UUUCCGCCUCAUUUGAUGCAGAGAAUGAAAAGUUUCCAU____GCACCGC .....((((((((.(((...(((((.(((((((...))))))).)))))..........................((.((....)).)).)))))))))))................... (-21.60 = -21.52 + -0.08)



| Location | 9,395,156 – 9,395,270 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.93 |
| Mean single sequence MFE | -23.79 |
| Consensus MFE | -21.26 |
| Energy contribution | -20.93 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.723271 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9395156 114 - 27905053 GGGGUGC----AUGGAAACUUUUCAUUCUCUGCAUCAAAUGAGGCAGAAA--AAAUUUAUACAAAGGCAAACGUCAAGUUUGAAUCACUCAAACGUAACGGACGAGAAAUGAAAAAGGAA .......----..(....)((((((((((((((.((....)).)))))..--...................((((..((((((.....))))))......)))).).))))))))..... ( -23.80) >DroVir_CAF1 76266 116 - 1 GCGGUGC----AUGAAAACUUUUCAUUCUCUGCAUCAAAUGAGGCGGAAAAAAAAUGUAUACAAAGGCAACCGUCAAGUUUGAAUCACUCAAACGUAACGGACGAGAAAUGAAAACGGAA .((((((----(((((.....)))....(((((.((....)).)))))......))))))..........((((...((((((.....))))))...))))..............))... ( -23.90) >DroPse_CAF1 27196 114 - 1 GCGGUGU----GUGGAAACUUUUCAUUCUCUGCACCAAAUGAGGCAGAAA--AAAUUUAUACAAAGGCAAACGUCAAGUUUGAAUCACUCAAACGUAACGGACGAGAAAUGAAAAAGGAA ((..(((----(((....).........(((((..(....)..)))))..--......)))))...))...((((..((((((.....))))))......))))................ ( -24.20) >DroGri_CAF1 77037 115 - 1 GCGAUGC----AAGAAAACUUUUCAUUCUCUGCAUCAAAUGAGGCGGAAA-AAAAUGUAUACAAAGGCAACCGUCAAGUUUGAAUCACUCAAACGUAACGGACGAGAAAUGAAAACGGAA .......----........((((((((((((((.((....)).)))....-...................((((...((((((.....))))))...))))..))).))))))))..... ( -23.30) >DroWil_CAF1 21959 118 - 1 AAGGUGCAGCUAGGGAAACUUUUCAUUCUCUGCAUCAAAUGAGGCAGAAA--AAAUUUAUACAAAGGCAAACGUCAAGUUUGAAUCACUCAAACGUAACGGACGAGAAAUGAAAACGGAA ....(((.(..(((....)))..)....(((((.((....)).)))))..--..............)))..((((..((((((.....))))))......))))................ ( -24.20) >DroMoj_CAF1 85485 115 - 1 GCGGUGC----AUGAAAACUUUUCAUUCUGUGGAUCAAAUGAGCCGGAAA-AAAAUGUAUACAAAGGCAACCGUCAAGUUUGAAUCACUCAAACGUAACGGACGAGAAAUGAAAGCGGAA (((((..----......)))(((((((((((...........(((.....-..............)))..((((...((((((.....))))))...)))))).)).))))))))).... ( -23.31) >consensus GCGGUGC____AUGAAAACUUUUCAUUCUCUGCAUCAAAUGAGGCAGAAA__AAAUGUAUACAAAGGCAAACGUCAAGUUUGAAUCACUCAAACGUAACGGACGAGAAAUGAAAACGGAA ...................((((((((((((((.((....)).))))).......................((((..((((((.....))))))......)))).).))))))))..... (-21.26 = -20.93 + -0.33)

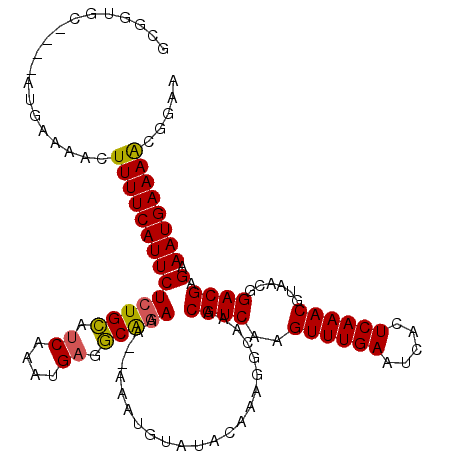

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:02:09 2006