
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 9,362,822 – 9,362,925 |
| Length | 103 |
| Max. P | 0.972620 |

| Location | 9,362,822 – 9,362,925 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 76.19 |
| Mean single sequence MFE | -24.99 |
| Consensus MFE | -16.28 |
| Energy contribution | -16.98 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.690135 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9362822 103 + 27905053 CCAGCC--CGCCCACAUCAGGCCAUCAACCC-ACUGCCUUU-----CCCGUAAUUGGGUCACUUGUAUGCGAAAAAGCAGCGAACUGUUGCAUGCAAAGUAAAUGCAACAU-------- ......--.(((.......)))........(-.((((.(((-----(.((((...((....))..)))).))))..)))).)...((((((((.........)))))))).-------- ( -23.10) >DroSim_CAF1 141264 103 + 1 CCACCU--CGUCCACAUCAGCCCAUCAACCC-ACUGCCUUU-----CCCGUAAUUGGGUCACUUGUAUGCGAAAAAGCAGCGAACUGUUGCAUGCAAAGUACAUGCAACAU-------- .....(--(((..(((...(((((.......-.........-----........)))))....))).(((......)))))))..(((((((((.......))))))))).-------- ( -23.56) >DroEre_CAF1 132961 96 + 1 -----C--CGCCCACAUCUGCCCACU--UCC-ACUGCCUUU-----CCCGUAAUUGGGUCACUUGUAUGCGAAAAAGCAGCGAACUGUUGCAUGCAAAGUACAUGCAACAU-------- -----.--(((..(((..((((((.(--(.(-.........-----...).)).)))).))..))).(((......))))))...(((((((((.......))))))))).-------- ( -24.60) >DroYak_CAF1 133259 98 + 1 C-A--C--CCCCUACAUUUGCCCACU--CCC-ACUGCCUUU-----GCCGUAAUUGGGUCACUUGUAUGCGAAAAAGCAGCGAACUGUUGCAUGCAAAGUACAUGCAACAU-------- .-.--.--....((((..((((((..--...-((.((....-----)).))...)))).))..))))(((......)))......(((((((((.......))))))))).-------- ( -24.80) >DroAna_CAF1 127310 98 + 1 --------------UACCUGCCCCUU--UUCUGUCCCCUCU-----UGGGUAAUUGGGUCACUUGUAUGCGAAAAAGCAGCGAACUGUUGCAUGCGAAGUACAUGCAACAUCAAGGAUA --------------..(((((((..(--(.....(((....-----.))).))..))))...((((.(((......)))))))..(((((((((.......)))))))))...)))... ( -25.90) >DroPer_CAF1 133145 115 + 1 CCGCCCUGCGACUGCCCCUGCCCCU---UCC-CCUGCCCCUACCCCUCUGUAAUUGGGUCACUUCUAUGCGAAAAAGCAGCGAACUGUUGCAUGCAAAGUACAUGCGACAUCCAGGAGA ....((((((.((((...(((....---...-...((((.(((......)))...)))).........))).....))))))...(((((((((.......)))))))))..))))... ( -28.01) >consensus C_A__C__CGCCCACAUCUGCCCAUU__CCC_ACUGCCUUU_____CCCGUAAUUGGGUCACUUGUAUGCGAAAAAGCAGCGAACUGUUGCAUGCAAAGUACAUGCAACAU________ ...................(((((..............................)))))...((((.(((......)))))))..(((((((((.......)))))))))......... (-16.28 = -16.98 + 0.70)

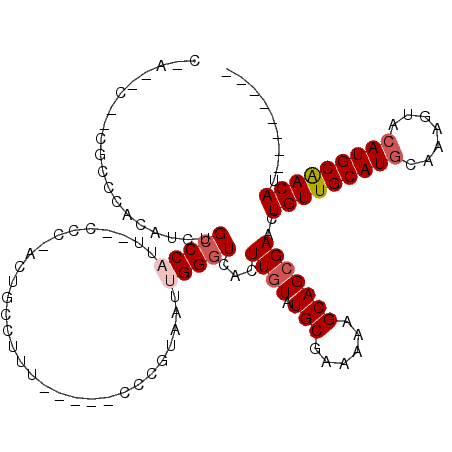

| Location | 9,362,822 – 9,362,925 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 76.19 |
| Mean single sequence MFE | -36.41 |
| Consensus MFE | -21.38 |
| Energy contribution | -22.10 |
| Covariance contribution | 0.73 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.59 |
| SVM decision value | 1.70 |
| SVM RNA-class probability | 0.972620 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9362822 103 - 27905053 --------AUGUUGCAUUUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGG-----AAAGGCAGU-GGGUUGAUGGCCUGAUGUGGGCG--GGCUGG --------.(((((((.......)))).))).(((..((((((((((((((......))))(((......)))-----)))))))))-)..)))..((((((.......))--)))).. ( -38.60) >DroSim_CAF1 141264 103 - 1 --------AUGUUGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGG-----AAAGGCAGU-GGGUUGAUGGGCUGAUGUGGACG--AGGUGG --------.((((((((((.....))))))))))..(((((((((((((((......))))(((......)))-----)))))))))-))(((.(((......))).))).--...... ( -36.50) >DroEre_CAF1 132961 96 - 1 --------AUGUUGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGG-----AAAGGCAGU-GGA--AGUGGGCAGAUGUGGGCG--G----- --------.((((((((((.....)))))))))).((((((((((((((((......))))(((......)))-----)))))))))-)))--.((..((....))..)).--.----- ( -37.20) >DroYak_CAF1 133259 98 - 1 --------AUGUUGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGC-----AAAGGCAGU-GGG--AGUGGGCAAAUGUAGGGG--G--U-G --------.((((((((((.....))))))))))(..(.((((....((((......))))((((.((((.((-----....)).))-)).--..)))).....)))).).--.--)-. ( -34.30) >DroAna_CAF1 127310 98 - 1 UAUCCUUGAUGUUGCAUGUACUUCGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACCCA-----AGAGGGGACAGAA--AAGGGGCAGGUA-------------- ...(((...((((((((((.....))))))))))....(((.(((((((((......))))........(((.-----...))).....))--))).))))))..-------------- ( -30.80) >DroPer_CAF1 133145 115 - 1 UCUCCUGGAUGUCGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUAGAAGUGACCCAAUUACAGAGGGGUAGGGGCAGG-GGA---AGGGGCAGGGGCAGUCGCAGGGCGG ..((((((((((.((((((.....)))))).)).((((.((((((((((.(.........((((..........))))........)-.))---)))))))))))).))).)))))... ( -41.03) >consensus ________AUGUUGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGG_____AAAGGCAGU_GGA__AAUGGGCAGAUGUGGGCG__G__U_G .........((((((((((.....))))))))))(..((((((((((((((......)))).((......))......))))))))).........)..)................... (-21.38 = -22.10 + 0.73)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:37 2006