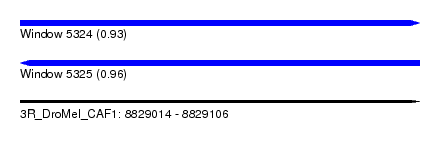

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 8,829,014 – 8,829,106 |
| Length | 92 |
| Max. P | 0.964186 |

| Location | 8,829,014 – 8,829,106 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 79.70 |
| Mean single sequence MFE | -46.20 |
| Consensus MFE | -34.03 |
| Energy contribution | -35.40 |
| Covariance contribution | 1.37 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.927114 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8829014 92 + 27905053 GAGGAC---GACAGUCUACUGCGUCUGGACAGCCUCUGCAAGAGCCUGCAGAGGCACACGGAUGCCACAGACGCCUCGGCUGCG------------------GCUGCUGCUCU ((((..---....((((...(((((((....(((((((((......)))))))))...)))))))...)))).))))(((.((.------------------...)).))).. ( -41.40) >DroSec_CAF1 3358 95 + 1 GAGGACGACGACAGUCUACUGCGUCUGGACAGCCUCUGCAAGAGCCUGCAGAGGCACACGGAUGCCACAGACGCCUCGGCUGCG------------------GCUGCUGCUCU (((.(((.((.(((((....(((((((....(((((((((......)))))))))....((...)).)))))))...)))))))------------------..)).).))). ( -41.90) >DroEre_CAF1 3495 113 + 1 GAGGGUGAUAACAGUCUUUUGCGUCUGGACAGCCUCUGCAAGCGGCUGCAGAUGCACACGGAUGCCACAGAUGCCUCGACUGCGGCAACGGCGGCCGAAGCGGCCGCUGCUGU ((((((....)).((((...(((((((....((.((((((......)))))).))...)))))))...)))).))))....((((((...(((((((...))))))))))))) ( -49.80) >DroYak_CAF1 3420 113 + 1 GAGGGCGACAACAGUCUAUUGCGUCUGGACAGCCUCUGCAAGAGGCUGCAGAUGCACACCGAUGCCACGGACGCCUCGACUGCGGCAACGGCGGCCGAAGCGGCAGCUGCUGU ...((((((....)))...((((((((..(((((((.....)))))))))))))))(.(((.((((.(((.((...)).))).)))).))).))))...(((((....))))) ( -51.70) >consensus GAGGACGACAACAGUCUACUGCGUCUGGACAGCCUCUGCAAGAGCCUGCAGAGGCACACGGAUGCCACAGACGCCUCGACUGCG__________________GCUGCUGCUCU ((((.........((((...(((((((....(((((((((......)))))))))...)))))))...)))).))))(((.((.((..((.....))..)).)).)))..... (-34.03 = -35.40 + 1.37)

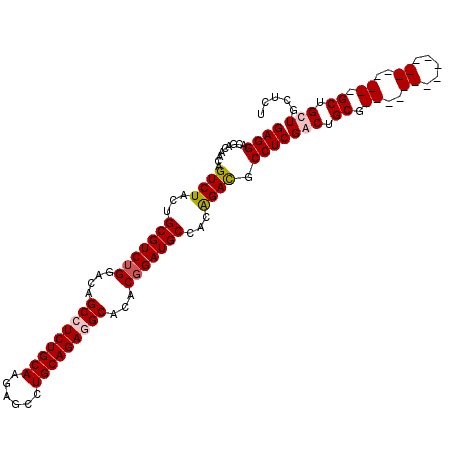
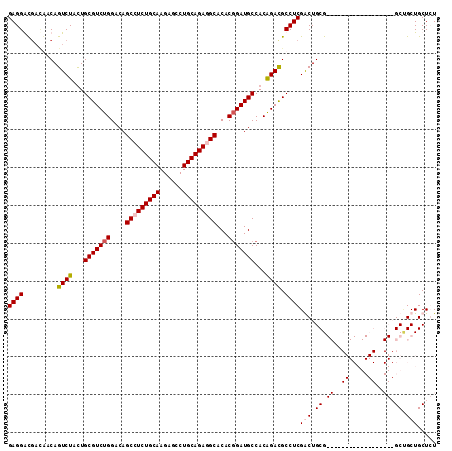
| Location | 8,829,014 – 8,829,106 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 79.70 |
| Mean single sequence MFE | -47.12 |
| Consensus MFE | -33.66 |
| Energy contribution | -35.16 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.71 |
| SVM decision value | 1.56 |
| SVM RNA-class probability | 0.964186 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8829014 92 - 27905053 AGAGCAGCAGC------------------CGCAGCCGAGGCGUCUGUGGCAUCCGUGUGCCUCUGCAGGCUCUUGCAGAGGCUGUCCAGACGCAGUAGACUGUC---GUCCUC .(..((((.((------------------(((((.((...)).)))))))..........((((((((....))))))))))))..).(((((((....))).)---)))... ( -41.60) >DroSec_CAF1 3358 95 - 1 AGAGCAGCAGC------------------CGCAGCCGAGGCGUCUGUGGCAUCCGUGUGCCUCUGCAGGCUCUUGCAGAGGCUGUCCAGACGCAGUAGACUGUCGUCGUCCUC .(..((((.((------------------(((((.((...)).)))))))..........((((((((....))))))))))))..).(((((((....))).))))...... ( -42.50) >DroEre_CAF1 3495 113 - 1 ACAGCAGCGGCCGCUUCGGCCGCCGUUGCCGCAGUCGAGGCAUCUGUGGCAUCCGUGUGCAUCUGCAGCCGCUUGCAGAGGCUGUCCAGACGCAAAAGACUGUUAUCACCCUC ...((((((((.((....)).)))))))).((((((...((.((((.((((.(((....).(((((((....))))))))).)))))))).))....)))))).......... ( -52.10) >DroYak_CAF1 3420 113 - 1 ACAGCAGCUGCCGCUUCGGCCGCCGUUGCCGCAGUCGAGGCGUCCGUGGCAUCGGUGUGCAUCUGCAGCCUCUUGCAGAGGCUGUCCAGACGCAAUAGACUGUUGUCGCCCUC ((((((((((..((....))(((((.((((((.(.((...)).).)))))).)))))(((.(((((((((((.....)))))))..)))).))).))).)))))))....... ( -52.30) >consensus ACAGCAGCAGC__________________CGCAGCCGAGGCGUCUGUGGCAUCCGUGUGCAUCUGCAGCCUCUUGCAGAGGCUGUCCAGACGCAAUAGACUGUCGUCGCCCUC ..............................((((((...(((((((.((((.......((((((((((....)))))))))))))))))))))....)))))).......... (-33.66 = -35.16 + 1.50)
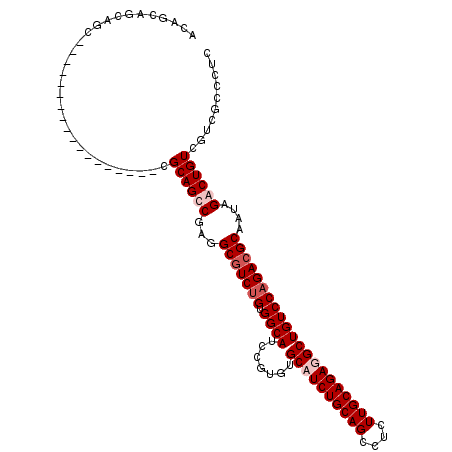


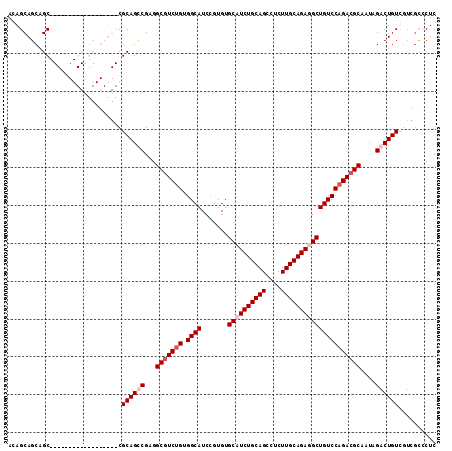
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:57:53 2006