
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,957,315 – 7,957,455 |
| Length | 140 |
| Max. P | 0.894618 |

| Location | 7,957,315 – 7,957,415 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 89.14 |
| Mean single sequence MFE | -25.13 |
| Consensus MFE | -20.11 |
| Energy contribution | -20.33 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.88 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.894618 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7957315 100 + 27905053 -----AAAUGAAAAUUAUCCUUGCGGCAGCAAAAAAAGAUUGUUGUCAAAUCAUAUUGGCUGUGUUCCUUAAACAAUAAGCAGGCAGC-UAGUUGCGAGCACAAAC -----..............(((((((((((((.......)))))))).......(((((((((.(..((((.....)))).).)))))-)))).)))))....... ( -26.30) >DroSec_CAF1 23711 104 + 1 ACAAGAAAUGAAAAUUAUGUUUGCGGCAGCAAAAAAAGAAA--AGUCCAACCAUAUUGGCUGUCUUCCUUAAACAAUAAGCAGGCAGCACAGUUGCGAGCACAAAC .................((((((((((..............--....(((.....)))(((((((..((((.....)))).)))))))...))))))))))..... ( -24.10) >DroSim_CAF1 23959 101 + 1 --AAGAAAUGAAAAUUAUGUUUGCGGCAGCAAAAAAAGAAA--AGUCCAACCAUAUUGGCUGUCUUCCUUAAACAAUAAGCAGGCAGC-UAGUUGCGAGCACAAAC --...............((((((((((..........((..--..)).........(((((((((..((((.....)))).)))))))-))))))))))))..... ( -25.00) >consensus __AAGAAAUGAAAAUUAUGUUUGCGGCAGCAAAAAAAGAAA__AGUCCAACCAUAUUGGCUGUCUUCCUUAAACAAUAAGCAGGCAGC_UAGUUGCGAGCACAAAC .................((((((((((..........((......)).........(((((((((..((((.....)))).))))))).))))))))))))..... (-20.11 = -20.33 + 0.23)


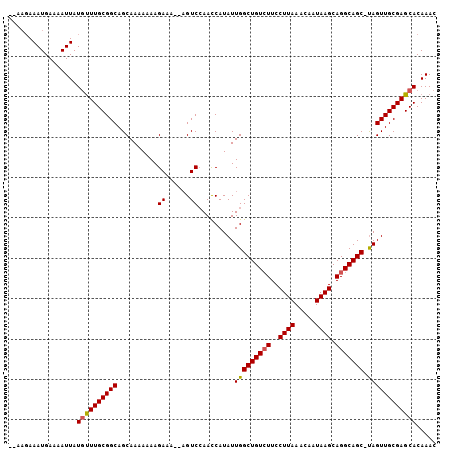
| Location | 7,957,350 – 7,957,455 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 79.84 |
| Mean single sequence MFE | -26.12 |
| Consensus MFE | -15.11 |
| Energy contribution | -15.18 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.813205 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7957350 105 + 27905053 UGUUGUCAAAUCAUAUUGGCUGUGUUCCUUAAACAAUAAGCAGGCAGC-UAGUUGCGAGCA-CAAACGCUAUUUGCCAUGGCAACGAAAAUUGAAUGGGGAAAAAUA .(((((((...((...(((((((((((((((.....))))...((((.-...)))))))))-))...))))..))...)))))))...................... ( -24.70) >DroSec_CAF1 23751 104 + 1 A--AGUCCAACCAUAUUGGCUGUCUUCCUUAAACAAUAAGCAGGCAGCACAGUUGCGAGCA-CAAACGCUUCUUGCCAUGGCAACGAAAAUUGAAAGGGGAAAAAUA .--..(((..((......(((((((..((((.....)))).))))))).((((((((....-....)))(((((((....)))).))))))))...)))))...... ( -26.40) >DroSim_CAF1 23997 103 + 1 A--AGUCCAACCAUAUUGGCUGUCUUCCUUAAACAAUAAGCAGGCAGC-UAGUUGCGAGCA-CAAACGCUUCUUGUCAUGGCAACGAAAAUUGAAAGGGAAAAAAUA .--..(((........(((((((((..((((.....)))).)))))))-))(((((.(..(-(((.......))))..).)))))............)))....... ( -25.70) >DroEre_CAF1 24752 86 + 1 ------------------GUUGUGUUCCUUAGACAAUAAGAGGGCAGC-UUGUUGCAAGCAGCAAACGCCCUUUGCCAUGGCAACUAAAAUUGCAAGGGAA--AAUA ------------------......((((((.(.((((.(((((((.(.-((((((....)))))).))))))))((....)).......))))).))))))--.... ( -27.70) >consensus A__AGUCCAACCAUAUUGGCUGUCUUCCUUAAACAAUAAGCAGGCAGC_UAGUUGCGAGCA_CAAACGCUUCUUGCCAUGGCAACGAAAAUUGAAAGGGAAAAAAUA ........................((((((...((((..((..(((((...)))))..))............((((....)))).....))))..))))))...... (-15.11 = -15.18 + 0.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:49:18 2006