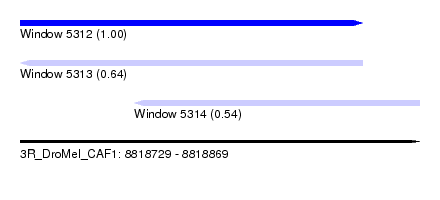

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 8,818,729 – 8,818,869 |
| Length | 140 |
| Max. P | 0.995553 |

| Location | 8,818,729 – 8,818,849 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.33 |
| Mean single sequence MFE | -39.53 |
| Consensus MFE | -33.44 |
| Energy contribution | -34.20 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.85 |
| SVM decision value | 2.59 |
| SVM RNA-class probability | 0.995553 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8818729 120 + 27905053 AUCCUUAUAGCCAUCGCCGCGAUUGUAAAAGUCCAACUGCUUCCACUUGAAGACCUCAAUCAUGGGAUCUCCAUCUGCAGAGAUUAGCAAUUGGAACCCUGAGAUUGCGCACAGGAGCAG .((((....((....))(((((((.......(((((.((((.(((.((((.....))))...)))((((((........)))))))))).))))).......)))))))...)))).... ( -31.04) >DroSec_CAF1 238725 120 + 1 AUCCUUAUAGCCAUCGCCGCGAUUGUAAAAGUCCAGUUGCUUCCACUUGAAGACCUCAAUCAUGGGAUCUCCAUCUGCGGAGAUUAGCAGUUGGAAACCUGCGAUUGCGCAGAGGAGCAG .(((((...((....))((((((((((....((((..((((.(((.((((.....))))...)))(((((((......)))))))))))..))))....))))))))))..))))).... ( -42.20) >DroSim_CAF1 257001 120 + 1 AUCCUUAUAGCCAUCGCCGCGAUUGUAAAAGUCCAGUUGCUUCCACUUGAAGACCUCAAUCAUGGGAUCUCCAUCUGCGGAGAUUAGCAGUUGGAAACCUGCGAUUGCGCAGAGGAGCAG .(((((...((....))((((((((((....((((..((((.(((.((((.....))))...)))(((((((......)))))))))))..))))....))))))))))..))))).... ( -42.20) >DroEre_CAF1 258968 120 + 1 AUCCAUAUAGUCAUCUCCGCGGUUGUAAAAGUCCAGCUGCUUCCACUUGAAAACCUCUAUCAUAGGAUCUCCGUCUGCCGAGAUUAGCAGUUGGAAACCUGCAAUUGCGCAGAUGAGCAG ..........((((((.((((((((((....(((((((((((((...(((.........)))..)))((((.(....).))))..))))))))))....)))))))))).)))))).... ( -43.90) >DroYak_CAF1 252036 120 + 1 AUCCAUAUAGCCAUCCUUGCCAUUGUAAAAGUCCAGCUGCUUCCACUUGAAGACCUCUAUCAUGGGAUCUCCAUCUGCGGAGAUUAGCAAGUGGAGCCCUGCGAUUGCGCAGAGGAGCAG .........((..(((((.....(((...(((((((..(((.(((((((....((........))(((((((......)))))))..)))))))))).))).))))..)))))))))).. ( -38.30) >consensus AUCCUUAUAGCCAUCGCCGCGAUUGUAAAAGUCCAGCUGCUUCCACUUGAAGACCUCAAUCAUGGGAUCUCCAUCUGCGGAGAUUAGCAGUUGGAAACCUGCGAUUGCGCAGAGGAGCAG .........(((.((..((((((((((....(((((((((((((...(((.........)))..)))(((((......)))))..))))))))))....))))))))))..)).).)).. (-33.44 = -34.20 + 0.76)



| Location | 8,818,729 – 8,818,849 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.33 |
| Mean single sequence MFE | -39.28 |
| Consensus MFE | -30.18 |
| Energy contribution | -30.98 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.637287 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8818729 120 - 27905053 CUGCUCCUGUGCGCAAUCUCAGGGUUCCAAUUGCUAAUCUCUGCAGAUGGAGAUCCCAUGAUUGAGGUCUUCAAGUGGAAGCAGUUGGACUUUUACAAUCGCGGCGAUGGCUAUAAGGAU ....((((..(((.......(((((.(((((((((....((..(...((((((((.((....)).)))))))).)..))))))))))))))))......)))(((....)))...)))). ( -37.72) >DroSec_CAF1 238725 120 - 1 CUGCUCCUCUGCGCAAUCGCAGGUUUCCAACUGCUAAUCUCCGCAGAUGGAGAUCCCAUGAUUGAGGUCUUCAAGUGGAAGCAACUGGACUUUUACAAUCGCGGCGAUGGCUAUAAGGAU ....((((....((.(((((.((((((((.((((........)))).)))))))).(.((((((((((((.....((....))...))))))...)))))).)))))).))....)))). ( -40.20) >DroSim_CAF1 257001 120 - 1 CUGCUCCUCUGCGCAAUCGCAGGUUUCCAACUGCUAAUCUCCGCAGAUGGAGAUCCCAUGAUUGAGGUCUUCAAGUGGAAGCAACUGGACUUUUACAAUCGCGGCGAUGGCUAUAAGGAU ....((((....((.(((((.((((((((.((((........)))).)))))))).(.((((((((((((.....((....))...))))))...)))))).)))))).))....)))). ( -40.20) >DroEre_CAF1 258968 120 - 1 CUGCUCAUCUGCGCAAUUGCAGGUUUCCAACUGCUAAUCUCGGCAGACGGAGAUCCUAUGAUAGAGGUUUUCAAGUGGAAGCAGCUGGACUUUUACAACCGCGGAGAUGACUAUAUGGAU ....((((((.(((.(((..((((((((..(((((......)))))..))))).)))..)))...((((.....(((((((........)))))))))))))).)))))).......... ( -38.80) >DroYak_CAF1 252036 120 - 1 CUGCUCCUCUGCGCAAUCGCAGGGCUCCACUUGCUAAUCUCCGCAGAUGGAGAUCCCAUGAUAGAGGUCUUCAAGUGGAAGCAGCUGGACUUUUACAAUGGCAAGGAUGGCUAUAUGGAU ....(((.(((((....)))))((((...((((((((((((((....)))))))....((.(((((((((...((((....)).))))))))))))).)))))))...))))....))). ( -39.50) >consensus CUGCUCCUCUGCGCAAUCGCAGGUUUCCAACUGCUAAUCUCCGCAGAUGGAGAUCCCAUGAUUGAGGUCUUCAAGUGGAAGCAGCUGGACUUUUACAAUCGCGGCGAUGGCUAUAAGGAU ((((..(((((((....))))))).((((.(((((....(((((...((((((((.(......).)))))))).)))))))))).))))...........))))................ (-30.18 = -30.98 + 0.80)
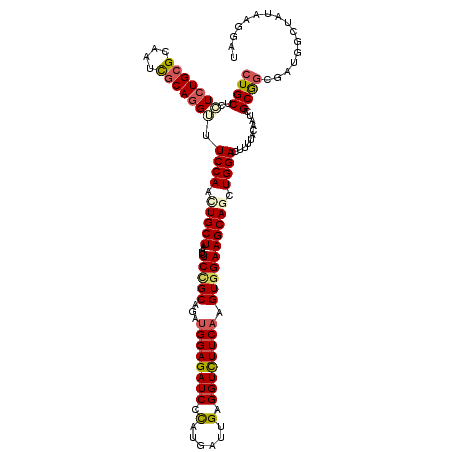


| Location | 8,818,769 – 8,818,869 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.01 |
| Mean single sequence MFE | -33.14 |
| Consensus MFE | -25.02 |
| Energy contribution | -26.66 |
| Covariance contribution | 1.64 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.538927 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8818769 100 - 27905053 --------------------AAAUGUUAUCGCUGGACGUUCUGCUCCUGUGCGCAAUCUCAGGGUUCCAAUUGCUAAUCUCUGCAGAUGGAGAUCCCAUGAUUGAGGUCUUCAAGUGGAA --------------------........(((((((((.....((......)).(((((...((((((((.((((........)))).)))))..)))..)))))..))))...))))).. ( -28.10) >DroSec_CAF1 238765 120 - 1 GUUUUUAAACGAAGCUAUAUAAAUGUUAUCGCUGGAGGUUCUGCUCCUCUGCGCAAUCGCAGGUUUCCAACUGCUAAUCUCCGCAGAUGGAGAUCCCAUGAUUGAGGUCUUCAAGUGGAA ..........((((((.............(((.(((((.......))))))))((((((..((((((((.((((........)))).)))))).))..)))))).)).))))........ ( -35.60) >DroSim_CAF1 257041 113 - 1 -------AACGAAGCCAUAUAAAUGUUAUCGCUGGAGGUUCUGCUCCUCUGCGCAAUCGCAGGUUUCCAACUGCUAAUCUCCGCAGAUGGAGAUCCCAUGAUUGAGGUCUUCAAGUGGAA -------...((((((.............(((.(((((.......))))))))((((((..((((((((.((((........)))).)))))).))..)))))).)).))))........ ( -38.20) >DroEre_CAF1 259008 113 - 1 GUUU-----CAUAUACA--UAAAUGUUAUCGCUGAAGUUUCUGCUCAUCUGCGCAAUUGCAGGUUUCCAACUGCUAAUCUCGGCAGACGGAGAUCCUAUGAUAGAGGUUUUCAAGUGGAA ....-----........--.........(((((((((..((((.(((((((((....)))))((((((..(((((......)))))..))))))...))))))))..))))..))))).. ( -30.10) >DroYak_CAF1 252076 120 - 1 GUGUUUAAACAAAGCCAUAAAAAUGUUAUCGCUGCACGUUCUGCUCCUCUGCGCAAUCGCAGGGCUCCACUUGCUAAUCUCCGCAGAUGGAGAUCCCAUGAUAGAGGUCUUCAAGUGGAA ((((........(((.((((.....)))).))))))).(((..((((((((((....))).((((((((.((((........)))).)))))..))).....)))))......))..))) ( -33.70) >consensus GU_U___AACAAAGCCAUAUAAAUGUUAUCGCUGGAGGUUCUGCUCCUCUGCGCAAUCGCAGGUUUCCAACUGCUAAUCUCCGCAGAUGGAGAUCCCAUGAUUGAGGUCUUCAAGUGGAA ............................(((((.((((.(((((......)).((((((..((((((((.((((........)))).))))))))...))))))))).)))).))))).. (-25.02 = -26.66 + 1.64)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:57:43 2006