
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 8,817,481 – 8,817,681 |
| Length | 200 |
| Max. P | 0.999963 |

| Location | 8,817,481 – 8,817,601 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.17 |
| Mean single sequence MFE | -38.08 |
| Consensus MFE | -34.16 |
| Energy contribution | -34.76 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.90 |
| SVM decision value | 4.01 |
| SVM RNA-class probability | 0.999756 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
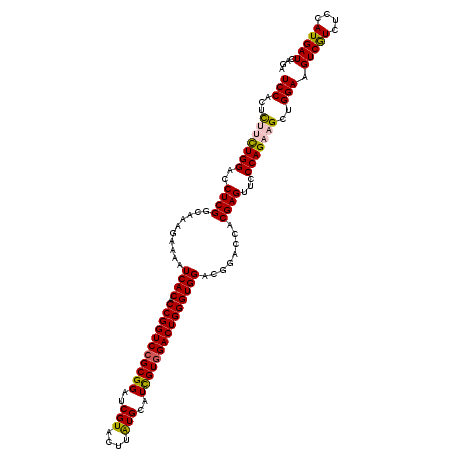
>3R_DroMel_CAF1 8817481 120 + 27905053 UAAAUCAUCUCCGAUGAAUUGGAAUAAACGGAGCCAUGGUUUUCCGUUGAGAACGGCUUGCUGGUUUUCCAGCAGCCCACUCCACUCUUCUGGACCUCGGCAAAGAAAAUCACCCCGGUC .........(((((....))))).....(((.(...((((((((.((((((...((((.(((((....)))))))))...((((......)))).))))))...)))))))).))))... ( -37.90) >DroSec_CAF1 237458 120 + 1 UAAAUCAUCUCCGAUGAAUUGGAAUAAACGGAGCCAUGGUUUUCGGUUGAGAACGUCUUGCUGGUUUUCCAGCAGCCCACUCCACUCUUUUGGACCUCGGCAAAGAAAAUCACACCGGUC .........(((((....))))).....(((.(...((((((((.((((((...(..(((((((....)))))))..)..((((......)))).))))))...))))))))).)))... ( -35.80) >DroSim_CAF1 255750 120 + 1 UAAAUCAUCUCCGAUGAAUUGGAAUAAACGGAGCCAUGGUUUUCGGUUGAGAACGGCUUGCUGGUUUUCCAGCAGCCCACUCCACUUUUCUGGACCUCGGCAAAGAAAAUCACACCGGUC .........(((((....))))).....(((.(...((((((((.((((((...((((.(((((....)))))))))...((((......)))).))))))...))))))))).)))... ( -39.90) >DroEre_CAF1 257525 120 + 1 UAAAUCAUCUCUGAUGAAUUGGAAAAAACGGAGCCAUGGUUUUCAGUUGAGAAAGGCUUGCUGGUUUUCCAGCAGCCCACUCCACUCUUCUGGACCUCGGCAAAGAAAAUCACUCCGGUC ....(((((...)))))...........(((((...((((((((.((((((...((((.(((((....)))))))))...((((......)))).))))))...)))))))))))))... ( -41.90) >DroYak_CAF1 250654 120 + 1 UAAAUCAUCUCCGAUGAAUUUGAAUAAACAGAGCCAUGGUUUUCCGUUGAAAAAGGCUUGCUGGUUUUCCAGCAGCCUACUCCACUCUUCUGGACCUCGGCAAAGAAAAUCACUCCGGUC ..........((((((..((((......))))..)))(((((((.(((((...(((((.(((((....))))))))))..((((......))))..)))))...)))))))....))).. ( -34.90) >consensus UAAAUCAUCUCCGAUGAAUUGGAAUAAACGGAGCCAUGGUUUUCCGUUGAGAACGGCUUGCUGGUUUUCCAGCAGCCCACUCCACUCUUCUGGACCUCGGCAAAGAAAAUCACACCGGUC ....(((((...)))))...........(((.(...((((((((.((((((...((((.(((((....)))))))))...((((......)))).))))))...))))))))).)))... (-34.16 = -34.76 + 0.60)



| Location | 8,817,481 – 8,817,601 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.17 |
| Mean single sequence MFE | -37.90 |
| Consensus MFE | -31.78 |
| Energy contribution | -31.50 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.820654 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8817481 120 - 27905053 GACCGGGGUGAUUUUCUUUGCCGAGGUCCAGAAGAGUGGAGUGGGCUGCUGGAAAACCAGCAAGCCGUUCUCAACGGAAAACCAUGGCUCCGUUUAUUCCAAUUCAUCGGAGAUGAUUUA ((((..((..(......)..))..))))..((.(((((((((((((((((((....))))).))))......((((((..........)))))))))))).)))).))............ ( -42.40) >DroSec_CAF1 237458 120 - 1 GACCGGUGUGAUUUUCUUUGCCGAGGUCCAAAAGAGUGGAGUGGGCUGCUGGAAAACCAGCAAGACGUUCUCAACCGAAAACCAUGGCUCCGUUUAUUCCAAUUCAUCGGAGAUGAUUUA ..((((((.....((((((((....))...))))))((((((((((((((((....))))))....(((.((....)).))).........))))))))))...)))))).......... ( -32.60) >DroSim_CAF1 255750 120 - 1 GACCGGUGUGAUUUUCUUUGCCGAGGUCCAGAAAAGUGGAGUGGGCUGCUGGAAAACCAGCAAGCCGUUCUCAACCGAAAACCAUGGCUCCGUUUAUUCCAAUUCAUCGGAGAUGAUUUA ..((((((...((((((..((....))..)))))).((((((((((((((((....))))))(((((((.((....)).)))...))))..))))))))))...)))))).......... ( -39.30) >DroEre_CAF1 257525 120 - 1 GACCGGAGUGAUUUUCUUUGCCGAGGUCCAGAAGAGUGGAGUGGGCUGCUGGAAAACCAGCAAGCCUUUCUCAACUGAAAACCAUGGCUCCGUUUUUUCCAAUUCAUCAGAGAUGAUUUA ...((((((.(((((((..((....))..)))))(((.(((.((((((((((....))))).)))))..))).))).......)).))))))...........(((((...))))).... ( -38.70) >DroYak_CAF1 250654 120 - 1 GACCGGAGUGAUUUUCUUUGCCGAGGUCCAGAAGAGUGGAGUAGGCUGCUGGAAAACCAGCAAGCCUUUUUCAACGGAAAACCAUGGCUCUGUUUAUUCAAAUUCAUCGGAGAUGAUUUA ((((...(..(......)..)...))))..((((((..((((((((((((((....))))).)))))(((((....))))).....))))..))).)))....(((((...))))).... ( -36.50) >consensus GACCGGAGUGAUUUUCUUUGCCGAGGUCCAGAAGAGUGGAGUGGGCUGCUGGAAAACCAGCAAGCCGUUCUCAACCGAAAACCAUGGCUCCGUUUAUUCCAAUUCAUCGGAGAUGAUUUA ((((...(..(......)..)...)))).....((..(((((.(((((((((....))))).)))).(((......))).......)))))..))........(((((...))))).... (-31.78 = -31.50 + -0.28)



| Location | 8,817,521 – 8,817,641 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.50 |
| Mean single sequence MFE | -48.82 |
| Consensus MFE | -45.04 |
| Energy contribution | -45.24 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.09 |
| Structure conservation index | 0.92 |
| SVM decision value | 4.93 |
| SVM RNA-class probability | 0.999963 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8817521 120 + 27905053 UUUCCGUUGAGAACGGCUUGCUGGUUUUCCAGCAGCCCACUCCACUCUUCUGGACCUCGGCAAAGAAAAUCACCCCGGUCCGCGGAUCGUAUUUAUGCAUUGUGGACUGGGUGGACGGAC ..(((((((((...((((.(((((....)))))))))...((((......)))).))))))........((((((.(((((((((..(((....)))..)))))))))))))))..))). ( -50.30) >DroSec_CAF1 237498 120 + 1 UUUCGGUUGAGAACGUCUUGCUGGUUUUCCAGCAGCCCACUCCACUCUUUUGGACCUCGGCAAAGAAAAUCACACCGGUCCGCGGAUCGUACUUGUGCAUCGUGGACUGGGUGGACGGAC .((((((((((...(..(((((((....)))))))..)..((((......)))).))))))........((((.(((((((((((...(((....))).))))))))))))))).)))). ( -48.00) >DroSim_CAF1 255790 120 + 1 UUUCGGUUGAGAACGGCUUGCUGGUUUUCCAGCAGCCCACUCCACUUUUCUGGACCUCGGCAAAGAAAAUCACACCGGUCCGCGGAUCGUAUUUGUGCAUCGUGGACUGGGUGGACGGAC .((((((((((...((((.(((((....)))))))))...((((......)))).))))))........((((.(((((((((((...(((....))).))))))))))))))).)))). ( -51.20) >DroEre_CAF1 257565 120 + 1 UUUCAGUUGAGAAAGGCUUGCUGGUUUUCCAGCAGCCCACUCCACUCUUCUGGACCUCGGCAAAGAAAAUCACUCCGGUCAGCGGAUCGUACUUAUGUAUCGUGGACUGGGUGGACGGAC .(((.((((((...((((.(((((....)))))))))...((((......)))).))))))........((((.((((((.(((....((((....))))))).))))))))))..))). ( -43.90) >DroYak_CAF1 250694 120 + 1 UUUCCGUUGAAAAAGGCUUGCUGGUUUUCCAGCAGCCUACUCCACUCUUCUGGACCUCGGCAAAGAAAAUCACUCCGGUCCGCGGAUCGUACUUAUGCAUCGUGGACUGGGUGGACGGAC ..((((((((...(((((.(((((....))))))))))..((((......))))..)))))........((((.(((((((((((..(((....)))..)))))))))))))))..))). ( -50.70) >consensus UUUCCGUUGAGAACGGCUUGCUGGUUUUCCAGCAGCCCACUCCACUCUUCUGGACCUCGGCAAAGAAAAUCACACCGGUCCGCGGAUCGUACUUAUGCAUCGUGGACUGGGUGGACGGAC ((((.((((((...((((.(((((....)))))))))...((((......)))).))))))...)))).((((.(((((((((((..(((....)))..)))))))))))))))...... (-45.04 = -45.24 + 0.20)



| Location | 8,817,521 – 8,817,641 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.50 |
| Mean single sequence MFE | -42.37 |
| Consensus MFE | -33.96 |
| Energy contribution | -34.56 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.796619 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8817521 120 - 27905053 GUCCGUCCACCCAGUCCACAAUGCAUAAAUACGAUCCGCGGACCGGGGUGAUUUUCUUUGCCGAGGUCCAGAAGAGUGGAGUGGGCUGCUGGAAAACCAGCAAGCCGUUCUCAACGGAAA .(((((.....((.(((((...((.............))(((((..((..(......)..))..)))))......))))).))(((((((((....))))).)))).......))))).. ( -48.62) >DroSec_CAF1 237498 120 - 1 GUCCGUCCACCCAGUCCACGAUGCACAAGUACGAUCCGCGGACCGGUGUGAUUUUCUUUGCCGAGGUCCAAAAGAGUGGAGUGGGCUGCUGGAAAACCAGCAAGACGUUCUCAACCGAAA (..((((..((((.(((((...((.............))(((((...(..(......)..)...)))))......))))).)))).((((((....)))))).))))..).......... ( -43.22) >DroSim_CAF1 255790 120 - 1 GUCCGUCCACCCAGUCCACGAUGCACAAAUACGAUCCGCGGACCGGUGUGAUUUUCUUUGCCGAGGUCCAGAAAAGUGGAGUGGGCUGCUGGAAAACCAGCAAGCCGUUCUCAACCGAAA (..((....((((.(((((....((((....((.((....)).)).)))).((((((..((....))..))))))))))).)))).((((((....))))))...))..).......... ( -38.80) >DroEre_CAF1 257565 120 - 1 GUCCGUCCACCCAGUCCACGAUACAUAAGUACGAUCCGCUGACCGGAGUGAUUUUCUUUGCCGAGGUCCAGAAGAGUGGAGUGGGCUGCUGGAAAACCAGCAAGCCUUUCUCAACUGAAA ....(((((((((.((..((.(((....))))).....((((((...(..(......)..)...))).)))..)).))).))))))((((((....)))))).....(((......))). ( -36.60) >DroYak_CAF1 250694 120 - 1 GUCCGUCCACCCAGUCCACGAUGCAUAAGUACGAUCCGCGGACCGGAGUGAUUUUCUUUGCCGAGGUCCAGAAGAGUGGAGUAGGCUGCUGGAAAACCAGCAAGCCUUUUUCAACGGAAA .(((((.(((((.((((.((...(........)...)).)))).)).))).((((((..((....))..)))))).(((((.((((((((((....))))).))))).)))))))))).. ( -44.60) >consensus GUCCGUCCACCCAGUCCACGAUGCAUAAGUACGAUCCGCGGACCGGAGUGAUUUUCUUUGCCGAGGUCCAGAAGAGUGGAGUGGGCUGCUGGAAAACCAGCAAGCCGUUCUCAACCGAAA .........((((.(((((...((.............))(((((...(..(......)..)...)))))......))))).)))).((((((....)))))).................. (-33.96 = -34.56 + 0.60)



| Location | 8,817,561 – 8,817,681 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.33 |
| Mean single sequence MFE | -41.19 |
| Consensus MFE | -36.65 |
| Energy contribution | -36.69 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.26 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.709751 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8817561 120 + 27905053 UCCACUCUUCUGGACCUCGGCAAAGAAAAUCACCCCGGUCCGCGGAUCGUAUUUAUGCAUUGUGGACUGGGUGGACGGACCACGAGUUCCCAGAAGAUGGAAGUCGUCUCCAUGAUCAGA ((((.((((((((..((((..............((((((((((((..(((....)))..))))))))))))(((.....)))))))...)))))))))))).(((((....))))).... ( -44.50) >DroSec_CAF1 237538 120 + 1 UCCACUCUUUUGGACCUCGGCAAAGAAAAUCACACCGGUCCGCGGAUCGUACUUGUGCAUCGUGGACUGGGUGGACGGACCACGAGUUCCCAGUCGCCGGAAGUCAUCUCCAUGAUCAGA ((((......))))..(((((.........(((.(((((((((((...(((....))).))))))))))))))((((((.(....).)))..))))))))..(((((....))))).... ( -40.10) >DroSim_CAF1 255830 120 + 1 UCCACUUUUCUGGACCUCGGCAAAGAAAAUCACACCGGUCCGCGGAUCGUAUUUGUGCAUCGUGGACUGGGUGGACGGACCACGAGUUCCCAGUCGCCGGAAGUCAUCUCCAUGAUCAGA ........(((((((.(((((.........(((.(((((((((((...(((....))).))))))))))))))((((((.(....).)))..))))))))..)))(((.....))))))) ( -40.90) >DroEre_CAF1 257605 120 + 1 UCCACUCUUCUGGACCUCGGCAAAGAAAAUCACUCCGGUCAGCGGAUCGUACUUAUGUAUCGUGGACUGGGUGGACGGACCACGAGUUCCCAGAAGCUGGAAGUCGUCUCCAUGAUCAGA ((((..(((((((..((((..........((((.((((((.(((....((((....))))))).))))))))))........))))...))))))).)))).(((((....))))).... ( -37.57) >DroYak_CAF1 250734 120 + 1 UCCACUCUUCUGGACCUCGGCAAAGAAAAUCACUCCGGUCCGCGGAUCGUACUUAUGCAUCGUGGACUGGGUGGACGGACCACGAGUUCCCAGAAGCUGGAAGUCGUCUCCAUGAUCAGA ((((..(((((((..((((..........((((.(((((((((((..(((....)))..)))))))))))))))........))))...))))))).)))).(((((....))))).... ( -42.87) >consensus UCCACUCUUCUGGACCUCGGCAAAGAAAAUCACACCGGUCCGCGGAUCGUACUUAUGCAUCGUGGACUGGGUGGACGGACCACGAGUUCCCAGAAGCUGGAAGUCGUCUCCAUGAUCAGA (((...(((((((..((((..........((((.(((((((((((..(((....)))..)))))))))))))))........))))...)))))))..))).(((((....))))).... (-36.65 = -36.69 + 0.04)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:57:37 2006