
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,434,131 – 7,434,225 |
| Length | 94 |
| Max. P | 0.860477 |

| Location | 7,434,131 – 7,434,225 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 93.02 |
| Mean single sequence MFE | -28.38 |
| Consensus MFE | -24.46 |
| Energy contribution | -24.71 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.860477 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
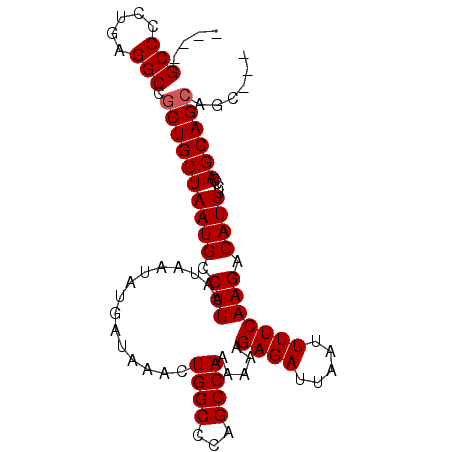
>3R_DroMel_CAF1 7434131 94 + 27905053 CCUACUGCUGCUGGGAAAUGUCUUGAAAAUUAAUCUCUUUUUUGGCUGGGCCAGUUUAUCAUAUUAUUAAGGCAUUAGCAGCGGCCUCAGGGGC---- (((...(((((((...((((((((((.(((...........(((((...)))))........))).))))))))))..)))))))...)))...---- ( -28.21) >DroSec_CAF1 16931 94 + 1 CCUACUGCUGCUGGGAAAUGUCUUGAAAAUUAAUCUCUUUUUUGGCUGGGCCAGUUCAUCAUAUUAUUAAGGCAUUAGCAGCGGCCUCAGGGGC---- (((...(((((((...((((((((((.(((...........(((((...)))))........))).))))))))))..)))))))...)))...---- ( -28.21) >DroEre_CAF1 16920 88 + 1 ---G---CUGCUGGGAAAUGUCUUGAAAAUUAAUCUCUUUUUUGGCUGGGCCAGUUUAUCAUAUUAUUAAGGCAUUAGCAGCGGCCUCAGGGGC---- ---(---((((((....(((((((((.(((...........(((((...)))))........))).)))))))))))))))).(((.....)))---- ( -25.91) >DroYak_CAF1 17532 95 + 1 ---GCUGCUGCUGGGAAAUGUCUUGAAAAUUAAUCUCUUUUUUGGCUGGGCCAGUUUAUCAUAUUAUUAAGGCAUUAGCAGCGGCCUCAGGGGCCACC ---...(((((((....(((((((((.(((...........(((((...)))))........))).))))))))))))))))((((.....))))... ( -31.21) >consensus ___ACUGCUGCUGGGAAAUGUCUUGAAAAUUAAUCUCUUUUUUGGCUGGGCCAGUUUAUCAUAUUAUUAAGGCAUUAGCAGCGGCCUCAGGGGC____ ......(((((((....(((((((((.(((...........(((((...)))))........))).)))))))))))))))).(((.....))).... (-24.46 = -24.71 + 0.25)



| Location | 7,434,131 – 7,434,225 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 93.02 |
| Mean single sequence MFE | -22.73 |
| Consensus MFE | -19.48 |
| Energy contribution | -19.72 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.654999 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7434131 94 - 27905053 ----GCCCCUGAGGCCGCUGCUAAUGCCUUAAUAAUAUGAUAAACUGGCCCAGCCAAAAAAGAGAUUAAUUUUCAAGACAUUUCCCAGCAGCAGUAGG ----...((((..((.((((..((((.(((...............((((...)))).....((((.....))))))).))))...)))).))..)))) ( -23.00) >DroSec_CAF1 16931 94 - 1 ----GCCCCUGAGGCCGCUGCUAAUGCCUUAAUAAUAUGAUGAACUGGCCCAGCCAAAAAAGAGAUUAAUUUUCAAGACAUUUCCCAGCAGCAGUAGG ----...((((..((.((((..((((.(((...............((((...)))).....((((.....))))))).))))...)))).))..)))) ( -23.00) >DroEre_CAF1 16920 88 - 1 ----GCCCCUGAGGCCGCUGCUAAUGCCUUAAUAAUAUGAUAAACUGGCCCAGCCAAAAAAGAGAUUAAUUUUCAAGACAUUUCCCAGCAG---C--- ----(((.....))).((((((((((.(((...............((((...)))).....((((.....))))))).))))....)))))---)--- ( -19.50) >DroYak_CAF1 17532 95 - 1 GGUGGCCCCUGAGGCCGCUGCUAAUGCCUUAAUAAUAUGAUAAACUGGCCCAGCCAAAAAAGAGAUUAAUUUUCAAGACAUUUCCCAGCAGCAGC--- ...((((.....))))((((((((((.(((...............((((...)))).....((((.....))))))).))))....))))))...--- ( -25.40) >consensus ____GCCCCUGAGGCCGCUGCUAAUGCCUUAAUAAUAUGAUAAACUGGCCCAGCCAAAAAAGAGAUUAAUUUUCAAGACAUUUCCCAGCAGCAGC___ ....(((.....))).((((((((((.(((...............((((...)))).....((((.....))))))).))))....))))))...... (-19.48 = -19.72 + 0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:44:32 2006