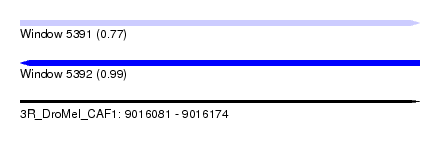

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 9,016,081 – 9,016,174 |
| Length | 93 |
| Max. P | 0.993468 |

| Location | 9,016,081 – 9,016,174 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 91.07 |
| Mean single sequence MFE | -29.89 |
| Consensus MFE | -22.18 |
| Energy contribution | -22.98 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.773052 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9016081 93 + 27905053 UCGGCUUUACCACUUUUUAUUGCGUAUACGCUGGGGCACCCCCAUUACGUAUACGCAGUGUAUUACGCCCCUCCACUCACAGCAGGCAAAUGG .........(((.....(((((((((((((.(((((...)))))...)))))))))))))......(((.((........))..)))...))) ( -29.30) >DroSec_CAF1 130095 93 + 1 UCGGCGUUACCACUUUUUAUUGCGUAUACGCCGUGGCGGCCCCAUCGCGUAUACGCAGUGUAUUACGCCCCUCCAGUCACAGCAGGCAAAUGG ..(((((..........((((((((((((((.((((.....)))).))))))))))))))....)))))((....(((......)))....)) ( -34.24) >DroSim_CAF1 133319 93 + 1 UCGGCGUUACCACUUUUUAUUGCGUACACGCCGGGGCGGACCCAUCGCGUAUACGCAGUGUAUUACGCCCCUCCAGUCACAGCAGACAAAUGG ..((((((((((........)).))).)))))((((((((..(((.(((....))).)))..)).))))))....(((......)))...... ( -32.80) >DroEre_CAF1 136145 92 + 1 UUGGCCUUACCACUUUUUAUUGCGUAUACGCAGGG-AGGCCCCAUCGCGUAUACGCAGUGUAUUACGCCCCUCCAGUCACAGCAUGCAAAUGG ((((.............((((((((((((((.(((-....)))...))))))))))))))............))))......(((....))). ( -27.21) >DroYak_CAF1 141563 91 + 1 UCGGCCUUACCACUUUUUAUUGCGUAUACGCAGUG-AGGGCCCAUCGCGUAUACGCAGUGUAUUACGCCCC-CCAGUCACAGCAUGCAAAUGG ..(((..(((........(((((((((((((.(((-......))).))))))))))))))))....)))..-..........(((....))). ( -25.90) >consensus UCGGCCUUACCACUUUUUAUUGCGUAUACGCAGGGGCGGCCCCAUCGCGUAUACGCAGUGUAUUACGCCCCUCCAGUCACAGCAGGCAAAUGG ..(((.............(((((((((((((.(((.....)))...)))))))))))))(.....))))........................ (-22.18 = -22.98 + 0.80)



| Location | 9,016,081 – 9,016,174 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 91.07 |
| Mean single sequence MFE | -35.11 |
| Consensus MFE | -28.61 |
| Energy contribution | -28.53 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.81 |
| SVM decision value | 2.40 |
| SVM RNA-class probability | 0.993468 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
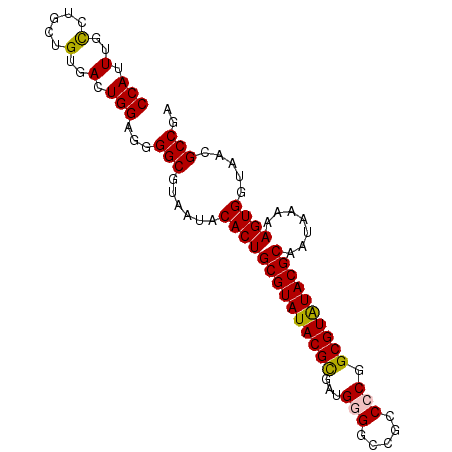
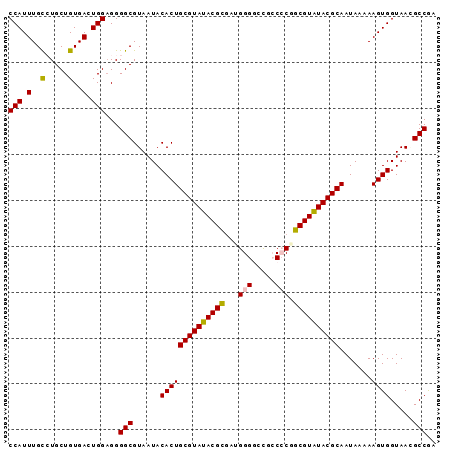
>3R_DroMel_CAF1 9016081 93 - 27905053 CCAUUUGCCUGCUGUGAGUGGAGGGGCGUAAUACACUGCGUAUACGUAAUGGGGGUGCCCCAGCGUAUACGCAAUAAAAAGUGGUAAAGCCGA (((((..(.....)..)))))...(((......((((((((((((((..(((((...))))))))))))))).......)))).....))).. ( -37.71) >DroSec_CAF1 130095 93 - 1 CCAUUUGCCUGCUGUGACUGGAGGGGCGUAAUACACUGCGUAUACGCGAUGGGGCCGCCACGGCGUAUACGCAAUAAAAAGUGGUAACGCCGA .......(((.(.......).)))(((((....((((((((((((((..(((.....)))..)))))))))).......))))...))))).. ( -35.41) >DroSim_CAF1 133319 93 - 1 CCAUUUGUCUGCUGUGACUGGAGGGGCGUAAUACACUGCGUAUACGCGAUGGGUCCGCCCCGGCGUGUACGCAAUAAAAAGUGGUAACGCCGA ((.(..(((......)))..).))(((((....((((((((((((((...(((....)))..)))))))))).......))))...))))).. ( -37.11) >DroEre_CAF1 136145 92 - 1 CCAUUUGCAUGCUGUGACUGGAGGGGCGUAAUACACUGCGUAUACGCGAUGGGGCCU-CCCUGCGUAUACGCAAUAAAAAGUGGUAAGGCCAA (((.(..(.....)..).)))...(((.(....(((((((((((((((..(((....-)))))))))))))).......))))...).))).. ( -33.31) >DroYak_CAF1 141563 91 - 1 CCAUUUGCAUGCUGUGACUGG-GGGGCGUAAUACACUGCGUAUACGCGAUGGGCCCU-CACUGCGUAUACGCAAUAAAAAGUGGUAAGGCCGA (((.(..(.....)..).)))-..(((.(....(((((((((((((((.(((....)-)).))))))))))).......))))...).))).. ( -32.01) >consensus CCAUUUGCCUGCUGUGACUGGAGGGGCGUAAUACACUGCGUAUACGCGAUGGGGCCGCCCCGGCGUAUACGCAAUAAAAAGUGGUAACGCCGA (((.(..(.....)..).)))...(((......((((((((((((((...(((.....))).)))))))))).......)))).....))).. (-28.61 = -28.53 + -0.08)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:58:58 2006