
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,846,006 – 7,846,096 |
| Length | 90 |
| Max. P | 0.983271 |

| Location | 7,846,006 – 7,846,096 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.73 |
| Mean single sequence MFE | -31.32 |
| Consensus MFE | -16.35 |
| Energy contribution | -16.55 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.87 |
| Structure conservation index | 0.52 |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.983271 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
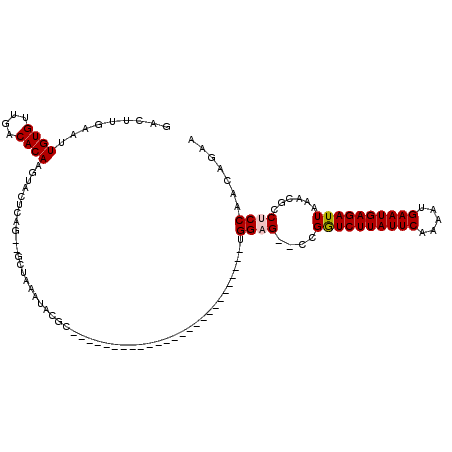
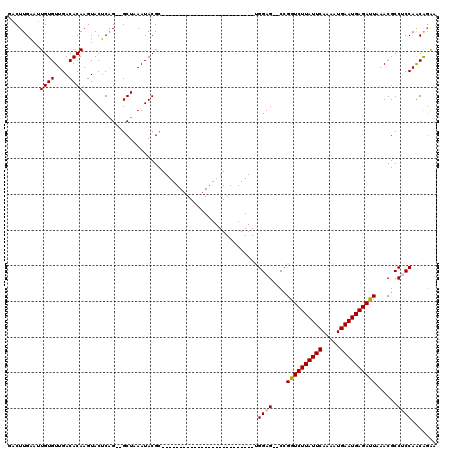
>3R_DroMel_CAF1 7846006 90 + 27905053 GUCUUGGAAUGUGUUGACACAAGUACUCAG--GCUAAAUACGC--------------------------UGGAG--CCGGUCUUAUUCAAAAUGAAUGAGAUUAAACGCCUCCAACGGAA ((((.((..((((....))))....)).))--))......((.--------------------------(((((--.(((((((((((.....)))))))))....)).))))).))... ( -26.40) >DroPse_CAF1 9928 118 + 1 GACUUGAAUUGUGUUGACACAAGUACUCGG--GCUAAAUACGUAUGUACUCGAAGGUAUGUGUGUACGCCGGAGCGCUGGUCUUAUUCAAAAUGAAUGAGAUUAGACGCCACCAGCAGAC ..(((((.(((((....)))))....))))--)........(((..(((.(....)...)))..)))((.((.(((((((((((((((.....)))))))))))).)))..)).)).... ( -39.30) >DroEre_CAF1 9209 90 + 1 AUCUUGAAGUGUGUUGACACAAGUACUCAG--GCUAAAUACGC--------------------------UGGAG--CCGGUCUUAUUCAAAAUGAAUGAGAUUAAACGCCUCCAACGUAA ..(((((..((((....)))).....))))--).....((((.--------------------------(((((--.(((((((((((.....)))))))))....)).))))).)))). ( -28.40) >DroYak_CAF1 9312 90 + 1 GUCUUGAAAUGUGUUGACACAAGUACUCAG--GCUAAAUACGC--------------------------UGGAG--CCGGUCUUAUUCAAAAUGAAUGAGAUUAAACGCCUCCAACGGAA ((((.((..((((....)))).....))))--))......((.--------------------------(((((--.(((((((((((.....)))))))))....)).))))).))... ( -25.30) >DroAna_CAF1 9918 92 + 1 GACUUGAAUUGUGUUGACACAAGUACUCAGGGGCUAAAUACGU--------------------------AGGAG--CUGAUCUUAUUCAAAAUGAAUGAGAUUAGACGCCUCCAACAGGC ..(((((.(((((....)))))....)))))..........((--------------------------.((((--((((((((((((.....))))))))))))....)))).)).... ( -29.20) >DroPer_CAF1 9961 118 + 1 GACUUGAAUUGUGUUGACACAAGUACUCGG--GCUAAAUACGUAUGUACUCGAAGGUAUGUGUGUACGCCGGAGCGCUGGUCUUAUUCAAAAUGAAUGAGAUUAGACGCCACCAGCAGAC ..(((((.(((((....)))))....))))--)........(((..(((.(....)...)))..)))((.((.(((((((((((((((.....)))))))))))).)))..)).)).... ( -39.30) >consensus GACUUGAAUUGUGUUGACACAAGUACUCAG__GCUAAAUACGC__________________________UGGAG__CCGGUCUUAUUCAAAAUGAAUGAGAUUAAACGCCUCCAACAGAA .........((((....)))).................................................((((....((((((((((.....))))))))))......))))....... (-16.35 = -16.55 + 0.20)

| Location | 7,846,006 – 7,846,096 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.73 |
| Mean single sequence MFE | -26.25 |
| Consensus MFE | -14.78 |
| Energy contribution | -14.95 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.853326 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7846006 90 - 27905053 UUCCGUUGGAGGCGUUUAAUCUCAUUCAUUUUGAAUAAGACCGG--CUCCA--------------------------GCGUAUUUAGC--CUGAGUACUUGUGUCAACACAUUCCAAGAC ....(((((((.((.....(((.((((.....)))).))).)).--)))))--------------------------))((((((...--..)))))).((((....))))......... ( -24.40) >DroPse_CAF1 9928 118 - 1 GUCUGCUGGUGGCGUCUAAUCUCAUUCAUUUUGAAUAAGACCAGCGCUCCGGCGUACACACAUACCUUCGAGUACAUACGUAUUUAGC--CCGAGUACUUGUGUCAACACAAUUCAAGUC ((.((((((.((((.((..(((.((((.....)))).)))..)))))))))))).))........(((.((((......((((((...--..)))))).((((....))))))))))).. ( -30.70) >DroEre_CAF1 9209 90 - 1 UUACGUUGGAGGCGUUUAAUCUCAUUCAUUUUGAAUAAGACCGG--CUCCA--------------------------GCGUAUUUAGC--CUGAGUACUUGUGUCAACACACUUCAAGAU .((((((((((.((.....(((.((((.....)))).))).)).--)))))--------------------------)))))......--.((((....((((....)))).)))).... ( -25.90) >DroYak_CAF1 9312 90 - 1 UUCCGUUGGAGGCGUUUAAUCUCAUUCAUUUUGAAUAAGACCGG--CUCCA--------------------------GCGUAUUUAGC--CUGAGUACUUGUGUCAACACAUUUCAAGAC ....(((((((.((.....(((.((((.....)))).))).)).--)))))--------------------------))((((((...--..)))))).((((....))))......... ( -24.40) >DroAna_CAF1 9918 92 - 1 GCCUGUUGGAGGCGUCUAAUCUCAUUCAUUUUGAAUAAGAUCAG--CUCCU--------------------------ACGUAUUUAGCCCCUGAGUACUUGUGUCAACACAAUUCAAGUC ....((.((((....((.((((.((((.....)))).)))).))--)))).--------------------------))((((((((...))))))))(((((....)))))........ ( -21.40) >DroPer_CAF1 9961 118 - 1 GUCUGCUGGUGGCGUCUAAUCUCAUUCAUUUUGAAUAAGACCAGCGCUCCGGCGUACACACAUACCUUCGAGUACAUACGUAUUUAGC--CCGAGUACUUGUGUCAACACAAUUCAAGUC ((.((((((.((((.((..(((.((((.....)))).)))..)))))))))))).))........(((.((((......((((((...--..)))))).((((....))))))))))).. ( -30.70) >consensus GUCCGUUGGAGGCGUCUAAUCUCAUUCAUUUUGAAUAAGACCAG__CUCCA__________________________ACGUAUUUAGC__CUGAGUACUUGUGUCAACACAAUUCAAGAC .......((((....((..(((.((((.....)))).)))..))..)))).............................((((((((...)))))))).((((....))))......... (-14.78 = -14.95 + 0.17)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:47:53 2006