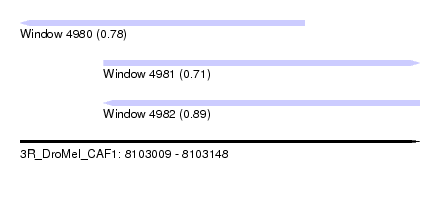

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 8,103,009 – 8,103,148 |
| Length | 139 |
| Max. P | 0.888869 |

| Location | 8,103,009 – 8,103,108 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 89.67 |
| Mean single sequence MFE | -21.07 |
| Consensus MFE | -16.50 |
| Energy contribution | -16.58 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.782803 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8103009 99 - 27905053 CUGAAAUUGUAUUCUGUUGCUCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCACAAAAA----------ACGAAAUUCAACAGCAACAAAAACUUUUGUUGUUAC .((((.((((.((.(((.(((.((..((((((......))))))......))))).))).)).----------))))..))))..(((((((((....))))))))).. ( -23.40) >DroSec_CAF1 54710 104 - 1 CUGAAAUUGUAUUCUGUUGCUCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCAAAAAAAA-----AACAACGAAAUUCAACAGCAACUAAAACUUUUGUUGUUAC ...............((((..((((.((((((......))))))..)))).)))).........-----((((((((((..................)))))))))).. ( -16.47) >DroSim_CAF1 26801 109 - 1 CUGAAAUUGUAUUCUGUUGCUCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCACAAAAAAAAAACAACAACGAAAUUCAACGGCAACAAAAACUUUUGUUGUUAC ..............(((.(((.((..((((((......))))))......))))).)))..........((((((((((......(....)......)))))))))).. ( -21.30) >DroEre_CAF1 29741 107 - 1 CUGAAAUUGUAUUCCGUUGCUCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCACAAAAAAAAG--AAAAAACAAAUUCAACAGCAACAAAAACCUUUGUUGUUAG .((((.((((((((((.....)))))).(((..(((....))).....)))................--.....)))).))))..(((((((((....))))))))).. ( -23.00) >DroYak_CAF1 23274 99 - 1 CUGAAAUUGUAUUCUGUUGCCCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCACAAAA----------AACCAAAUUCAACAGCAAGAAACACUUUUGUUGUUAA ..(((.(((..((.(((.((.((((.((((((......))))))..))))...)).))).))----------...))).)))((((((((((.....)))))))))).. ( -21.20) >consensus CUGAAAUUGUAUUCUGUUGCUCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCACAAAAAA_____AA_AACGAAAUUCAACAGCAACAAAAACUUUUGUUGUUAC .((((...(.((((((.....)))))))(((..(((....))).....)))............................))))..(((((((((....))))))))).. (-16.50 = -16.58 + 0.08)


| Location | 8,103,038 – 8,103,148 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.26 |
| Mean single sequence MFE | -25.87 |
| Consensus MFE | -21.90 |
| Energy contribution | -21.94 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.710984 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8103038 110 + 27905053 AUUUCGU----------UUUUUGUGGCUGGCGAAUUUCACAUUUGUGUACGCGACUCCGAGCAACAGAAUACAAUUUCAGCAUUUUCCAGCCACGACCAGCUCCACUUUCACAAUGAAUC .....((----------(..((((((((((.((((.......((((((.(((........))....).)))))).......)))).))))))))))..)))................... ( -24.84) >DroSec_CAF1 54739 115 + 1 AUUUCGUUGUU-----UUUUUUUUGGCUGGCGAAUUUCACAUUUGUGUACGCGACUCCGAGCAACAGAAUACAAUUUCAGCAUUUUCCAGCCACGACCAGCUCCACUUUCACAAUGAAUC ..((((((((.-----.......(((((((.((((.......((((((.(((........))....).)))))).......)))).))))))).((..((.....)).)))))))))).. ( -23.24) >DroSim_CAF1 26830 120 + 1 AUUUCGUUGUUGUUUUUUUUUUGUGGCUGGCGAAUUUCACAUUUGUGUACGCGACUCCGAGCAACAGAAUACAAUUUCAGCAUUUUCCAGCCACGACCAGCUCCACUUUCACAAUGAAUC ..((((((((.((.......((((((((((.((((.......((((((.(((........))....).)))))).......)))).))))))))))........))....)))))))).. ( -28.40) >DroEre_CAF1 29770 117 + 1 AUUUGUUUUUU--CUUUUUUUUGUGGCUGGCGAAUUUCACAUUUGUGUACGCGACUCCGAGCAACGGAAUACAAUUUCAGCAUUUUCCAGCCACGGCC-GCUCCACUUUCACAAUGAAUC ..((((.....--.........((((((((.((((.......((((((.......((((.....)))))))))).......)))).))))))))((..-...))......))))...... ( -25.45) >DroYak_CAF1 23303 110 + 1 AUUUGGUU----------UUUUGUGGCUGGCGAAUUUCACAUUUGUGUACGCGACUCCGGGCAACAGAAUACAAUUUCAGCAUUUUCCAGCCACGACCAGCUCCACUUUCACAAUGAAUC ..((((((----------....((((((((.((((..(((....)))...((.......(....).(((......))).)))))).)))))))))))))).................... ( -27.40) >consensus AUUUCGUU_UU_____UUUUUUGUGGCUGGCGAAUUUCACAUUUGUGUACGCGACUCCGAGCAACAGAAUACAAUUUCAGCAUUUUCCAGCCACGACCAGCUCCACUUUCACAAUGAAUC ....................((((((((((.((((.......((((((.(((........))....).)))))).......)))).))))))))))........................ (-21.90 = -21.94 + 0.04)



| Location | 8,103,038 – 8,103,148 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.26 |
| Mean single sequence MFE | -29.52 |
| Consensus MFE | -28.26 |
| Energy contribution | -28.30 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.888869 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8103038 110 - 27905053 GAUUCAUUGUGAAAGUGGAGCUGGUCGUGGCUGGAAAAUGCUGAAAUUGUAUUCUGUUGCUCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCACAAAAA----------ACGAAAU ..((((((.....)))))).......((((((((..(((............(((((.....)))))((((((......)))))).)))..)))))))).....----------....... ( -28.90) >DroSec_CAF1 54739 115 - 1 GAUUCAUUGUGAAAGUGGAGCUGGUCGUGGCUGGAAAAUGCUGAAAUUGUAUUCUGUUGCUCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCAAAAAAAA-----AACAACGAAAU ..(((.((((.......(.(((((((((((((....(((((.......)))))(((.....))))))))))..(((....)))......)))))))........-----.)))).))).. ( -26.99) >DroSim_CAF1 26830 120 - 1 GAUUCAUUGUGAAAGUGGAGCUGGUCGUGGCUGGAAAAUGCUGAAAUUGUAUUCUGUUGCUCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCACAAAAAAAAAACAACAACGAAAU ..(((.((((((.((.....))..))((((((((..(((............(((((.....)))))((((((......)))))).)))..))))))))............)))).))).. ( -30.40) >DroEre_CAF1 29770 117 - 1 GAUUCAUUGUGAAAGUGGAGC-GGCCGUGGCUGGAAAAUGCUGAAAUUGUAUUCCGUUGCUCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCACAAAAAAAAG--AAAAAACAAAU ..((((((.....))))))..-....((((((((..(((............(((((.....)))))((((((......)))))).)))..)))))))).........--........... ( -31.10) >DroYak_CAF1 23303 110 - 1 GAUUCAUUGUGAAAGUGGAGCUGGUCGUGGCUGGAAAAUGCUGAAAUUGUAUUCUGUUGCCCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCACAAAA----------AACCAAAU ..((((((.....))))))..((((.((((((((..(((((.......)))))........((((.((((((......))))))..))))))))))))....----------.))))... ( -30.20) >consensus GAUUCAUUGUGAAAGUGGAGCUGGUCGUGGCUGGAAAAUGCUGAAAUUGUAUUCUGUUGCUCGGAGUCGCGUACACAAAUGUGAAAUUCGCCAGCCACAAAAAA_____AA_AACGAAAU ..((((((.....)))))).......((((((((..(((............(((((.....)))))((((((......)))))).)))..))))))))...................... (-28.26 = -28.30 + 0.04)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:52:19 2006