
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 8,101,786 – 8,102,170 |
| Length | 384 |
| Max. P | 0.999387 |

| Location | 8,101,786 – 8,101,906 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.00 |
| Mean single sequence MFE | -29.80 |
| Consensus MFE | -24.50 |
| Energy contribution | -25.26 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.672238 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8101786 120 - 27905053 CCUAUCGUAGCAACCCCCGUAAGAUCUCUGGACAUUGUCCAGAGUCUAUACGAAUUUUCUGCUCGAUUUUCUUAAAAUUUUACAGCAUAUUAUGGCAGCGCCAAAAGGGUUCCGCCUGUC ...((((.((((.....(((((((.((((((((...))))))))))).)))).......))))))))..........................(((.(.(((.....))).).))).... ( -32.30) >DroSec_CAF1 53440 120 - 1 CCUAUGGUAGCAACCCCCGUAAGAUCGCUGGACAUUGUCCAGUGUCUAUACAAAUUUUCUGCUCGAUUUUCUUAAAAUUUUACAGCAUAUUAUGGCAGCGCCAAAAGGGUUCCGCUUGUC .....((..(.(((((..((((((.((((((((...))))))))))).)))........((((.(((((.....)))))....)))).....((((...))))...))))))..)).... ( -29.60) >DroSim_CAF1 25526 120 - 1 CCUAUGGUAGCAACCCCCGUAAGAUCGCUGGACAUUGUCCAGUGUCUAUACGAAUUUUCUGCUCGAUUUUCUUAAAAUUUUACAGCAUAUUAUGGCAGCGCCAAAAGGGUUCCGCUUGUC .....((..(.(((((.(((((((.((((((((...))))))))))).)))).......((((.(((((.....)))))....)))).....((((...))))...))))))..)).... ( -31.40) >DroEre_CAF1 28499 116 - 1 CCU--AGUAGCAACCCCCGAAAUAU--CUGGACAUUGUCCAGCGUCUGUACGAAUUUUCUGCUCGAUUUUCUUGAAAUUUUACAGCAUAUUUGGGCAGCGCCAAAAGGGUUCCGCUUGUC ...--((..(.(((((.........--.(((((...)))))(((.(((..(((((...(((...((((((...))))))...)))...)))))..)))))).....))))))..)).... ( -29.20) >DroYak_CAF1 21920 120 - 1 CCUAUAGUAGCAACCCCCGAAAUAUCUCUGGACAUUGUCCAGAGUCUAUACGAAUUUUCUGCUCGAUUUUCUUAAAAUUUUACAGCAUAUUAUGGCAGCGCCAAAAGGGUUCCGCUUGUC .....((..(.(((((.((......((((((((...))))))))......)).......((((.(((((.....)))))....)))).....((((...))))...))))))..)).... ( -26.50) >consensus CCUAUAGUAGCAACCCCCGUAAGAUCGCUGGACAUUGUCCAGAGUCUAUACGAAUUUUCUGCUCGAUUUUCUUAAAAUUUUACAGCAUAUUAUGGCAGCGCCAAAAGGGUUCCGCUUGUC .....((..(.(((((.((((....((((((((...))))))))....)))).......((((.((((((...))))))....)))).....((((...))))...))))))..)).... (-24.50 = -25.26 + 0.76)


| Location | 8,101,866 – 8,101,970 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.78 |
| Mean single sequence MFE | -44.30 |
| Consensus MFE | -32.24 |
| Energy contribution | -32.84 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.93 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.89 |
| SVM RNA-class probability | 0.981349 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
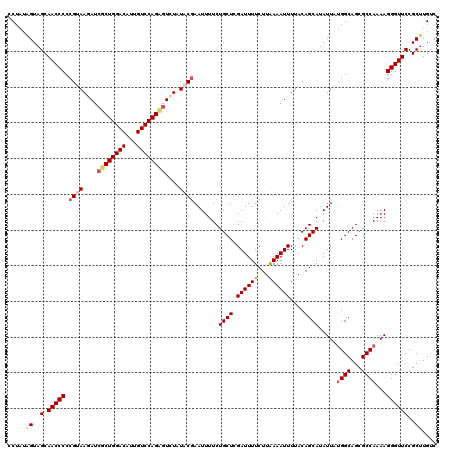
>3R_DroMel_CAF1 8101866 104 + 27905053 GGACAAUGUCCAGAGAUCUUACGGGGGUUGCUACGAUAGGGACUCGAGCUUGAGUCUGCC----UUCGGCAG-----AGGUUAAGUCCAUCGCUGGACACUCGAA-------CUCGAGUU ((((...))))......((((((..(....)..)).))))((((((((.(((((((((((----...)))))-----)......(((((....))))).))))).-------)))))))) ( -39.60) >DroSec_CAF1 53520 104 + 1 GGACAAUGUCCAGCGAUCUUACGGGGGUUGCUACCAUAGGGACUCGAGCUUGCUUCUGCC----UUCGGCAG-----AGGUUAAGUCCAUCGCUGGACACUCGAA-------CUCGAGUU .......((((((((((.......(((((.(((...))).)))))(..((((((((((((----...)))))-----))).))))..)))))))))))(((((..-------..))))). ( -47.90) >DroSim_CAF1 25606 104 + 1 GGACAAUGUCCAGCGAUCUUACGGGGGUUGCUACCAUAGGGACUCGAGCUUGCUUCUGCC----UUCGGCGG-----AGGUUAAGUCCAUCGCUGGACACUCGAA-------CUCGAGUU .......((((((((((.......(((((.(((...))).)))))(..((((((((((((----...)))))-----))).))))..)))))))))))(((((..-------..))))). ( -47.20) >DroEre_CAF1 28579 100 + 1 GGACAAUGUCCAG--AUAUUUCGGGGGUUGCUACU--AGGGACUCGAGCUUGCUUCUGGC----UUCGGCAG-----AGGUUAAGUCCGUCGCUGGGCACUCGAG-------CUCGAGUU ((.((((.(((.(--(....))))).))))))...--...((((((((((((((((((.(----....))))-----))))...(((((....)))))....)))-------)))))))) ( -41.20) >DroYak_CAF1 22000 120 + 1 GGACAAUGUCCAGAGAUAUUUCGGGGGUUGCUACUAUAGGGACUCGAGCUUGCUGCCUGCCCUCGUCGGCAGAGGAGAGGUUAAGUCCAUCGCUGGACACUCGACAAUAAGAGUCGAGUU .......((((((.(((....(...(((....)))...)(((((..(.((..((..(((((......))))).))..)).)..)))))))).))))))(((((((.......))))))). ( -45.60) >consensus GGACAAUGUCCAGAGAUCUUACGGGGGUUGCUACCAUAGGGACUCGAGCUUGCUUCUGCC____UUCGGCAG_____AGGUUAAGUCCAUCGCUGGACACUCGAA_______CUCGAGUU .......((((((.(((....(...(((....)))...)(((((..(.((.....(((((.......))))).....)).)..)))))))).))))))((((((.........)))))). (-32.24 = -32.84 + 0.60)


| Location | 8,101,866 – 8,101,970 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.78 |
| Mean single sequence MFE | -35.32 |
| Consensus MFE | -27.78 |
| Energy contribution | -28.62 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.863116 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
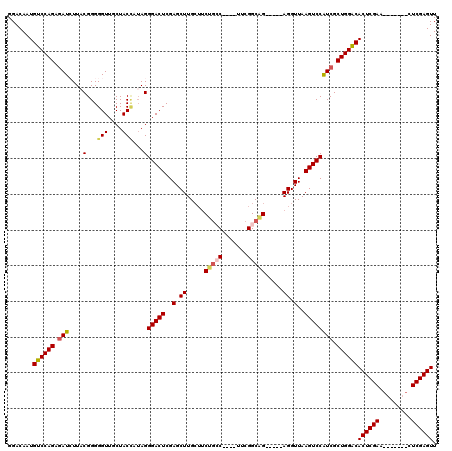
>3R_DroMel_CAF1 8101866 104 - 27905053 AACUCGAG-------UUCGAGUGUCCAGCGAUGGACUUAACCU-----CUGCCGAA----GGCAGACUCAAGCUCGAGUCCCUAUCGUAGCAACCCCCGUAAGAUCUCUGGACAUUGUCC .(((((..-------..)))))((((((.(((((........(-----(((((...----)))))).....((((((.......))).)))..))..(....)))).))))))....... ( -34.20) >DroSec_CAF1 53520 104 - 1 AACUCGAG-------UUCGAGUGUCCAGCGAUGGACUUAACCU-----CUGCCGAA----GGCAGAAGCAAGCUCGAGUCCCUAUGGUAGCAACCCCCGUAAGAUCGCUGGACAUUGUCC .(((((..-------..)))))((((((((((((((((....(-----(((((...----))))))((....)).)))))).(((((.........)))))..))))))))))....... ( -42.40) >DroSim_CAF1 25606 104 - 1 AACUCGAG-------UUCGAGUGUCCAGCGAUGGACUUAACCU-----CCGCCGAA----GGCAGAAGCAAGCUCGAGUCCCUAUGGUAGCAACCCCCGUAAGAUCGCUGGACAUUGUCC .(((((..-------..)))))((((((((((((((((.....-----..(((...----))).((.(....))))))))).(((((.........)))))..))))))))))....... ( -38.00) >DroEre_CAF1 28579 100 - 1 AACUCGAG-------CUCGAGUGCCCAGCGACGGACUUAACCU-----CUGCCGAA----GCCAGAAGCAAGCUCGAGUCCCU--AGUAGCAACCCCCGAAAUAU--CUGGACAUUGUCC .(((((((-------((....(((...((..(((.(.......-----..))))..----)).....))))))))))))....--....((((..((.((....)--).))...)))).. ( -23.50) >DroYak_CAF1 22000 120 - 1 AACUCGACUCUUAUUGUCGAGUGUCCAGCGAUGGACUUAACCUCUCCUCUGCCGACGAGGGCAGGCAGCAAGCUCGAGUCCCUAUAGUAGCAACCCCCGAAAUAUCUCUGGACAUUGUCC .(((((((.......)))))))((((((.(((((((((..........(((((......)))))((.....))..)))))).....((....)).........))).))))))....... ( -38.50) >consensus AACUCGAG_______UUCGAGUGUCCAGCGAUGGACUUAACCU_____CUGCCGAA____GGCAGAAGCAAGCUCGAGUCCCUAUAGUAGCAACCCCCGUAAGAUCGCUGGACAUUGUCC .(((((((.......)))))))((((((.((((((((..(.((.....(((((.......))))).....)).)..))))).....((....)).........))).))))))....... (-27.78 = -28.62 + 0.84)



| Location | 8,101,906 – 8,102,010 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.50 |
| Mean single sequence MFE | -42.74 |
| Consensus MFE | -30.20 |
| Energy contribution | -31.32 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.613191 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8101906 104 + 27905053 GACUCGAGCUUGAGUCUGCC----UUCGGCAG-----AGGUUAAGUCCAUCGCUGGACACUCGAA-------CUCGAGUUCAACCGGCUACCUGGGAUGCUGGUGAUGCAUCGCCAAGGG ((((((((.(((((((((((----...)))))-----)......(((((....))))).))))).-------))))))))..........((((((((((.......))))).)).))). ( -45.20) >DroSec_CAF1 53560 104 + 1 GACUCGAGCUUGCUUCUGCC----UUCGGCAG-----AGGUUAAGUCCAUCGCUGGACACUCGAA-------CUCGAGUUCAAUCGGCUACCUGGGAUGCUGGUGAUGCAUCGCCAAGAA ((((((((...(((((((((----...)))))-----))))...(((((....))))).......-------))))))))..........(.((((((((.......))))).))).).. ( -40.50) >DroSim_CAF1 25646 104 + 1 GACUCGAGCUUGCUUCUGCC----UUCGGCGG-----AGGUUAAGUCCAUCGCUGGACACUCGAA-------CUCGAGUUCAAUCGGCUACCUGGGAUGCUGGUGAUGCCUGGCCAAGGG ((((((((...(((((((((----...)))))-----))))...(((((....))))).......-------))))))))..((((((..(....)..))))))....(((.....))). ( -38.20) >DroEre_CAF1 28615 104 + 1 GACUCGAGCUUGCUUCUGGC----UUCGGCAG-----AGGUUAAGUCCGUCGCUGGGCACUCGAG-------CUCGAGUUCAACCGGCUACCUGGGAUGCUGCUGAUGCAUCGCCAAGAG ((((((((((((((((((.(----....))))-----))))...(((((....)))))....)))-------))))))))...((((....))))(((((.......)))))........ ( -43.40) >DroYak_CAF1 22040 120 + 1 GACUCGAGCUUGCUGCCUGCCCUCGUCGGCAGAGGAGAGGUUAAGUCCAUCGCUGGACACUCGACAAUAAGAGUCGAGUUCAAGCGGCUACCUGGGAUGCUGCUGAUGCAUCGCCAAGAG ((((..(.((..((..(((((......))))).))..)).)..))))..(((((.((.(((((((.......))))))))).)))))...(.((((((((.......))))).))).).. ( -46.40) >consensus GACUCGAGCUUGCUUCUGCC____UUCGGCAG_____AGGUUAAGUCCAUCGCUGGACACUCGAA_______CUCGAGUUCAACCGGCUACCUGGGAUGCUGGUGAUGCAUCGCCAAGAG ((((..(.((.....(((((.......))))).....)).)..))))....(((((..((((((.........))))))....)))))..(.((((((((.......))))).))).).. (-30.20 = -31.32 + 1.12)



| Location | 8,101,906 – 8,102,010 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.50 |
| Mean single sequence MFE | -38.84 |
| Consensus MFE | -28.34 |
| Energy contribution | -28.98 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.536159 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8101906 104 - 27905053 CCCUUGGCGAUGCAUCACCAGCAUCCCAGGUAGCCGGUUGAACUCGAG-------UUCGAGUGUCCAGCGAUGGACUUAACCU-----CUGCCGAA----GGCAGACUCAAGCUCGAGUC ..(((((.(((((.......))))))))))...........(((((((-------((.(((((((((....))))).......-----(((((...----))))))))).))))))))). ( -44.40) >DroSec_CAF1 53560 104 - 1 UUCUUGGCGAUGCAUCACCAGCAUCCCAGGUAGCCGAUUGAACUCGAG-------UUCGAGUGUCCAGCGAUGGACUUAACCU-----CUGCCGAA----GGCAGAAGCAAGCUCGAGUC ..(((((.(((((.......))))))))))...........(((((((-------((.(...(((((....)))))......(-----(((((...----))))))..).))))))))). ( -39.80) >DroSim_CAF1 25646 104 - 1 CCCUUGGCCAGGCAUCACCAGCAUCCCAGGUAGCCGAUUGAACUCGAG-------UUCGAGUGUCCAGCGAUGGACUUAACCU-----CCGCCGAA----GGCAGAAGCAAGCUCGAGUC ..((((((...((.(((((.........))).(((..(((.(((((..-------..)))))(((((....))))).......-----....))).----))).)).))..)).)))).. ( -29.60) >DroEre_CAF1 28615 104 - 1 CUCUUGGCGAUGCAUCAGCAGCAUCCCAGGUAGCCGGUUGAACUCGAG-------CUCGAGUGCCCAGCGACGGACUUAACCU-----CUGCCGAA----GCCAGAAGCAAGCUCGAGUC (((((((.(((((.......)))))))))).))........(((((((-------((....(((...((..(((.(.......-----..))))..----)).....)))))))))))). ( -34.40) >DroYak_CAF1 22040 120 - 1 CUCUUGGCGAUGCAUCAGCAGCAUCCCAGGUAGCCGCUUGAACUCGACUCUUAUUGUCGAGUGUCCAGCGAUGGACUUAACCUCUCCUCUGCCGACGAGGGCAGGCAGCAAGCUCGAGUC (((((((.(((((.......)))))))))).))..(((((.(((((((.......)))))))(((((....)))))....(((.(((((.......))))).)))...)))))....... ( -46.00) >consensus CUCUUGGCGAUGCAUCACCAGCAUCCCAGGUAGCCGAUUGAACUCGAG_______UUCGAGUGUCCAGCGAUGGACUUAACCU_____CUGCCGAA____GGCAGAAGCAAGCUCGAGUC ..(((((.(((((.......))))))))))....((.(((.(((((((.......)))))))...)))))...((((..(.((.....(((((.......))))).....)).)..)))) (-28.34 = -28.98 + 0.64)



| Location | 8,101,937 – 8,102,050 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.13 |
| Mean single sequence MFE | -40.70 |
| Consensus MFE | -37.66 |
| Energy contribution | -37.06 |
| Covariance contribution | -0.60 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.93 |
| SVM decision value | 3.56 |
| SVM RNA-class probability | 0.999387 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8101937 113 + 27905053 UUAAGUCCAUCGCUGGACACUCGAA-------CUCGAGUUCAACCGGCUACCUGGGAUGCUGGUGAUGCAUCGCCAAGGGUUUAUGGGCAUCUGGUCUAGUGAUCUCUUAUUGUUAACGC .((((...(((((((((((((((..-------..)))))....(((...(((((((((((.......))))).))..))))...))).......))))))))))..)))).......... ( -39.20) >DroSec_CAF1 53591 113 + 1 UUAAGUCCAUCGCUGGACACUCGAA-------CUCGAGUUCAAUCGGCUACCUGGGAUGCUGGUGAUGCAUCGCCAAGAAUUUUUGGGCAUCUGGUCCAGUGAUCUCUUAUUGUUAACGC .((((...(((((((((((((((..-------..))))).....(((...((..(((..(((((((....))))).))...)))..))...)))))))))))))..)))).......... ( -39.40) >DroSim_CAF1 25677 113 + 1 UUAAGUCCAUCGCUGGACACUCGAA-------CUCGAGUUCAAUCGGCUACCUGGGAUGCUGGUGAUGCCUGGCCAAGGGUUUUUGGGCAUCUGGUCCAGUGAUCUCUUAUUGUUAACGC .((((...(((((((((((((((..-------..)))))...((((((..(....)..))))))(((((((((((...))))...)))))))..))))))))))..)))).......... ( -42.20) >DroEre_CAF1 28646 112 + 1 UUAAGUCCGUCGCUGGGCACUCGAG-------CUCGAGUUCAACCGGCUACCUGGGAUGCUGCUGAUGCAUCGCCAAGAGUUU-UGGGCAUCUGGUCCAGUGAUCUCUUAUUGUUAACGC .((((...(((((((((((((((..-------..)))))....((((((.(.((((((((.......))))).))).))))..-))).......))))))))))..)))).......... ( -38.90) >DroYak_CAF1 22080 119 + 1 UUAAGUCCAUCGCUGGACACUCGACAAUAAGAGUCGAGUUCAAGCGGCUACCUGGGAUGCUGCUGAUGCAUCGCCAAGAGUUU-UGGGCAUCUGGUCCAGUGAUCUCUUAUUGUUAACGC .((((...(((((((((((((((((.......)))))))...((((((..(....)..))))))(((((....(((((...))-))))))))..))))))))))..)))).......... ( -43.80) >consensus UUAAGUCCAUCGCUGGACACUCGAA_______CUCGAGUUCAACCGGCUACCUGGGAUGCUGGUGAUGCAUCGCCAAGAGUUU_UGGGCAUCUGGUCCAGUGAUCUCUUAUUGUUAACGC .((((...((((((((((((((((.........))))))...((((((..(....)..))))))(((((....(((........))))))))..))))))))))..)))).......... (-37.66 = -37.06 + -0.60)



| Location | 8,102,050 – 8,102,170 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.50 |
| Mean single sequence MFE | -30.70 |
| Consensus MFE | -26.78 |
| Energy contribution | -27.58 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.777334 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8102050 120 - 27905053 AAAAGCGGCCAACUGUAUGCAAAAGAUUAAUGCAAAAUAUUCAAAUGGAAUAUGUACAUAUUUUGUGACCCCGGCCAAGUUUGCCUAAUUGCGUGAGACAGCCAAAGGUCAGAGGCUGCA ....((((((...((((((((.........)))...((((((.....))))))))))).....(.(((((..(((....((..((.....).)..))...)))...))))).))))))). ( -33.40) >DroSec_CAF1 53704 120 - 1 AAAAUCGGCCAACUGUAUGCAAAAGAUUAAUGCAAAAUAUUCAAAUGGAAUAUGUACAUAUCUUGUGACCCCGGCCAAGUUUGCCUAAUUGCGUGAGACAGCCAAAGGUCAGAGGCGGCA .......(((...((((((((.........)))...((((((.....)))))))))))...(((.(((((..(((....((..((.....).)..))...)))...))))).))).))). ( -29.10) >DroSim_CAF1 25790 120 - 1 AAAAGCGGCCAACUGUAUGCAAAAGAUUAAUGCAAAAUAUUCAAAUGGAAUAUGUACAUAUCUUGUGACCCCGGCCAAGUUUGCCUAAUUGCGUGAGACAGCCAAAGGUCAGAGGCGGCA ....((.(((...((((((((.........)))...((((((.....))))))))))).....(.(((((..(((....((..((.....).)..))...)))...))))).)))).)). ( -31.30) >DroEre_CAF1 28758 120 - 1 AAAAGCGGCCAACUGUAUGCAAAAGAUUGAGGCAAAAUAUUCAAAUGGAAUAUGCACAUAUUUUGUGACCCCGGCCAAGUUUGCCUAAUUGCGUGAGACAGCCAAAGGUCAGAGGCGGCA ....((.(((.....(((((((.......(((((((((((((.....)))))).((((.....))))............)))))))..))))))).(((........)))...))).)). ( -33.40) >DroYak_CAF1 22199 120 - 1 AAAAGCGGCCAACUGUAUGCAAAAGAUUAAUGCAAAAUAUUCAAAUGGAAUAUGUACAUAUUUUGUGACCCCGGCCAAGUUUGCCUAAUUGCGUGAGACAGCCAAAGGUCAGAGGCGACA .......(((...((((((((.........)))...((((((.....))))))))))).....(.(((((..(((....((..((.....).)..))...)))...))))).)))).... ( -26.30) >consensus AAAAGCGGCCAACUGUAUGCAAAAGAUUAAUGCAAAAUAUUCAAAUGGAAUAUGUACAUAUUUUGUGACCCCGGCCAAGUUUGCCUAAUUGCGUGAGACAGCCAAAGGUCAGAGGCGGCA ....((.(((...((((((((.........)))...((((((.....))))))))))).....(.(((((..(((....((..((.....).)..))...)))...))))).)))).)). (-26.78 = -27.58 + 0.80)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:52:13 2006