
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 1,249,534 – 1,249,627 |
| Length | 93 |
| Max. P | 0.889316 |

| Location | 1,249,534 – 1,249,627 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 79.21 |
| Mean single sequence MFE | -22.72 |
| Consensus MFE | -7.28 |
| Energy contribution | -9.52 |
| Covariance contribution | 2.24 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.32 |
| SVM decision value | 0.96 |
| SVM RNA-class probability | 0.889316 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
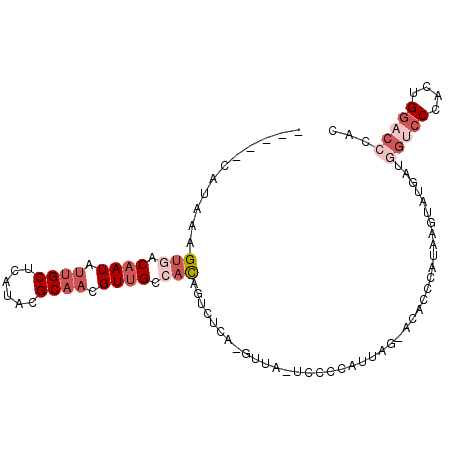
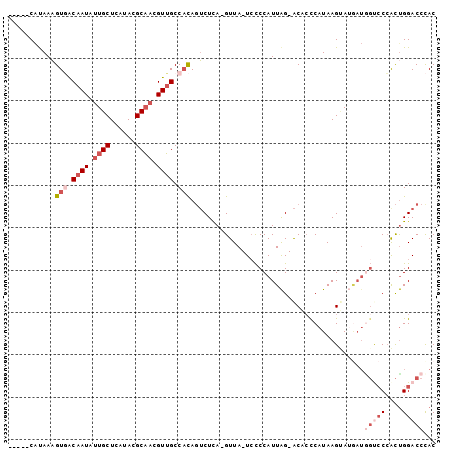
>3R_DroMel_CAF1 1249534 93 + 27905053 -----CAUAAAGUGACAAUAUUGCUCAUACGCAACGUUGCCACAGUCUCA-GUUA-UCCCCAUUAG-ACACCCAUAAGCAUAAUGGUACCGCUGGACCCAC -----......(((.(((..((((......))))..))).))).(((.((-((..-...((((((.-.(........)..))))))....))))))).... ( -23.30) >DroSec_CAF1 24734 93 + 1 -----CAUAAAGUGACAAUAUUGCUCAUACGCAACGUUGCCCCAGUCUCA-GUUA-UCCCCAUUAG-ACACCCAUAAGGAUGAUGGUCCCGCUGGACCCAC -----......(((.(((..((((......))))..))).....(((.((-((..-...((((((.-.(........)..))))))....))))))).))) ( -21.50) >DroEre_CAF1 24077 93 + 1 -----CAUAAAGUGGCAAUAUUGCUUAUACGCAACGUUGCCACAGUCUCA-GUUA-UGCCCAUUAG-ACACCCAUAAGUAAGAUAGCCCCACUGGACCCAC -----......(((((((..((((......))))..))))))).(((.((-((..-.((..(((..-((........))..))).))...))))))).... ( -26.70) >DroYak_CAF1 28194 93 + 1 -----CAUAAAGUUACAAUAUUGCUUAUACGCAACGUUGCCACAGUCUUA-GUUA-UCCCCAUUGG-ACAAUCAUAAGUAAGAUGGCCCGACCGGACCUAC -----......((..(((..((((......))))..)))..)).((((((-.(((-(((.....))-)......))).))))))((.((....)).))... ( -18.00) >DroAna_CAF1 23457 97 + 1 ACAGUCGCAAUGUCGCAAUAUUGCUCAUACGCCCUGUGGCCAUACACUCAAGUUACUCUCCAUUAGGGCGCUCAUAAGUAU----UUCCCAUUGGACCCAC ...(((.(((((..(.((((((.......((((((((((......((....))......))).)))))))......)))))----).).)))))))).... ( -24.12) >consensus _____CAUAAAGUGACAAUAUUGCUCAUACGCAACGUUGCCACAGUCUCA_GUUA_UCCCCAUUAG_ACACCCAUAAGUAUGAUGGUCCCACUGGACCCAC ...........(((.((((.((((......)))).)))).))).........................................(((((....)))))... ( -7.28 = -9.52 + 2.24)

| Location | 1,249,534 – 1,249,627 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 79.21 |
| Mean single sequence MFE | -29.94 |
| Consensus MFE | -15.84 |
| Energy contribution | -17.28 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.718622 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
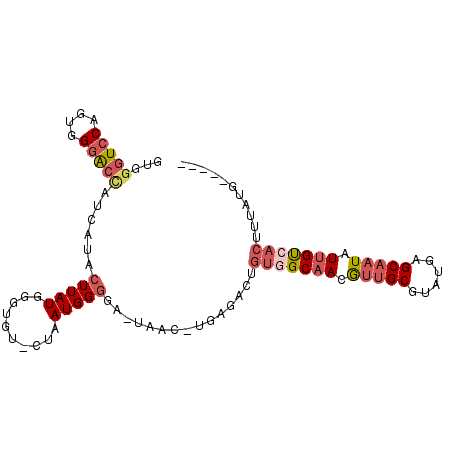
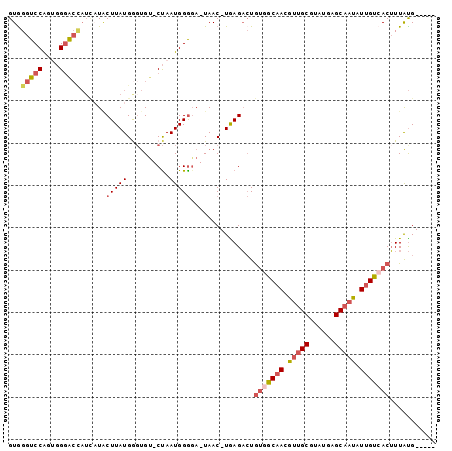
>3R_DroMel_CAF1 1249534 93 - 27905053 GUGGGUCCAGCGGUACCAUUAUGCUUAUGGGUGU-CUAAUGGGGA-UAAC-UGAGACUGUGGCAACGUUGCGUAUGAGCAAUAUUGUCACUUUAUG----- ...(((((((..((.((((((.((........))-.))))))..)-)..)-)).))))(((((((..((((......))))..)))))))......----- ( -31.90) >DroSec_CAF1 24734 93 - 1 GUGGGUCCAGCGGGACCAUCAUCCUUAUGGGUGU-CUAAUGGGGA-UAAC-UGAGACUGGGGCAACGUUGCGUAUGAGCAAUAUUGUCACUUUAUG----- .(.(((((((..(..(((((((((....))))).-...))))...-)..)-)).)))).)(((((..((((......))))..)))))........----- ( -28.60) >DroEre_CAF1 24077 93 - 1 GUGGGUCCAGUGGGGCUAUCUUACUUAUGGGUGU-CUAAUGGGCA-UAAC-UGAGACUGUGGCAACGUUGCGUAUAAGCAAUAUUGCCACUUUAUG----- ...((((((((..(.((((..((((....)))).-...)))).).-..))-)).))))(((((((..((((......))))..)))))))......----- ( -33.60) >DroYak_CAF1 28194 93 - 1 GUAGGUCCGGUCGGGCCAUCUUACUUAUGAUUGU-CCAAUGGGGA-UAAC-UAAGACUGUGGCAACGUUGCGUAUAAGCAAUAUUGUAACUUUAUG----- ...(((((....)))))......((((.(.((((-((.....)))-))))-))))...((.((((..((((......))))..)))).))......----- ( -26.60) >DroAna_CAF1 23457 97 - 1 GUGGGUCCAAUGGGAA----AUACUUAUGAGCGCCCUAAUGGAGAGUAACUUGAGUGUAUGGCCACAGGGCGUAUGAGCAAUAUUGCGACAUUGCGACUGU ...(((((((((...(----((((((((..(((((((..(((...(((((....)).)))..))).))))))))))))...))))....))))).)))).. ( -29.00) >consensus GUGGGUCCAGUGGGACCAUCAUACUUAUGGGUGU_CUAAUGGGGA_UAAC_UGAGACUGUGGCAACGUUGCGUAUGAGCAAUAUUGUCACUUUAUG_____ ...(((((....)))))......(((((..........)))))...............(((((((.(((((......))))).)))))))........... (-15.84 = -17.28 + 1.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:40:28 2006