
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 9,305,194 – 9,305,287 |
| Length | 93 |
| Max. P | 0.658565 |

| Location | 9,305,194 – 9,305,287 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 81.07 |
| Mean single sequence MFE | -26.98 |
| Consensus MFE | -18.27 |
| Energy contribution | -19.33 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.576328 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9305194 93 + 27905053 ---UGAGCAGGCAA---UCC-----GGAUACCCUACGGAAUUGUGGGAGCCUGCUUCGGCAGGAUACAUAUGUCUCUAUGUAUAUUC---------AUGAGCAAUAUAUUUUA ---...((..((((---(((-----(.........))).))))).(((.(((((....)))))(((((((......))))))).)))---------....))........... ( -26.60) >DroPse_CAF1 73687 99 + 1 CGAAGAGCAGAGAAGAGAGC-----CGAGCCCCUUGGGGGUUAAGGAAGCGUGAGCCAGCAGCAUACAUAUGCCUAUAUGUAUAUUC---------AUGAGCAAUAUAUUUUA .(((..((...........(-----(.((((((...))))))..))..((....))..))...((((((((....)))))))).)))---------................. ( -22.60) >DroSim_CAF1 77705 93 + 1 ---UGAGCAGGCAA---UCC-----GGAUACCCUACGGAAUUGUGGGAGCCUGCUCCGGCAGGAUACAUAUGUCUCUAUGUAUAUUC---------AUGAGCAAUAUAUUUUA ---.((((((((..---...-----(....)((((((....)))))).))))))))....((((((.((.((.(((.(((......)---------))))))))).)))))). ( -26.20) >DroEre_CAF1 77187 102 + 1 ---UGAGCGGGCAA---GCC-----GGAUACCCUACGGAAUUGUGGGAGCCUGCUCCGGCAGGAUACAUAUGUCUCUAUGUAUAUUCAUGAGAUUCAUGAGCAAUAUAUUUUA ---.((((((((..---...-----(....)((((((....)))))).))))))))..((...(((((((......)))))))..(((((.....)))))))........... ( -31.60) >DroYak_CAF1 77187 93 + 1 ---UGAGCAGGCAA---UCC-----GGAUACCCUACGGAAUUGUGGGAGCCUGCUCCGGCAGGAUACAUAUGUCUCUAUGUAUAUUC---------AUGAGCAAUAUAUUUUA ---.((((((((..---...-----(....)((((((....)))))).))))))))....((((((.((.((.(((.(((......)---------))))))))).)))))). ( -26.20) >DroAna_CAF1 75184 97 + 1 ---UGAGCACUCG----UCUGGAAAGGAAGCCCGGCGGGAUUGCAGGAGCGUGCUCCGGCAGGAUACAUAUGUCUAUAUGUAUAUUC---------AUGAGCAAUAUAUUUUA ---...((.((((----((.((........)).)))))).((((.((((....)))).)))).((((((((....))))))))....---------....))........... ( -28.70) >consensus ___UGAGCAGGCAA___UCC_____GGAUACCCUACGGAAUUGUGGGAGCCUGCUCCGGCAGGAUACAUAUGUCUCUAUGUAUAUUC_________AUGAGCAAUAUAUUUUA ....((((((((.....(((.....)))...((((((....)))))).)))))))).......(((((((......))))))).............................. (-18.27 = -19.33 + 1.06)



| Location | 9,305,194 – 9,305,287 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 81.07 |
| Mean single sequence MFE | -23.78 |
| Consensus MFE | -16.03 |
| Energy contribution | -16.26 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.658565 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9305194 93 - 27905053 UAAAAUAUAUUGCUCAU---------GAAUAUACAUAGAGACAUAUGUAUCCUGCCGAAGCAGGCUCCCACAAUUCCGUAGGGUAUCC-----GGA---UUGCCUGCUCA--- ....((((((..(((((---------(......))).)))..))))))........((.((((((.(((((......)).))).....-----...---..)))))))).--- ( -22.40) >DroPse_CAF1 73687 99 - 1 UAAAAUAUAUUGCUCAU---------GAAUAUACAUAUAGGCAUAUGUAUGCUGCUGGCUCACGCUUCCUUAACCCCCAAGGGGCUCG-----GCUCUCUUCUCUGCUCUUCG ...........((....---------((((((((((((....)))))))))..((((((....))........((((...))))..))-----))....)))...))...... ( -18.80) >DroSim_CAF1 77705 93 - 1 UAAAAUAUAUUGCUCAU---------GAAUAUACAUAGAGACAUAUGUAUCCUGCCGGAGCAGGCUCCCACAAUUCCGUAGGGUAUCC-----GGA---UUGCCUGCUCA--- ....((((((..(((((---------(......))).)))..)))))).........((((((((.(((((......)).))).....-----...---..)))))))).--- ( -25.90) >DroEre_CAF1 77187 102 - 1 UAAAAUAUAUUGCUCAUGAAUCUCAUGAAUAUACAUAGAGACAUAUGUAUCCUGCCGGAGCAGGCUCCCACAAUUCCGUAGGGUAUCC-----GGC---UUGCCCGCUCA--- ...........(((((((.....)))))..(((((((......)))))))...))(((.((((((((((((......)).))).....-----)))---)))))))....--- ( -28.30) >DroYak_CAF1 77187 93 - 1 UAAAAUAUAUUGCUCAU---------GAAUAUACAUAGAGACAUAUGUAUCCUGCCGGAGCAGGCUCCCACAAUUCCGUAGGGUAUCC-----GGA---UUGCCUGCUCA--- ....((((((..(((((---------(......))).)))..)))))).........((((((((.(((((......)).))).....-----...---..)))))))).--- ( -25.90) >DroAna_CAF1 75184 97 - 1 UAAAAUAUAUUGCUCAU---------GAAUAUACAUAUAGACAUAUGUAUCCUGCCGGAGCACGCUCCUGCAAUCCCGCCGGGCUUCCUUUCCAGA----CGAGUGCUCA--- ...........((....---------....((((((((....))))))))..(((.((((....)))).))).....)).(((((((.((....))----.))).)))).--- ( -21.40) >consensus UAAAAUAUAUUGCUCAU_________GAAUAUACAUAGAGACAUAUGUAUCCUGCCGGAGCAGGCUCCCACAAUUCCGUAGGGUAUCC_____GGA___UUGCCUGCUCA___ ..............................(((((((......))))))).......((((((((.(((((......)).)))..................)))))))).... (-16.03 = -16.26 + 0.23)


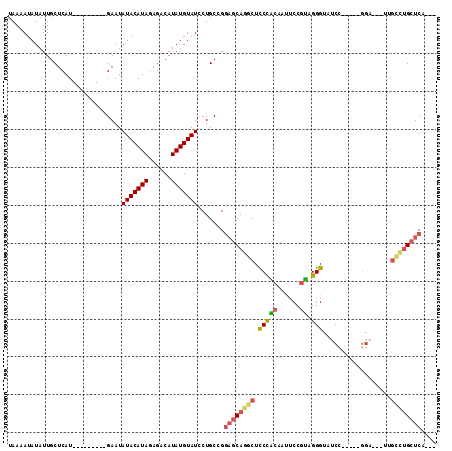
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:00:48 2006