
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,930,988 – 7,931,182 |
| Length | 194 |
| Max. P | 0.824860 |

| Location | 7,930,988 – 7,931,082 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 88.30 |
| Mean single sequence MFE | -20.28 |
| Consensus MFE | -14.16 |
| Energy contribution | -15.08 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.70 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.528807 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7930988 94 - 27905053 CUGUUUUCCAGGGCUAAGAUAUCAGUGGUUUGAUAAAAUAGAAUUUGCAUUUUUAAAUUAUUUCAUUUCACGUGCUUGUCAGAGCCUUGAUUUA ........(((((((..((((.((((((..((((((..(((((.......)))))..)))..)))..)))).))..))))..)))))))..... ( -21.70) >DroSec_CAF1 62562 94 - 1 CUGUUUUCCAGGGCUAAGAUAUCAGUGGUUUGAUAAAAUAGAAUUUGCAUUUUUAAAUUAUUUCAUUUCACGUGCUUGUCAGAGCCUUGAUUUA ........(((((((..((((.((((((..((((((..(((((.......)))))..)))..)))..)))).))..))))..)))))))..... ( -21.70) >DroSim_CAF1 64654 79 - 1 CUGUUUUCCGGG--UAAAAUA--AGUGGUUG----AAAAAAAAAUUGCCUU-UUAAAUUU--UUCUUUCCCG-GCUUGUCA-AGCCUUG--UUA .........(((--(...(((--(((.(..(----(((.(((((((.....-...)))))--)).)))).).-))))))..-.))))..--... ( -15.00) >DroEre_CAF1 63463 94 - 1 CUGUUUUCCAGGGCUAAGAUAUCAGUGGGUUGAUAAAAUAGAAUUUGCAUUUUUAAAUUAUUUCAUUUCACGUGCUUGUCAGAGCCUUGAUUUA ........(((((((..((((.(((((((..(((((..(((((.......)))))..)))))....))))).))..))))..)))))))..... ( -21.50) >DroYak_CAF1 64842 94 - 1 CUGUUUUCCAGGGCUAAGAUAUCAGUGGGUUGAUAAAAUAGAAUUUGCAUUUUUAAAUUAUUUCAUUUCACGUGCUUGUCAGAGCCUUGAUUUA ........(((((((..((((.(((((((..(((((..(((((.......)))))..)))))....))))).))..))))..)))))))..... ( -21.50) >consensus CUGUUUUCCAGGGCUAAGAUAUCAGUGGUUUGAUAAAAUAGAAUUUGCAUUUUUAAAUUAUUUCAUUUCACGUGCUUGUCAGAGCCUUGAUUUA ........(((((((..((((...((((..(((........((((((......))))))...)))..)))).....))))..)))))))..... (-14.16 = -15.08 + 0.92)


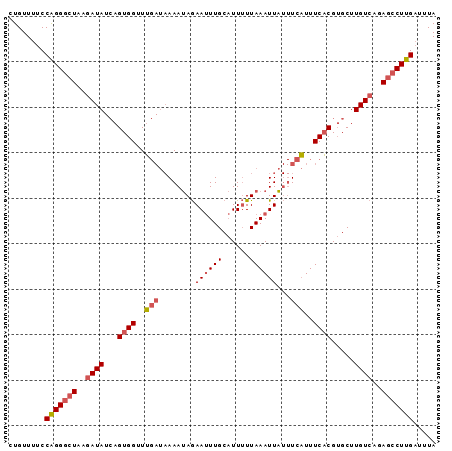
| Location | 7,931,042 – 7,931,153 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 84.83 |
| Mean single sequence MFE | -30.03 |
| Consensus MFE | -20.72 |
| Energy contribution | -22.21 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.824860 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7931042 111 + 27905053 UAUUUUAUCAAACCACUGAUAUCUUAGCCCUGGAAAACAGUACCAAGUCCCAGUCCUUUUAUCUCGGCCCACCUUUGUUGGCCAUUGAACUGGCUGGCAUUGGUGGUUGGU ...........((((((((.((...((..((((...((........)).))))..))...)).)))).(((((..(((..((((......))))..)))..))))).)))) ( -34.30) >DroSec_CAF1 62616 111 + 1 UAUUUUAUCAAACCACUGAUAUCUUAGCCCUGGAAAACAGUACCAACUCCCAGUCCUUUCAUCUCGGCCCACCUUUGUUGGCCAUUGAACUGGCUGCCAUUGGUGGUUGGU ......(((((.((((..((......(..(((.....)))..).......(((((..((((....((((.((....)).))))..))))..)))))..))..))))))))) ( -29.90) >DroSim_CAF1 64701 100 + 1 UUUUU----CAACCACU--UAUUUUA--CCCGGAAAACAGUACC-ACUCCCAG-CCUUUCAUUUGGGCCCACCUUUGUUGGCCAU-GGACUGGCUCCCAUUGGUGGUGGGU .....----........--......(--((((............-.......(-(((.......))))(((((..((..(((((.-....)))))..))..)))))))))) ( -29.30) >DroEre_CAF1 63517 108 + 1 UAUUUUAUCAACCCACUGAUAUCUUAGCCCUGGAAAACAGUACCAAAUCCCAGUCCGUGCAACUCGGGCCCCCAUUGUUGGCCAUUGAACUGGCUGCCAUUGGUGGUU--- .....(((((......))))).....((((((.....))((((.............)))).....))))(((((.((..(((((......)))))..)).))).))..--- ( -26.62) >consensus UAUUUUAUCAAACCACUGAUAUCUUAGCCCUGGAAAACAGUACCAACUCCCAGUCCUUUCAUCUCGGCCCACCUUUGUUGGCCAUUGAACUGGCUGCCAUUGGUGGUUGGU ...............((((......((..((((................))))..))......)))).(((((..((..(((((......)))))..))..)))))..... (-20.72 = -22.21 + 1.50)


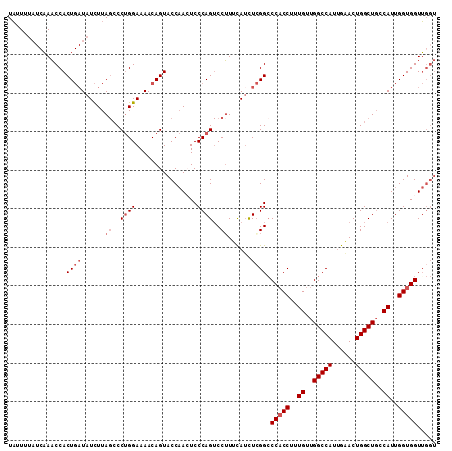
| Location | 7,931,082 – 7,931,182 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 85.67 |
| Mean single sequence MFE | -30.30 |
| Consensus MFE | -19.64 |
| Energy contribution | -19.95 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.619063 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7931082 100 + 27905053 UACCAAGUCCCAGUCCUUUUAUCUCGGCCCACCUUUGUUGGCCAUUGAACUGGCUGGCAUUGGUGGUUGGU----UCACCGCAAAUGAGUUCGUUUUCCUCUCA .((((((.((.((.........)).)))(((((..(((..((((......))))..)))..))))))))))----..........((((...........)))) ( -29.00) >DroSec_CAF1 62656 104 + 1 UACCAACUCCCAGUCCUUUCAUCUCGGCCCACCUUUGUUGGCCAUUGAACUGGCUGCCAUUGGUGGUUGGUUGGUUCACCGCAAAUGAGUUCGUUUUCCUCUCA .(((((((((.((.........)).)).(((((..((..(((((......)))))..))..)))))..)))))))..........((((...........)))) ( -28.20) >DroSim_CAF1 64733 98 + 1 UACC-ACUCCCAG-CCUUUCAUUUGGGCCCACCUUUGUUGGCCAU-GGACUGGCUCCCAUUGGUGGUGGGUG--GUCACCGCAAAUGAGUUCGUU-UCCUCUCA .(((-((((...(-(((.......))))(((((..((..(((((.-....)))))..))..))))).)))))--))....(.(((((....))))-).)..... ( -34.40) >DroEre_CAF1 63557 100 + 1 UACCAAAUCCCAGUCCGUGCAACUCGGGCCCCCAUUGUUGGCCAUUGAACUGGCUGCCAUUGGUGGUU----GGUUCGCCGCAAAUGAGUUCAUUUUCCUCUCA ................(((.(((((((....)).((((.(((....((((..((..((...))..)).----.)))))))))))..)))))))).......... ( -29.60) >consensus UACCAACUCCCAGUCCUUUCAUCUCGGCCCACCUUUGUUGGCCAUUGAACUGGCUGCCAUUGGUGGUUGGU___UUCACCGCAAAUGAGUUCGUUUUCCUCUCA .........................((.(((((..((..(((((......)))))..))..)))))................(((((....))))).))..... (-19.64 = -19.95 + 0.31)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:49:06 2006