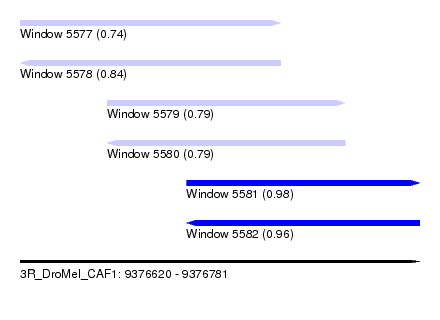

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 9,376,620 – 9,376,781 |
| Length | 161 |
| Max. P | 0.979003 |

| Location | 9,376,620 – 9,376,725 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 80.68 |
| Mean single sequence MFE | -24.77 |
| Consensus MFE | -14.18 |
| Energy contribution | -14.15 |
| Covariance contribution | -0.03 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.742391 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
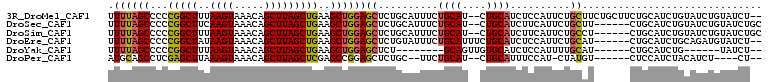
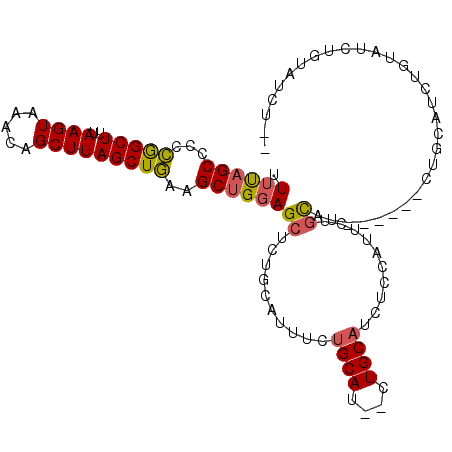
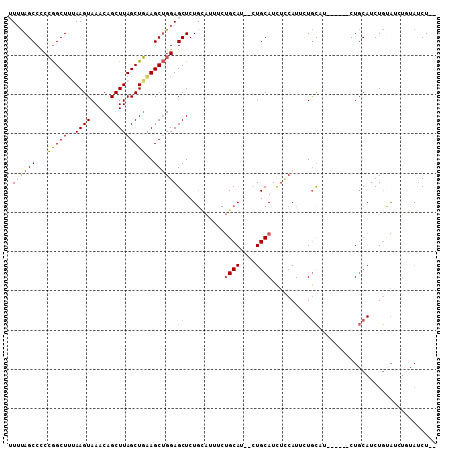
>3R_DroMel_CAF1 9376620 105 + 27905053 UUUUAGCCCCCGGCUUUAAGUAAACAGCUUAGCUGAAGCUGGAGCUCUGCAUUUCUGCAU--CUGCAUCUCCAUUCUGCUUCUGCUUCUGCAUCUGUAUCUGUAUCU-- ((..((((...))))..))....((((....((.(((((.(((((..........(((..--..)))..........))))).))))).))..))))..........-- ( -25.25) >DroSec_CAF1 6244 101 + 1 UUUUAGCCCCCGGCUUCAAGUAAACAGCUUAGCUGAAGCUGGAGCUCUGCAUUUCUGCAU--CUGCAUCUUCAUUCUGCUU------CUGCAUCUGUAUCUGUAUCUGC ....(((..(((((((((..(((.....)))..))))))))).)))..........(((.--.((((....((...(((..------..)))..))....))))..))) ( -23.40) >DroSim_CAF1 6305 101 + 1 UUUUAGCCCCCGGCUUUAAGUAAACAGCUUAGCUGAAGCUGGAGCUCUGCAUUUCUGCAU--CUGCAUCUUCAUUCUGCCU------CUGCAUCUGUAUCUGUAUCUGC ....(((..(((((((((..(((.....)))..))))))))).))).((((....((((.--.((((..............------.))))..))))..))))..... ( -20.06) >DroEre_CAF1 7009 101 + 1 UUUUAGCCCCCGGCUAUAAGUAAACAGCUUAGCUGAAGCUGGAGCUCUGUAUUUCUGCAUUUCUGCAUCUCCAUUCUGCAU------CUGCAUCUGCAGAUGUAUCU-- ...(((((...))))).(((((...((((((((....))).)))))...))))).((((....)))).........(((((------((((....)))))))))...-- ( -29.70) >DroYak_CAF1 6495 87 + 1 UUUUAGCCCCCGGCUUUAAGUAAACAGCUUAGCUGAAGCUGGAGCUCU--------GCAGUUGUGCAUCUCCAUUUUGCAU------CUGCAUCUG------UAUCU-- ....(((..(((((((((..(((.....)))..))))))))).))).(--------((((..(((((.........)))))------)))))....------.....-- ( -24.60) >DroPer_CAF1 10889 92 + 1 AUGCAGCCUCGAGCUUAAAGUAAACAGCUUAGCUCGAGCCGGAGCUCUGC--UUCUGCAU--CUGCAUUUCCAU-CUAUGU------CUCCAUCUACAUCU----CU-- ((((((..(((((((..((((.....)))))))))))((.(((((...))--))).))..--))))))......-......------..............----..-- ( -25.60) >consensus UUUUAGCCCCCGGCUUUAAGUAAACAGCUUAGCUGAAGCUGGAGCUCUGCAUUUCUGCAU__CUGCAUCUCCAUUCUGCAU______CUGCAUCUGUAUCUGUAUCU__ .((((((...(((((..((((.....)))))))))..))))))((..........((((....))))..........)).............................. (-14.18 = -14.15 + -0.03)
| Location | 9,376,620 – 9,376,725 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 80.68 |
| Mean single sequence MFE | -27.72 |
| Consensus MFE | -17.18 |
| Energy contribution | -17.27 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.837526 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
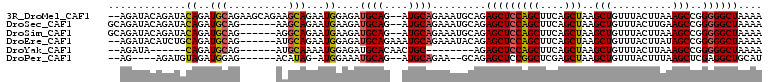
>3R_DroMel_CAF1 9376620 105 - 27905053 --AGAUACAGAUACAGAUGCAGAAGCAGAAGCAGAAUGGAGAUGCAG--AUGCAGAAAUGCAGAGCUCCAGCUUCAGCUAAGCUGUUUACUUAAAGCCGGGGGCUAAAA --................((((.(((.(((((....(((((......--.((((....))))...)))))))))).)))...))))........((((...)))).... ( -28.90) >DroSec_CAF1 6244 101 - 1 GCAGAUACAGAUACAGAUGCAG------AAGCAGAAUGAAGAUGCAG--AUGCAGAAAUGCAGAGCUCCAGCUUCAGCUAAGCUGUUUACUUGAAGCCGGGGGCUAAAA (((..(.((........(((..------..)))...)).)..)))..--.((((....)))).((((((.(((((((.((((...)))).)))))))..)))))).... ( -28.50) >DroSim_CAF1 6305 101 - 1 GCAGAUACAGAUACAGAUGCAG------AGGCAGAAUGAAGAUGCAG--AUGCAGAAAUGCAGAGCUCCAGCUUCAGCUAAGCUGUUUACUUAAAGCCGGGGGCUAAAA (((..(.((........(((..------..)))...)).)..)))..--.((((....)))).((((((.((((((((...))))........))))..)))))).... ( -25.10) >DroEre_CAF1 7009 101 - 1 --AGAUACAUCUGCAGAUGCAG------AUGCAGAAUGGAGAUGCAGAAAUGCAGAAAUACAGAGCUCCAGCUUCAGCUAAGCUGUUUACUUAUAGCCGGGGGCUAAAA --.....(((((((....))))------))).....((....((((....))))......)).(((((((((....)))..(((((......)))))..)))))).... ( -32.60) >DroYak_CAF1 6495 87 - 1 --AGAUA------CAGAUGCAG------AUGCAAAAUGGAGAUGCACAACUGC--------AGAGCUCCAGCUUCAGCUAAGCUGUUUACUUAAAGCCGGGGGCUAAAA --.....------....(((((------.((((.........))))...))))--------).((((((.((((((((...))))........))))..)))))).... ( -22.90) >DroPer_CAF1 10889 92 - 1 --AG----AGAUGUAGAUGGAG------ACAUAG-AUGGAAAUGCAG--AUGCAGAA--GCAGAGCUCCGGCUCGAGCUAAGCUGUUUACUUUAAGCUCGAGGCUGCAU --..----..(((((((((...------.)))..-.((((...((..--.(((....--)))..)))))).(((((((((((.......)))..)))))))).)))))) ( -28.30) >consensus __AGAUACAGAUACAGAUGCAG______AAGCAGAAUGGAGAUGCAG__AUGCAGAAAUGCAGAGCUCCAGCUUCAGCUAAGCUGUUUACUUAAAGCCGGGGGCUAAAA .............((..(((..........)))...))....((((....)))).........(((((((((....)))..(((..........)))..)))))).... (-17.18 = -17.27 + 0.08)
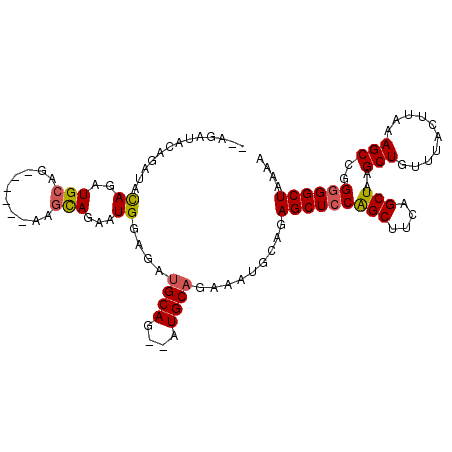
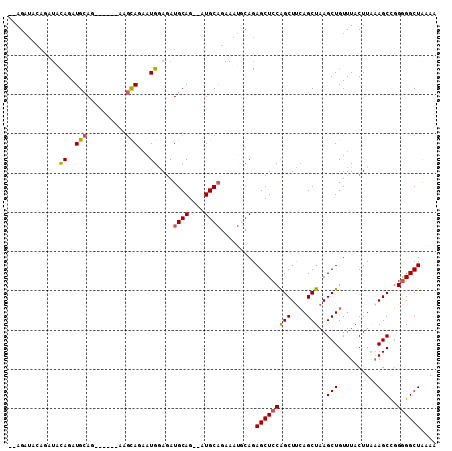
| Location | 9,376,655 – 9,376,751 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 89.72 |
| Mean single sequence MFE | -21.46 |
| Consensus MFE | -16.09 |
| Energy contribution | -16.40 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.786401 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9376655 96 + 27905053 AAGCUGGAGCUCUGCAUUUCUGCAU--CUGCAUCUCCAUUCUGCUUCUGCUUCUGCAUCUGUAUCUGUAUCU--GCAAAUGCACAUGUACGCUUGUAUUU ((((.(((((..((((....)))).--..(((.........)))....)))))(((((.(((((.(((....--))).))))).))))).))))...... ( -22.30) >DroSec_CAF1 6279 92 + 1 AAGCUGGAGCUCUGCAUUUCUGCAU--CUGCAUCUUCAUUCUGCUU------CUGCAUCUGUAUCUGUAUCUGCGCAAAUGCACAUGUACGCUUGUAUUU ((((.(((((..........(((..--..)))..........))))------)(((((.(((((.(((......))).))))).))))).))))...... ( -21.05) >DroSim_CAF1 6340 92 + 1 AAGCUGGAGCUCUGCAUUUCUGCAU--CUGCAUCUUCAUUCUGCCU------CUGCAUCUGUAUCUGUAUCUGCGCAAAUGCACAUGUACGCUUGUAUUU ((((.(..((..(((((((.((((.--.((((....((...(((..------..)))..))....))))..)))).)))))))...)).)))))...... ( -19.50) >DroEre_CAF1 7044 92 + 1 AAGCUGGAGCUCUGUAUUUCUGCAUUUCUGCAUCUCCAUUCUGCAU------CUGCAUCUGCAGAUGUAUCU--GCAAAUACACAUGUACACUUGUAUUU ..(((((((...........((((....)))).)))))...(((((------((((....)))))))))...--))(((((((..........))))))) ( -23.00) >consensus AAGCUGGAGCUCUGCAUUUCUGCAU__CUGCAUCUCCAUUCUGCUU______CUGCAUCUGUAUCUGUAUCU__GCAAAUGCACAUGUACGCUUGUAUUU (((((((((...((((............)))).)))))...............(((((.(((((.(((......))).))))).))))).))))...... (-16.09 = -16.40 + 0.31)

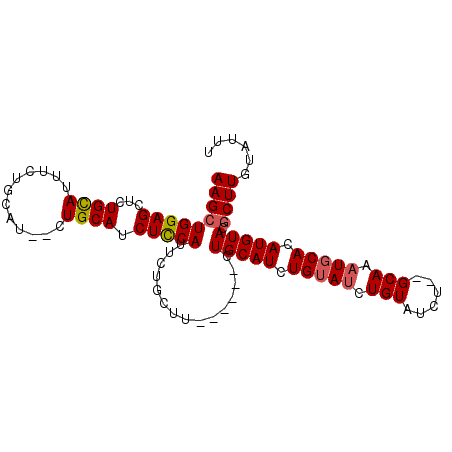
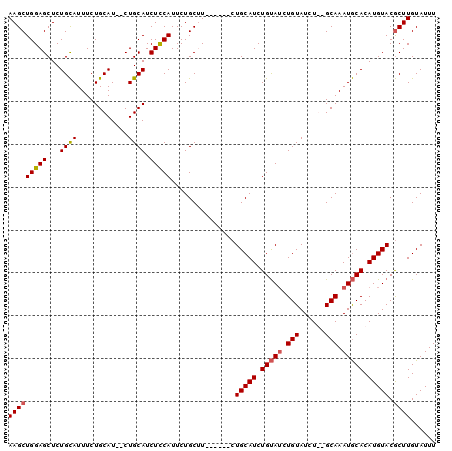
| Location | 9,376,655 – 9,376,751 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 89.72 |
| Mean single sequence MFE | -26.85 |
| Consensus MFE | -18.33 |
| Energy contribution | -17.77 |
| Covariance contribution | -0.56 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.21 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.794563 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9376655 96 - 27905053 AAAUACAAGCGUACAUGUGCAUUUGC--AGAUACAGAUACAGAUGCAGAAGCAGAAGCAGAAUGGAGAUGCAG--AUGCAGAAAUGCAGAGCUCCAGCUU ......((((.....(.(((.(((((--(..(........)..)))))).))).).......(((((......--.((((....))))...))))))))) ( -25.40) >DroSec_CAF1 6279 92 - 1 AAAUACAAGCGUACAUGUGCAUUUGCGCAGAUACAGAUACAGAUGCAG------AAGCAGAAUGAAGAUGCAG--AUGCAGAAAUGCAGAGCUCCAGCUU ......(((((((..(((((....)))))..))).....((..(((..------..)))...))..((.((..--.((((....))))..))))..)))) ( -24.80) >DroSim_CAF1 6340 92 - 1 AAAUACAAGCGUACAUGUGCAUUUGCGCAGAUACAGAUACAGAUGCAG------AGGCAGAAUGAAGAUGCAG--AUGCAGAAAUGCAGAGCUCCAGCUU ......(((((((..(((((....)))))..))).....((..(((..------..)))...))..((.((..--.((((....))))..))))..)))) ( -24.70) >DroEre_CAF1 7044 92 - 1 AAAUACAAGUGUACAUGUGUAUUUGC--AGAUACAUCUGCAGAUGCAG------AUGCAGAAUGGAGAUGCAGAAAUGCAGAAAUACAGAGCUCCAGCUU ......((((...(((.(((((((((--(((....)))))))))))).------))).....(((((.((((....))))...........))))))))) ( -32.50) >consensus AAAUACAAGCGUACAUGUGCAUUUGC__AGAUACAGAUACAGAUGCAG______AAGCAGAAUGAAGAUGCAG__AUGCAGAAAUGCAGAGCUCCAGCUU ........((((.....((((((((..............)))))))).....................((((....))))...)))).((((....)))) (-18.33 = -17.77 + -0.56)

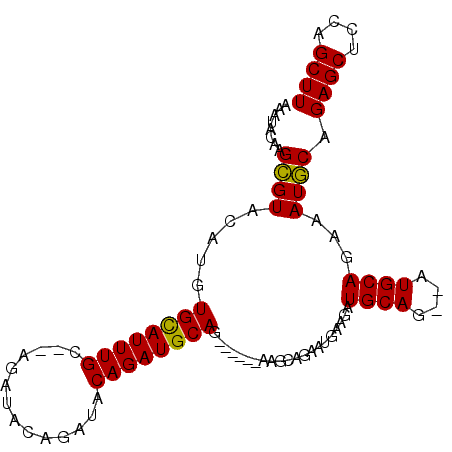
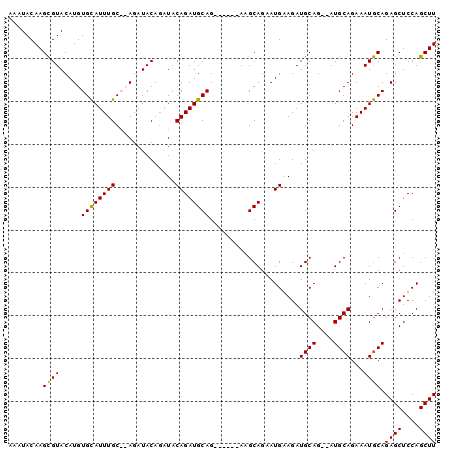
| Location | 9,376,687 – 9,376,781 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 83.26 |
| Mean single sequence MFE | -29.14 |
| Consensus MFE | -15.99 |
| Energy contribution | -16.67 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.51 |
| Structure conservation index | 0.55 |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.979003 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9376687 94 + 27905053 UCCAUUCUGCUUCUGCUUCUGCAUCUGUAUCUGUAUCU--GCAAAU--------------GCACAUGUACGCUUGUAUUUGUAUCUGAAGCCGAAGCAGACAAGAAUGGC .(((((((...((((((((.((.((.((((.(((....--))).))--------------))(((.((((....)))).)))....)).)).))))))))..))))))). ( -33.70) >DroSec_CAF1 6311 90 + 1 UUCAUUCUGCUU------CUGCAUCUGUAUCUGUAUCUGCGCAAAU--------------GCACAUGUACGCUUGUAUUUGUAUCUGAAGCCGAAGCAGACAAGAAUGGC (((..(((((((------(.((.((.((((..((((..(((....(--------------((....))))))..))))..))))..)).)).))))))))...))).... ( -25.10) >DroSim_CAF1 6372 90 + 1 UUCAUUCUGCCU------CUGCAUCUGUAUCUGUAUCUGCGCAAAU--------------GCACAUGUACGCUUGUAUUUGUAUCUGAAGCCGAAGCAGACAAGAAUGGC .(((((((...(------((((.((.((.((.((....))((((((--------------(((...(....).)))))))))....)).)).)).)))))..))))))). ( -25.00) >DroEre_CAF1 7078 88 + 1 UCCAUUCUGCAU------CUGCAUCUGCAGAUGUAUCU--GCAAAU--------------ACACAUGUACACUUGUAUUUGUAUCUGAAGCCGAAGCAGACAAGAAUGGC .(((((((((((------((((....)))))))).(((--((....--------------..(((.((((....)))).))).((.......)).)))))..))))))). ( -30.90) >DroYak_CAF1 6556 96 + 1 UCCAUUUUGCAU------CUGCAUCUG------UAUCU--GCAAAUGCACACGCACAUGUACACAUGCACAGUUGUAUUUGUAUCUGAAGCCGAAGCAGACAAGAAUGGC .(((((((...(------((((.((.(------(.(((--(((((((((.((...((((....))))....))))))))))))...)).)).)).)))))..))))))). ( -31.00) >consensus UCCAUUCUGCAU______CUGCAUCUGUAUCUGUAUCU__GCAAAU______________GCACAUGUACGCUUGUAUUUGUAUCUGAAGCCGAAGCAGACAAGAAUGGC .(((((((((..........)).(((((.((.((.((...((((((................(((........)))))))))....)).)).)).)))))..))))))). (-15.99 = -16.67 + 0.68)

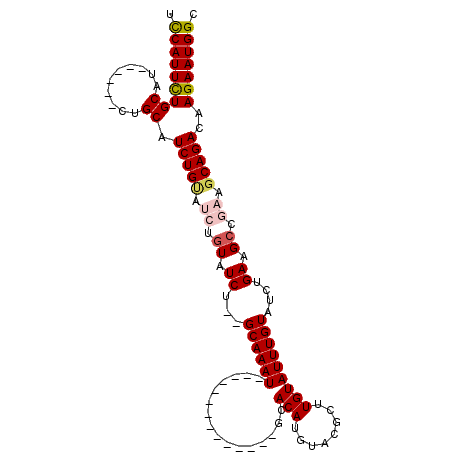
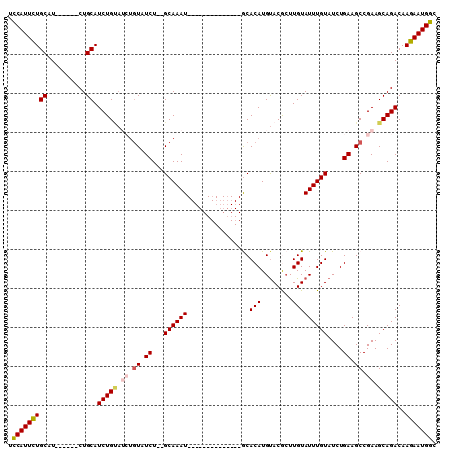
| Location | 9,376,687 – 9,376,781 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 83.26 |
| Mean single sequence MFE | -30.78 |
| Consensus MFE | -14.88 |
| Energy contribution | -15.32 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.55 |
| Structure conservation index | 0.48 |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.961130 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9376687 94 - 27905053 GCCAUUCUUGUCUGCUUCGGCUUCAGAUACAAAUACAAGCGUACAUGUGC--------------AUUUGC--AGAUACAGAUACAGAUGCAGAAGCAGAAGCAGAAUGGA .(((((((..((((((((.((((...((....))..)))).......(((--------------(((((.--...........))))))))))))))))...))))))). ( -32.40) >DroSec_CAF1 6311 90 - 1 GCCAUUCUUGUCUGCUUCGGCUUCAGAUACAAAUACAAGCGUACAUGUGC--------------AUUUGCGCAGAUACAGAUACAGAUGCAG------AAGCAGAAUGAA ..((((((..(((((.((.((((...((....))..))))(((..(((((--------------....)))))..))).......)).))))------)...)))))).. ( -27.30) >DroSim_CAF1 6372 90 - 1 GCCAUUCUUGUCUGCUUCGGCUUCAGAUACAAAUACAAGCGUACAUGUGC--------------AUUUGCGCAGAUACAGAUACAGAUGCAG------AGGCAGAAUGAA ..(((((((.(((((.((.((((...((....))..))))(((..(((((--------------....)))))..))).......)).))))------).).)))))).. ( -27.80) >DroEre_CAF1 7078 88 - 1 GCCAUUCUUGUCUGCUUCGGCUUCAGAUACAAAUACAAGUGUACAUGUGU--------------AUUUGC--AGAUACAUCUGCAGAUGCAG------AUGCAGAAUGGA .((((((((((((((....))...)))))..............(((.(((--------------((((((--(((....)))))))))))).------))).))))))). ( -34.40) >DroYak_CAF1 6556 96 - 1 GCCAUUCUUGUCUGCUUCGGCUUCAGAUACAAAUACAACUGUGCAUGUGUACAUGUGCGUGUGCAUUUGC--AGAUA------CAGAUGCAG------AUGCAAAAUGGA .(((((..(((((((....))...)))))...........(..((((....))))..)...(((((((((--(....------....)))))------))))).))))). ( -32.00) >consensus GCCAUUCUUGUCUGCUUCGGCUUCAGAUACAAAUACAAGCGUACAUGUGC______________AUUUGC__AGAUACAGAUACAGAUGCAG______AAGCAGAAUGGA .((((((.(((((((....))......((((..(((....)))..)))).......................)))))..........(((..........))))))))). (-14.88 = -15.32 + 0.44)

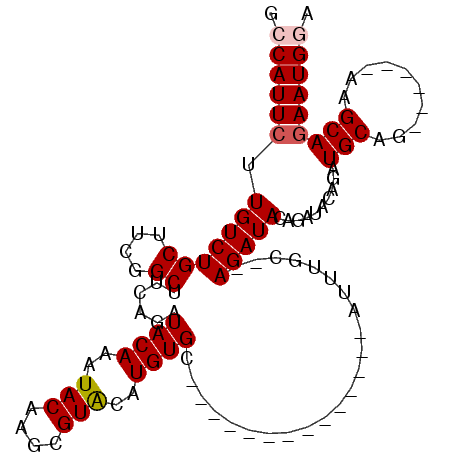
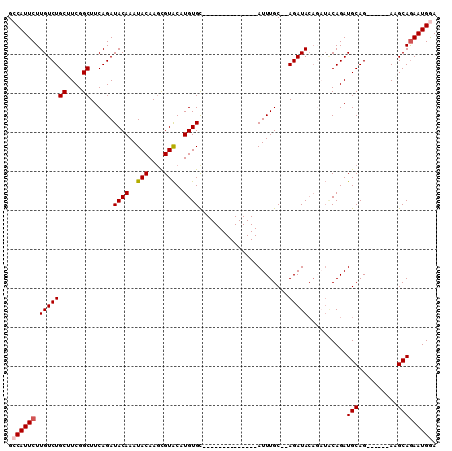
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:59 2006