
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 8,093,184 – 8,093,384 |
| Length | 200 |
| Max. P | 0.999901 |

| Location | 8,093,184 – 8,093,304 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.47 |
| Mean single sequence MFE | -33.46 |
| Consensus MFE | -32.96 |
| Energy contribution | -32.92 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.99 |
| SVM decision value | 2.96 |
| SVM RNA-class probability | 0.997902 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8093184 120 + 27905053 CAUAUAUAUGCAAUCUCAUAUGCAUUACUUUGCAUGUCUGGCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUUGAGGGGCAAAAAUGGCAGGGAGGGAGGUUUUUCUAA ...........(((((((((((((......)))))))(((.((..((...(..(((((((((...((((....)))).)))))))))...).))..)).))).....))))))....... ( -33.50) >DroSec_CAF1 44928 118 + 1 CAUA--UAUGCAAUCUCAUAUGCAUUACUUUGCAUGUCUGGCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUUGAGGGGCAGAAAUGGCAGGGAGGGAGGUUUUUCUAA ....--.....(((((((((((((......)))))))(((.((..((...(..(((((((((...((((....)))).)))))))))...).))..)).))).....))))))....... ( -33.90) >DroSim_CAF1 16962 118 + 1 CAUA--UAUGCAAUCUCAUAUGCAUUACUUUGCAUGUCUGGCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUUGAGGGGCAGAAAUGGCAGGGAGGGAGGUUUUUCUAA ....--.....(((((((((((((......)))))))(((.((..((...(..(((((((((...((((....)))).)))))))))...).))..)).))).....))))))....... ( -33.90) >DroEre_CAF1 19992 118 + 1 CAUA--CAUGCAAUCUCAUAUGCAUUACUUUGCAUGUCUGGCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUAGAGGGGCAAAAAUGGCAGGGAGGGAGGUUUUUCUAA ....--...((..((((...(((.....(((((.(.(((((((((((....))))).........((((....))))...)))))).).)))))....)))..))))...))........ ( -32.40) >DroYak_CAF1 13441 118 + 1 CAUA--UAUGCAAUCUCAUAUGCAUUACUUUGCAUGUCUGGCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUUGAGGGACAGAAAUGGCAGGGAGGGAGGUUUUUCUAU ....--.....((((((.((((((......))))))(((.((...........(((((((((...((((....)))).)))))))))...((....))))..)))..))))))....... ( -33.60) >consensus CAUA__UAUGCAAUCUCAUAUGCAUUACUUUGCAUGUCUGGCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUUGAGGGGCAGAAAUGGCAGGGAGGGAGGUUUUUCUAA ...........(((((((((((((......)))))))(((.((..((...(..(((((((((...((((....)))).)))))))))...).))..)).))).....))))))....... (-32.96 = -32.92 + -0.04)


| Location | 8,093,224 – 8,093,344 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.17 |
| Mean single sequence MFE | -41.24 |
| Consensus MFE | -39.54 |
| Energy contribution | -40.14 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.41 |
| Structure conservation index | 0.96 |
| SVM decision value | 4.45 |
| SVM RNA-class probability | 0.999901 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
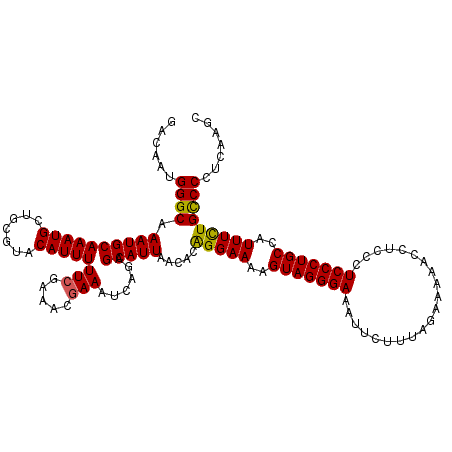
>3R_DroMel_CAF1 8093224 120 + 27905053 GCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUUGAGGGGCAAAAAUGGCAGGGAGGGAGGUUUUUCUAAAGAAUUUCCCUACUUUUCCUGUGUUAAUGCUCUGAUUUCG ..((((((.....(((((((((...((((....)))).)))))))))(((((......(((((((((((((((((.....))))))))))).....)))))).....))))))))))).. ( -41.90) >DroSec_CAF1 44966 120 + 1 GCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUUGAGGGGCAGAAAUGGCAGGGAGGGAGGUUUUUCUAAAGAAUUUCCCUACUUUUCCUGUGUUAAUGCUCUGAUUUCG ..((((((.....(((((((((...((((....)))).)))))))))(((((..((((.((((((((((((((((.....))))))))))).....)))))))))..))))))))))).. ( -43.10) >DroSim_CAF1 17000 120 + 1 GCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUUGAGGGGCAGAAAUGGCAGGGAGGGAGGUUUUUCUAAAGAAUUUCCCUACUUUUCCUGUGUUAAUGCUCUGAUUUCG ..((((((.....(((((((((...((((....)))).)))))))))(((((..((((.((((((((((((((((.....))))))))))).....)))))))))..))))))))))).. ( -43.10) >DroEre_CAF1 20030 120 + 1 GCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUAGAGGGGCAAAAAUGGCAGGGAGGGAGGUUUUUCUAAAGAAUUUCCCUACUUUUCCCGUGUUAAUGCUCUGAUUUCG ..(((((....))))).(((((...((((....)))).)))))...((((((....(((((((((((((((((((.....))))))))))......)))).))))).))))))....... ( -37.50) >DroYak_CAF1 13479 120 + 1 GCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUUGAGGGACAGAAAUGGCAGGGAGGGAGGUUUUUCUAUAGAAUUUCCCUACUUUUCCUGUGUUAAUGCUCUGAUUUCG ..((((.......(((((((((...((((....)))).)))))))))...((....))(((((((((((((((((.....))))))))))).....))))))......))))........ ( -40.60) >consensus GCGAGUUAAGGAACUCAAGCGUAAAGGUCAAAAGAUCAACGCUUGAGGGGCAGAAAUGGCAGGGAGGGAGGUUUUUCUAAAGAAUUUCCCUACUUUUCCUGUGUUAAUGCUCUGAUUUCG ..((((((.....(((((((((...((((....)))).)))))))))(((((..((((.((((((((((((((((.....))))))))))).....)))))))))..))))))))))).. (-39.54 = -40.14 + 0.60)



| Location | 8,093,224 – 8,093,344 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.17 |
| Mean single sequence MFE | -30.20 |
| Consensus MFE | -30.08 |
| Energy contribution | -29.88 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.55 |
| Structure conservation index | 1.00 |
| SVM decision value | 4.18 |
| SVM RNA-class probability | 0.999825 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8093224 120 - 27905053 CGAAAUCAGAGCAUUAACACAGGAAAAGUAGGGAAAUUCUUUAGAAAAACCUCCCUCCCUGCCAUUUUUGCCCCUCAAGCGUUGAUCUUUUGACCUUUACGCUUGAGUUCCUUAACUCGC ........(((........((((((..(((((((.....................)))))))..))))))...(((((((((.(.((....)).)...)))))))))........))).. ( -29.10) >DroSec_CAF1 44966 120 - 1 CGAAAUCAGAGCAUUAACACAGGAAAAGUAGGGAAAUUCUUUAGAAAAACCUCCCUCCCUGCCAUUUCUGCCCCUCAAGCGUUGAUCUUUUGACCUUUACGCUUGAGUUCCUUAACUCGC ........(((........((((((..(((((((.....................)))))))..))))))...(((((((((.(.((....)).)...)))))))))........))).. ( -31.60) >DroSim_CAF1 17000 120 - 1 CGAAAUCAGAGCAUUAACACAGGAAAAGUAGGGAAAUUCUUUAGAAAAACCUCCCUCCCUGCCAUUUCUGCCCCUCAAGCGUUGAUCUUUUGACCUUUACGCUUGAGUUCCUUAACUCGC ........(((........((((((..(((((((.....................)))))))..))))))...(((((((((.(.((....)).)...)))))))))........))).. ( -31.60) >DroEre_CAF1 20030 120 - 1 CGAAAUCAGAGCAUUAACACGGGAAAAGUAGGGAAAUUCUUUAGAAAAACCUCCCUCCCUGCCAUUUUUGCCCCUCUAGCGUUGAUCUUUUGACCUUUACGCUUGAGUUCCUUAACUCGC ........(((.........((((((((((((((.....................)))))))...)))).)))(((.(((((.(.((....)).)...))))).)))........))).. ( -25.90) >DroYak_CAF1 13479 120 - 1 CGAAAUCAGAGCAUUAACACAGGAAAAGUAGGGAAAUUCUAUAGAAAAACCUCCCUCCCUGCCAUUUCUGUCCCUCAAGCGUUGAUCUUUUGACCUUUACGCUUGAGUUCCUUAACUCGC ........(((.......(((((((..(((((((.....................)))))))..)))))))..(((((((((.(.((....)).)...)))))))))........))).. ( -32.80) >consensus CGAAAUCAGAGCAUUAACACAGGAAAAGUAGGGAAAUUCUUUAGAAAAACCUCCCUCCCUGCCAUUUCUGCCCCUCAAGCGUUGAUCUUUUGACCUUUACGCUUGAGUUCCUUAACUCGC ........(((........((((((..(((((((.....................)))))))..))))))...(((((((((.(.((....)).)...)))))))))........))).. (-30.08 = -29.88 + -0.20)



| Location | 8,093,264 – 8,093,384 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.00 |
| Mean single sequence MFE | -37.24 |
| Consensus MFE | -35.62 |
| Energy contribution | -35.54 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.96 |
| SVM decision value | 3.79 |
| SVM RNA-class probability | 0.999615 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8093264 120 + 27905053 GCUUGAGGGGCAAAAAUGGCAGGGAGGGAGGUUUUUCUAAAGAAUUUCCCUACUUUUCCUGUGUUAAUGCUCUGAUUUCGUUUCCAAAAAUGUACGCAGCAUUUGCAUUUGCCCAUUGUC .......(((((((....(((((((((((((((((.....))))))))))).....))))))((.((((((.((....(((((....)))))..)).)))))).)).)))))))...... ( -38.50) >DroSec_CAF1 45006 120 + 1 GCUUGAGGGGCAGAAAUGGCAGGGAGGGAGGUUUUUCUAAAGAAUUUCCCUACUUUUCCUGUGUUAAUGCUCUGAUUUCGUUUCGAAAAAUGUACGCAGCAUUUGCAUUUGCCCAUUGUC .......(((((((....(((((((((((((((((.....))))))))))).....))))))((.((((((.((....(((((....)))))..)).)))))).)).)))))))...... ( -38.60) >DroSim_CAF1 17040 120 + 1 GCUUGAGGGGCAGAAAUGGCAGGGAGGGAGGUUUUUCUAAAGAAUUUCCCUACUUUUCCUGUGUUAAUGCUCUGAUUUCGUUUCGAAAAAUGUACGCAGCAUUUGCAUUUGCCCAUUGUC .......(((((((....(((((((((((((((((.....))))))))))).....))))))((.((((((.((....(((((....)))))..)).)))))).)).)))))))...... ( -38.60) >DroEre_CAF1 20070 120 + 1 GCUAGAGGGGCAAAAAUGGCAGGGAGGGAGGUUUUUCUAAAGAAUUUCCCUACUUUUCCCGUGUUAAUGCUCUGAUUUCGUUUUGAAAAAUGUAUGCAGCAUUUGCAUUUGCCCAUUGUC .......(((((((.((((.(((((((((((((((.....))))))))))).))))..))))((.((((((.......(((((....))))).....)))))).)).)))))))...... ( -36.10) >DroYak_CAF1 13519 120 + 1 GCUUGAGGGACAGAAAUGGCAGGGAGGGAGGUUUUUCUAUAGAAUUUCCCUACUUUUCCUGUGUUAAUGCUCUGAUUUCGUUUCGAAAAAUGUAUGCAGCAUUUGCAUUUGCCCAUUGUC ......(((.((((....(((((((((((((((((.....))))))))))).....))))))((.((((((.......(((((....))))).....)))))).)).)))))))...... ( -34.40) >consensus GCUUGAGGGGCAGAAAUGGCAGGGAGGGAGGUUUUUCUAAAGAAUUUCCCUACUUUUCCUGUGUUAAUGCUCUGAUUUCGUUUCGAAAAAUGUACGCAGCAUUUGCAUUUGCCCAUUGUC .......(((((((....(((((((((((((((((.....))))))))))).....))))))((.((((((.((....(((((....)))))..)).)))))).)).)))))))...... (-35.62 = -35.54 + -0.08)



| Location | 8,093,264 – 8,093,384 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.00 |
| Mean single sequence MFE | -26.68 |
| Consensus MFE | -23.52 |
| Energy contribution | -23.16 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.767337 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 8093264 120 - 27905053 GACAAUGGGCAAAUGCAAAUGCUGCGUACAUUUUUGGAAACGAAAUCAGAGCAUUAACACAGGAAAAGUAGGGAAAUUCUUUAGAAAAACCUCCCUCCCUGCCAUUUUUGCCCCUCAAGC ......(((((((....((((((.(......(((((....)))))...)))))))....((((...((..(((...(((....)))...)))..)).)))).....)))))))....... ( -29.40) >DroSec_CAF1 45006 120 - 1 GACAAUGGGCAAAUGCAAAUGCUGCGUACAUUUUUCGAAACGAAAUCAGAGCAUUAACACAGGAAAAGUAGGGAAAUUCUUUAGAAAAACCUCCCUCCCUGCCAUUUCUGCCCCUCAAGC ......((((.......((((((.(.......(((((...)))))...))))))).....(((((..(((((((.....................)))))))..)))))))))....... ( -26.30) >DroSim_CAF1 17040 120 - 1 GACAAUGGGCAAAUGCAAAUGCUGCGUACAUUUUUCGAAACGAAAUCAGAGCAUUAACACAGGAAAAGUAGGGAAAUUCUUUAGAAAAACCUCCCUCCCUGCCAUUUCUGCCCCUCAAGC ......((((.......((((((.(.......(((((...)))))...))))))).....(((((..(((((((.....................)))))))..)))))))))....... ( -26.30) >DroEre_CAF1 20070 120 - 1 GACAAUGGGCAAAUGCAAAUGCUGCAUACAUUUUUCAAAACGAAAUCAGAGCAUUAACACGGGAAAAGUAGGGAAAUUCUUUAGAAAAACCUCCCUCCCUGCCAUUUUUGCCCCUCUAGC ......(((((((.(((((((((.(.......((((.....))))...))))))).....((((......((((..(((....))).....)))))))))))....)))))))....... ( -25.90) >DroYak_CAF1 13519 120 - 1 GACAAUGGGCAAAUGCAAAUGCUGCAUACAUUUUUCGAAACGAAAUCAGAGCAUUAACACAGGAAAAGUAGGGAAAUUCUAUAGAAAAACCUCCCUCCCUGCCAUUUCUGUCCCUCAAGC ......(((..((((((((((.......)))))((((...))))......)))))...(((((((..(((((((.....................)))))))..))))))))))...... ( -25.50) >consensus GACAAUGGGCAAAUGCAAAUGCUGCGUACAUUUUUCGAAACGAAAUCAGAGCAUUAACACAGGAAAAGUAGGGAAAUUCUUUAGAAAAACCUCCCUCCCUGCCAUUUCUGCCCCUCAAGC ......((((.((((((((((.......)))))(((.....)))......))))).....(((((..(((((((.....................)))))))..)))))))))....... (-23.52 = -23.16 + -0.36)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:51:38 2006