
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 9,379,226 – 9,379,459 |
| Length | 233 |
| Max. P | 0.995707 |

| Location | 9,379,226 – 9,379,324 |
|---|---|
| Length | 98 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 78.04 |
| Mean single sequence MFE | -20.89 |
| Consensus MFE | -15.16 |
| Energy contribution | -15.49 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.41 |
| Structure conservation index | 0.73 |
| SVM decision value | 2.45 |
| SVM RNA-class probability | 0.994097 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9379226 98 + 27905053 AGCAUU-----UCUUUGAUUAAAUGAAUUUGAAUUAAAUGAAUAUACUACUUUUUUUCACCGUGUAGCUACCAGCAGCAGCUCAAAGCUCACGGCACACAGCA .(((((-----(.(((((((.((.....)).))))))).))))................(((((.((((...(((....)))...)))))))))......)). ( -20.60) >DroEre_CAF1 9482 88 + 1 AGCAUU--------AUGAAUAAAUGAAUCCAC----AAUGA--AUACCAC-AUUUUUCACCGUGUAGCUACCAGCAGCAGCUCAAAGCUCACGGCACACAGCA .((...--------.((((.(((((.......----.....--......)-))))))))(((((.((((...(((....)))...)))))))))......)). ( -19.47) >DroYak_CAF1 9090 92 + 1 AGCAUUCGUGUGA-AUAAAUAAAUGUAC---C----AUUUU--AUAGCAC-UUUUUUCACCGUGUAGCUACCAGCAGCAGCUCAAAGCUCACGGCACACAGCA .......(((((.-....(((((.....---.----..)))--)).....-........(((((.((((...(((....)))...)))))))))))))).... ( -22.60) >consensus AGCAUU________AUGAAUAAAUGAAU___C____AAUGA__AUACCAC_UUUUUUCACCGUGUAGCUACCAGCAGCAGCUCAAAGCUCACGGCACACAGCA .((....................((((............................))))(((((.((((...(((....)))...)))))))))......)). (-15.16 = -15.49 + 0.33)



| Location | 9,379,226 – 9,379,324 |
|---|---|
| Length | 98 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 78.04 |
| Mean single sequence MFE | -27.70 |
| Consensus MFE | -19.70 |
| Energy contribution | -19.70 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.51 |
| Structure conservation index | 0.71 |
| SVM decision value | 2.61 |
| SVM RNA-class probability | 0.995707 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9379226 98 - 27905053 UGCUGUGUGCCGUGAGCUUUGAGCUGCUGCUGGUAGCUACACGGUGAAAAAAAGUAGUAUAUUCAUUUAAUUCAAAUUCAUUUAAUCAAAGA-----AAUGCU ((((...(((((((((((.(.(((....))).).)))).)))))))......))))((((.(((.(((.(((.(((....)))))).)))))-----))))). ( -22.60) >DroEre_CAF1 9482 88 - 1 UGCUGUGUGCCGUGAGCUUUGAGCUGCUGCUGGUAGCUACACGGUGAAAAAU-GUGGUAU--UCAUU----GUGGAUUCAUUUAUUCAU--------AAUGCU (((..(((((((((((((.(.(((....))).).)))).)))))).....))-)..))).--.((((----((((((......))))))--------)))).. ( -30.50) >DroYak_CAF1 9090 92 - 1 UGCUGUGUGCCGUGAGCUUUGAGCUGCUGCUGGUAGCUACACGGUGAAAAAA-GUGCUAU--AAAAU----G---GUACAUUUAUUUAU-UCACACGAAUGCU .(((((((((((((((((.(.(((....))).).)))).)))))........-(((((((--...))----)---))))..........-.))))))...)). ( -30.00) >consensus UGCUGUGUGCCGUGAGCUUUGAGCUGCUGCUGGUAGCUACACGGUGAAAAAA_GUGGUAU__UCAUU____G___AUUCAUUUAUUCAU________AAUGCU ........((((((((((.(.(((....))).).)))).)))))).......................................................... (-19.70 = -19.70 + -0.00)



| Location | 9,379,324 – 9,379,422 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.04 |
| Mean single sequence MFE | -29.28 |
| Consensus MFE | -19.75 |
| Energy contribution | -19.95 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.626845 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9379324 98 + 27905053 --------------------CGCCACUAAUUGGACCCAGUGGUUCGCCAUCAU--CAUCGUGAUCGCAUUUAAAUGUGCUGAUCAAUUUACUUAAAUGUUAAUGGCAGCCCAGGGCGUGU --------------------((((.((.((((....))))((((.(((((...--(((..(((((((((......)))).)))))..........)))...))))))))).))))))... ( -28.60) >DroVir_CAF1 64384 91 + 1 --------------------UGCCACUAAUUGGACCCAGCGGUUCGCCA--------UCGUGAUCGCAUUUAAAUGUGCUGAUCAAUUUGGUUAAAUGUUAAUAGC-GCCCUGGGCGAGC --------------------..(((.....))).(((((.(((..((((--------...(((((((((......)))).)))))...)))).....((.....))-))))))))..... ( -27.90) >DroGri_CAF1 63716 91 + 1 --------------------UGCCACUAAUUGGACCCAGUGGUUCACCA--------UCGUGAUCGCAUUUAAAUGUGCUGAUCAAUUUGGUUAAAUGUUAAUAGC-GCCCUGGGCGAGG --------------------..(((.....))).(((((.(((..((((--------...(((((((((......)))).)))))...)))).....((.....))-))))))))..... ( -27.60) >DroEre_CAF1 9570 98 + 1 --------------------CGCCACUAAUUGGGCCCAGUGGUUCGCCAUCAU--CAUCGUGAUCGCAUUUAAAUGUGCUGAUCAAUUUACUUAAAUGUUAAUGGCAGCCCAGGGCGUGU --------------------..(((.....)))((((.(.((((.(((((...--(((..(((((((((......)))).)))))..........)))...)))))))))).)))).... ( -30.30) >DroWil_CAF1 7888 98 + 1 --------------------CGCCACUAAUUGGACCCAGCGGCUUACCAUCAUCUCAACGUGAUCGCAUUUAAAUGUGCUGAUCAAUUUACUUAAAUGUUAAUGAU-GCCCUGGGCGAG- --------------------..(((.....))).(((((.(((.....(((((...(((.(((((((((......)))).)))))((((....))))))).)))))-))))))))....- ( -25.20) >DroMoj_CAF1 71846 111 + 1 GGCGGAGCUGCCGUUGCUUUGGCCACUAAUUGGACCCAGCGGUUGGGCA--------UCGUGAUCGCAUUUAAAUGUGCUGAUCAAUUUGGUUAAAUGUUAAUAGC-GCCCUGGGCGAGC .((...((((((((((....(((((.....))).))))))))).((((.--------...(((((((((......)))).))))).....((((........))))-))))..)))..)) ( -36.10) >consensus ____________________CGCCACUAAUUGGACCCAGCGGUUCGCCA________UCGUGAUCGCAUUUAAAUGUGCUGAUCAAUUUACUUAAAUGUUAAUAGC_GCCCUGGGCGAGC ......................(((.....))).(((((.(((.................(((((((((......)))).)))))((((....))))..........))))))))..... (-19.75 = -19.95 + 0.20)


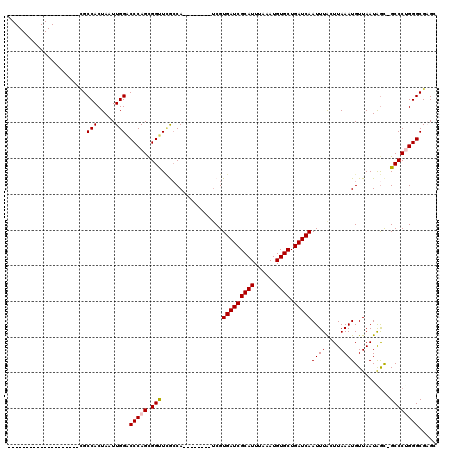
| Location | 9,379,324 – 9,379,422 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.04 |
| Mean single sequence MFE | -27.98 |
| Consensus MFE | -19.64 |
| Energy contribution | -19.28 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.762444 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9379324 98 - 27905053 ACACGCCCUGGGCUGCCAUUAACAUUUAAGUAAAUUGAUCAGCACAUUUAAAUGCGAUCACGAUG--AUGAUGGCGAACCACUGGGUCCAAUUAGUGGCG-------------------- ...((((..((..((((((((.((((.........(((((.(((........)))))))).))))--.))))))))..))(((((......)))))))))-------------------- ( -29.30) >DroVir_CAF1 64384 91 - 1 GCUCGCCCAGGGC-GCUAUUAACAUUUAACCAAAUUGAUCAGCACAUUUAAAUGCGAUCACGA--------UGGCGAACCGCUGGGUCCAAUUAGUGGCA-------------------- ((((.....))))-((((((((.......(((...(((((.(((........))))))))...--------))).(.(((....))).)..)))))))).-------------------- ( -25.60) >DroGri_CAF1 63716 91 - 1 CCUCGCCCAGGGC-GCUAUUAACAUUUAACCAAAUUGAUCAGCACAUUUAAAUGCGAUCACGA--------UGGUGAACCACUGGGUCCAAUUAGUGGCA-------------------- ....(((...)))-((((((((......(((....(((((.(((........))))))))((.--------(((....))).)))))....)))))))).-------------------- ( -24.90) >DroEre_CAF1 9570 98 - 1 ACACGCCCUGGGCUGCCAUUAACAUUUAAGUAAAUUGAUCAGCACAUUUAAAUGCGAUCACGAUG--AUGAUGGCGAACCACUGGGCCCAAUUAGUGGCG-------------------- ....(((((((..((((((((.((((.........(((((.(((........)))))))).))))--.))))))))..)))..))))(((.....)))..-------------------- ( -30.70) >DroWil_CAF1 7888 98 - 1 -CUCGCCCAGGGC-AUCAUUAACAUUUAAGUAAAUUGAUCAGCACAUUUAAAUGCGAUCACGUUGAGAUGAUGGUAAGCCGCUGGGUCCAAUUAGUGGCG-------------------- -.....(((...(-(((.(((((((((....))))(((((.(((........)))))))).))))))))).)))...((((((((......)))))))).-------------------- ( -27.90) >DroMoj_CAF1 71846 111 - 1 GCUCGCCCAGGGC-GCUAUUAACAUUUAACCAAAUUGAUCAGCACAUUUAAAUGCGAUCACGA--------UGCCCAACCGCUGGGUCCAAUUAGUGGCCAAAGCAACGGCAGCUCCGCC (((.(((..((((-(........((((....))))(((((.(((........))))))))...--------)))))....(((.((.(((.....)))))..)))...))))))...... ( -29.50) >consensus ACUCGCCCAGGGC_GCCAUUAACAUUUAACCAAAUUGAUCAGCACAUUUAAAUGCGAUCACGA________UGGCGAACCACUGGGUCCAAUUAGUGGCG____________________ ....((((((((..((((.....((((....))))(((((.(((........))))))))...........))))...)).))))))(((.....)))...................... (-19.64 = -19.28 + -0.36)


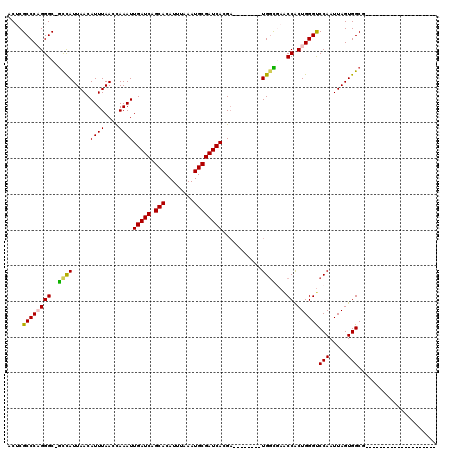
| Location | 9,379,344 – 9,379,459 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 94.45 |
| Mean single sequence MFE | -23.89 |
| Consensus MFE | -21.33 |
| Energy contribution | -21.09 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.868733 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9379344 115 - 27905053 -CCCCCAGACC--GCCCGCCCCACUAACCCAUACCCUUUGACACGCCCUGGGCUGCCAUUAACAUUUAAGUAAAUUGAUCAGCACAUUUAAAUGCGAUCACGAUGAUGAUGGCGAACC -..(((((..(--(.............................))..))))).((((((((.((((.........(((((.(((........)))))))).)))).)))))))).... ( -23.55) >DroSec_CAF1 8913 112 - 1 CCCCCCA------GUCCGCUCCACUAACCCAUACCCUUUGACACGCCCUGGGCUGCCAUUAACAUUUAAGUAAAUUGAUCAGCACAUUUAAAUGCGAUCACGAUGAUGAUGGCGAACC ...((((------(..((.........................))..))))).((((((((.((((.........(((((.(((........)))))))).)))).)))))))).... ( -24.61) >DroSim_CAF1 8952 112 - 1 CCCCCCA------GUCCGCUCCACUAACCCAUACCCUUUGACACGCCCUGGGCUGCCAUUAACAUUUAAGUAAAUUGAUCAGCACAUUUAAAUGCGAUCACGAUGAUGAUGGCGAACC ...((((------(..((.........................))..))))).((((((((.((((.........(((((.(((........)))))))).)))).)))))))).... ( -24.61) >DroEre_CAF1 9590 111 - 1 -CCCCCA------ACCAGCCCCACUAACCCAUACCCUUUGACACGCCCUGGGCUGCCAUUAACAUUUAAGUAAAUUGAUCAGCACAUUUAAAUGCGAUCACGAUGAUGAUGGCGAACC -......------...(((((........((.......)).........)))))(((((((.((((.........(((((.(((........)))))))).)))).)))))))..... ( -23.33) >DroYak_CAF1 9202 117 - 1 -CCCCCAGCCUGAACCAGCCCCACUAACCCAUACCCUUUGACACGCCCUGGGCUGCCAUUAACAUUUAAGUAAAUUGAUCAGCACAUUUAAAUGCGAUCACGAUGAUGAUGGCGAACC -...............(((((........((.......)).........)))))(((((((.((((.........(((((.(((........)))))))).)))).)))))))..... ( -23.33) >consensus _CCCCCA______GCCCGCCCCACUAACCCAUACCCUUUGACACGCCCUGGGCUGCCAUUAACAUUUAAGUAAAUUGAUCAGCACAUUUAAAUGCGAUCACGAUGAUGAUGGCGAACC .................((((........((.......)).........))))((((((((.((((.........(((((.(((........)))))))).)))).)))))))).... (-21.33 = -21.09 + -0.24)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:02:05 2006