
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,957,046 – 7,957,166 |
| Length | 120 |
| Max. P | 0.726505 |

| Location | 7,957,046 – 7,957,166 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.89 |
| Mean single sequence MFE | -40.11 |
| Consensus MFE | -37.59 |
| Energy contribution | -36.93 |
| Covariance contribution | -0.66 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.663255 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7957046 120 + 27905053 GUCGGGUGUUUUGGCGGGUGACCUUCACAUGCCCGGAUAGCUCAGUCGGUAGAGCAUUGGACUUUUAAUCCAAGGGUCCAGGGUUCAAGUCCCUGUUCGGGCGGCCAUCUUUUUUUGCCU ...(((((...((((((....))......((((((((((((((........))))..(((((((((.....)))))))))(((.......)))))))))))))))))........))))) ( -46.40) >DroSec_CAF1 23450 112 + 1 GUAAUGUUUUUUGGCGGACGACUUUCACCUGCUCGGAUAGCUCAGUCGGUAGAGCAUUGGACUUUUAAUCCAAGGGUCCAGGGUUCAAGUCCCUGUUCGGGCGGUCAUCUUU-------- ((..((((....))))....))......(((((((((((((((........))))..(((((((((.....)))))))))(((.......))))))))))))))........-------- ( -33.80) >DroSim_CAF1 23682 120 + 1 GCAAUGUUUUUUGGCGGACGACUUUCACCUGCCCGGAUAGCUCAGUCGGUAGAGCAUUGGACUUUUAAUCCAAGGGUCCAGGGUUCAAGUCCCUGUUCGGGCGGCCAUCUUUUUUUGAUU .(((.......(((((((.....)))...((((((((((((((........))))..(((((((((.....)))))))))(((.......))))))))))))))))).......)))... ( -40.14) >consensus GUAAUGUUUUUUGGCGGACGACUUUCACCUGCCCGGAUAGCUCAGUCGGUAGAGCAUUGGACUUUUAAUCCAAGGGUCCAGGGUUCAAGUCCCUGUUCGGGCGGCCAUCUUUUUUUG__U ...........(((((((.....)))...((((((((((((((........))))..(((((((((.....)))))))))(((.......)))))))))))))))))............. (-37.59 = -36.93 + -0.66)



| Location | 7,957,046 – 7,957,166 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.89 |
| Mean single sequence MFE | -34.87 |
| Consensus MFE | -30.34 |
| Energy contribution | -30.90 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.726505 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7957046 120 - 27905053 AGGCAAAAAAAGAUGGCCGCCCGAACAGGGACUUGAACCCUGGACCCUUGGAUUAAAAGUCCAAUGCUCUACCGACUGAGCUAUCCGGGCAUGUGAAGGUCACCCGCCAAAACACCCGAC .(((..............(((((..(((((.......))))).....((((((.....)))))).((((........))))....)))))..(((.....)))..)))............ ( -37.00) >DroSec_CAF1 23450 112 - 1 --------AAAGAUGACCGCCCGAACAGGGACUUGAACCCUGGACCCUUGGAUUAAAAGUCCAAUGCUCUACCGACUGAGCUAUCCGAGCAGGUGAAAGUCGUCCGCCAAAAAACAUUAC --------...((((((((((....(((((.......)))))....(((((((............((((........)))).)))))))..))))...))))))................ ( -30.52) >DroSim_CAF1 23682 120 - 1 AAUCAAAAAAAGAUGGCCGCCCGAACAGGGACUUGAACCCUGGACCCUUGGAUUAAAAGUCCAAUGCUCUACCGACUGAGCUAUCCGGGCAGGUGAAAGUCGUCCGCCAAAAAACAUUGC .............((((.(((((..(((((.......))))).....((((((.....)))))).((((........))))....))))).((((.....)).))))))........... ( -37.10) >consensus A__CAAAAAAAGAUGGCCGCCCGAACAGGGACUUGAACCCUGGACCCUUGGAUUAAAAGUCCAAUGCUCUACCGACUGAGCUAUCCGGGCAGGUGAAAGUCGUCCGCCAAAAAACAUUAC ...........((((((..(((.....)))(((((...((((((...((((((.....)))))).((((........))))..)))))))))))....))))))................ (-30.34 = -30.90 + 0.56)

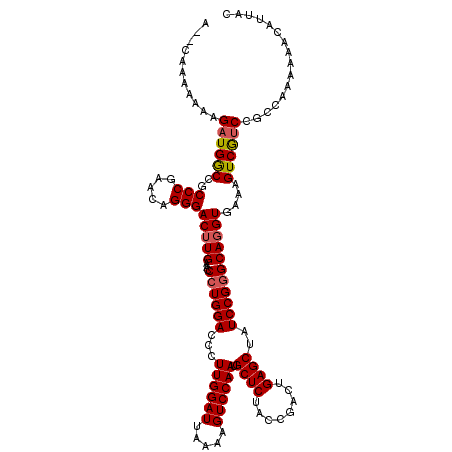

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:49:16 2006