
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,931,797 – 7,931,912 |
| Length | 115 |
| Max. P | 0.772958 |

| Location | 7,931,797 – 7,931,912 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.67 |
| Mean single sequence MFE | -30.51 |
| Consensus MFE | -17.50 |
| Energy contribution | -19.75 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.04 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.756486 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
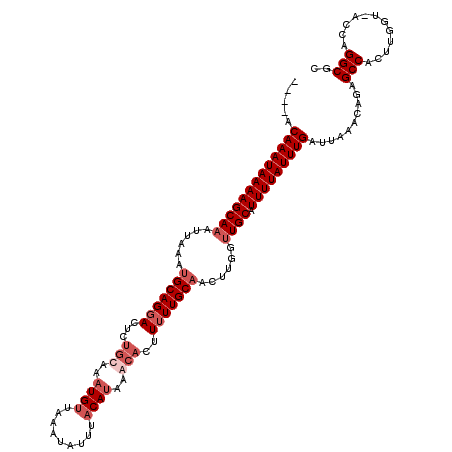
>3R_DroMel_CAF1 7931797 115 + 27905053 ----ACAAAUAAAAGCAAAUUAAAUGCAGGACUCUGCAAAUGUUAAAUAUUUACAUAAGCACUUUUUGCAACUUGGUUGCAUUUUAUUUGAUUAACCAGGGCCACUUGGU-ACCUGGCGC ----.(((((((((((((......(((((((...(((..((((.........))))..)))..)))))))......)))).))))))))).....((((((((....)))-.)))))... ( -33.30) >DroSim_CAF1 65448 116 + 1 ----ACAAAUAAAAGCAAAUUAAAUGCAGGACAUUGCAAAUGGUAAAUAUUUGCAUAAGCACUUUUUGCAACUUGGUUGCAUUUUAUUUGAUUAACCAGAGCCACUUGGUUAGCUGGCGC ----.(((((((((((((......(((((((...((((((((.....))))))))........)))))))......)))).))))))))).....((((.(((....)))...))))... ( -31.40) >DroEre_CAF1 64298 115 + 1 ----GCAAAUAAAAGCAAAUUUAAUGCAGUACUCCACAAAUGUUGCAUGUUUACACUAACACUUUUUGCAACUCGGCUGCAUUUUAUUUGAUUAAACAUAGCCACUUGGU-GCCAGGCGC ----((.........(((((..((((((((...........((((((((((......)))).....))))))...))))))))..)))))..........(((...((..-..))))))) ( -26.04) >DroYak_CAF1 65712 119 + 1 AAAUACAAAUAAAAGCAAAUUUAAUGCAGGACUCUGCAAAUGUUGUAUAUUUACAUAAACACUUUUUGCCACUUGGUUGCAUUUUAUUUGAUUAAACGCAGCCACUUGGU-GGCAGGCGC ..............((........(((((....)))))..(((((((....))))...)))....(((((((((((((((..((((......)))).)))))))...)))-))))))).. ( -31.30) >consensus ____ACAAAUAAAAGCAAAUUAAAUGCAGGACUCUGCAAAUGUUAAAUAUUUACAUAAACACUUUUUGCAACUUGGUUGCAUUUUAUUUGAUUAAACAGAGCCACUUGGU_ACCAGGCGC .....(((((((((((((......(((((((...(((..((((.........))))..)))..)))))))......)))).)))))))))..........(((............))).. (-17.50 = -19.75 + 2.25)


| Location | 7,931,797 – 7,931,912 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.67 |
| Mean single sequence MFE | -28.30 |
| Consensus MFE | -18.95 |
| Energy contribution | -18.45 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.772958 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
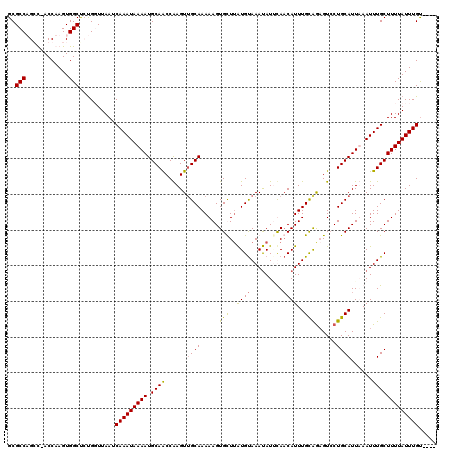
>3R_DroMel_CAF1 7931797 115 - 27905053 GCGCCAGGU-ACCAAGUGGCCCUGGUUAAUCAAAUAAAAUGCAACCAAGUUGCAAAAAGUGCUUAUGUAAAUAUUUAACAUUUGCAGAGUCCUGCAUUUAAUUUGCUUUUAUUUGU---- ..((((((.-.((....)).))))))....(((((((((((((((...))))))...((..(((.(((((((.......))))))))))..))(((.......)))))))))))).---- ( -30.20) >DroSim_CAF1 65448 116 - 1 GCGCCAGCUAACCAAGUGGCUCUGGUUAAUCAAAUAAAAUGCAACCAAGUUGCAAAAAGUGCUUAUGCAAAUAUUUACCAUUUGCAAUGUCCUGCAUUUAAUUUGCUUUUAUUUGU---- ..(((((....((....))..)))))....(((((.(((((((.....((((((((..(((..(((....)))..)))..))))))))....))))))).)))))...........---- ( -27.30) >DroEre_CAF1 64298 115 - 1 GCGCCUGGC-ACCAAGUGGCUAUGUUUAAUCAAAUAAAAUGCAGCCGAGUUGCAAAAAGUGUUAGUGUAAACAUGCAACAUUUGUGGAGUACUGCAUUAAAUUUGCUUUUAUUUGC---- .(((.((..-..)).)))((...((..((.(((((..(((((((.....(..(((...(((((.(((....)))..))))))))..)....)))))))..))))).))..))..))---- ( -24.00) >DroYak_CAF1 65712 119 - 1 GCGCCUGCC-ACCAAGUGGCUGCGUUUAAUCAAAUAAAAUGCAACCAAGUGGCAAAAAGUGUUUAUGUAAAUAUACAACAUUUGCAGAGUCCUGCAUUAAAUUUGCUUUUAUUUGUAUUU ((((..(((-((...))))).))))...........((((((((......(((((((((((((((((....)))).)))))))((((....))))......)))))).....)))))))) ( -31.70) >consensus GCGCCAGCC_ACCAAGUGGCUCUGGUUAAUCAAAUAAAAUGCAACCAAGUUGCAAAAAGUGCUUAUGUAAAUAUUCAACAUUUGCAGAGUCCUGCAUUAAAUUUGCUUUUAUUUGU____ ..(((............)))..........(((((((((.((((......((((.....(((..((((.........))))..)))......))))......)))))))))))))..... (-18.95 = -18.45 + -0.50)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:49:12 2006