
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,323,571 – 7,323,984 |
| Length | 413 |
| Max. P | 0.998441 |

| Location | 7,323,571 – 7,323,691 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.35 |
| Mean single sequence MFE | -35.39 |
| Consensus MFE | -35.19 |
| Energy contribution | -35.45 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724015 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7323571 120 - 27905053 CCGACAAAUUUUCCAACAUGAUGUAUGUUUUCGGGGCGUGUGCGUGUUUAUAUCGCGAAUACGUACCGCUGACUACACACACUGGCACUGCGACUCGAAUUAUCGGUGUCAAUUUUUAUG ................(((((((((.(((..(((..(((((.((((.......)))).)))))..)))..))))))).....((((((((.............))))))))....))))) ( -31.72) >DroSim_CAF1 3658 120 - 1 CCGACAAAUUUUCCAACAUGAUGUAUGUUUUCGGGGCGUGUGCGUGUUUAUAUCGCGUAUACGUACCGCUGACUACACACGCUGGCACUGCGACUCGAAUUAUCGGUGUCAAUUUUUAUG ............(((.(.((.((((.(((..(((..((((((((((.......))))))))))..)))..))))))))).).)))......(((((((....)))).))).......... ( -36.40) >DroEre_CAF1 3652 118 - 1 CCGACAAAUUUUCCGACAUGAUGUAUGUUUUCGGGGCGUGUGCGUGUUUAUAUCGCGUAUACGUACCGCUGACUACACA--CUGGCACUGCGACUCGAAUUAUCGGUGUCAAUUUUUAUG ................(((((((((.(((..(((..((((((((((.......))))))))))..)))..)))))))..--.((((((((.............))))))))....))))) ( -36.32) >DroYak_CAF1 3761 118 - 1 CCGACAAAUUUUUCGACAUGAUGUAUGUUUUCGGGGCGUGUGCGUGUUUAUAUCGCGUAUACGUACCGCUGACUACACA--CUGGCACUGCGACUCGAAUUAUCGGUGUCAAUUUUUAUG .(((........))).(((((((((.(((..(((..((((((((((.......))))))))))..)))..)))))))..--.((((((((.............))))))))....))))) ( -37.12) >consensus CCGACAAAUUUUCCAACAUGAUGUAUGUUUUCGGGGCGUGUGCGUGUUUAUAUCGCGUAUACGUACCGCUGACUACACA__CUGGCACUGCGACUCGAAUUAUCGGUGUCAAUUUUUAUG ................(((((((((.(((..(((..((((((((((.......))))))))))..)))..))))))).....((((((((.............))))))))....))))) (-35.19 = -35.45 + 0.25)



| Location | 7,323,611 – 7,323,723 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.77 |
| Mean single sequence MFE | -26.62 |
| Consensus MFE | -25.98 |
| Energy contribution | -26.22 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.96 |
| Structure conservation index | 0.98 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.541960 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7323611 112 - 27905053 CAUUCCCACUCCCUUCCC---UC-----CACCAUGCCAGACCGACAAAUUUUCCAACAUGAUGUAUGUUUUCGGGGCGUGUGCGUGUUUAUAUCGCGAAUACGUACCGCUGACUACACAC ..................---..-----...((((...((...........))...)))).((((.(((..(((..(((((.((((.......)))).)))))..)))..)))))))... ( -22.70) >DroSim_CAF1 3698 115 - 1 CAUUCCCACUCCCACCCCUCCUC-----CACCAUGCCAGACCGACAAAUUUUCCAACAUGAUGUAUGUUUUCGGGGCGUGUGCGUGUUUAUAUCGCGUAUACGUACCGCUGACUACACAC .......................-----...((((...((...........))...)))).((((.(((..(((..((((((((((.......))))))))))..)))..)))))))... ( -27.30) >DroEre_CAF1 3691 105 - 1 CA------C--------CCCACCCUGUGCGCCAUGCUAGACCGACAAAUUUUCCGACAUGAUGUAUGUUUUCGGGGCGUGUGCGUGUUUAUAUCGCGUAUACGUACCGCUGACUACACA- ..------.--------........(((((.((((...((...........))...)))).)))))(((..(((..((((((((((.......))))))))))..)))..)))......- ( -28.50) >DroYak_CAF1 3800 100 - 1 CA------C--------CUCCGC-----CGCCAUGCCAGACCGACAAAUUUUUCGACAUGAUGUAUGUUUUCGGGGCGUGUGCGUGUUUAUAUCGCGUAUACGUACCGCUGACUACACA- ..------.--------......-----...((((...((............))..)))).((((.(((..(((..((((((((((.......))))))))))..)))..)))))))..- ( -28.00) >consensus CA______C________CUCCUC_____CACCAUGCCAGACCGACAAAUUUUCCAACAUGAUGUAUGUUUUCGGGGCGUGUGCGUGUUUAUAUCGCGUAUACGUACCGCUGACUACACA_ ...............................((((.....................)))).((((.(((..(((..((((((((((.......))))))))))..)))..)))))))... (-25.98 = -26.22 + 0.25)



| Location | 7,323,723 – 7,323,843 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.83 |
| Mean single sequence MFE | -36.54 |
| Consensus MFE | -30.22 |
| Energy contribution | -30.90 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.752852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7323723 120 - 27905053 GAUGGGGGGCGACUGUCGUGUGUAAAAUUGAUAACUUUGUUUGUGUAAACAGCGACGCGACAUAUCGCCCUCCAUGGGAGUUUCCUCUCCAUUCCGCAUCCUGUGCCCCUCCACCUUUAC .(((((((((((.((((((((((......((((....))))(((....))))).))))))))..))))))))))).((((......)))).....((((...)))).............. ( -39.60) >DroSec_CAF1 56 120 - 1 GAUGGGGGGCGACUGUCGUGUGUAAAAUUGAUAACUUUGUUUGUGUAAACAGCGACGCGACAUAUCGCCCUCCAUGGGAGUUUCCUCUCCAUUCCGCAUCCUGUGCCCCUCCACCUUUAC .(((((((((((.((((((((((......((((....))))(((....))))).))))))))..))))))))))).((((......)))).....((((...)))).............. ( -39.60) >DroSim_CAF1 3813 120 - 1 AAUGGGGGGCGACUGUCGUGUGUAAAAUUGAUAACUUUGUUUGUGUAAACAGCGACGCGACAUAUCGCCCUCCAUGGGAGUUUCCUCUCCAUUCCACAUCCUGUGCCCCUCCACCUUUAC .(((((((((((.((((((((((......((((....))))(((....))))).))))))))..))))))))))).((((......))))....(((.....)))............... ( -38.90) >DroEre_CAF1 3796 120 - 1 GGGGGUGGGCGACUGUCGUGUGUAAAAUUGAUAACUUUGUUUGUGUAAACAGCGACGCGACAUAUCGCCCUCCAAGGGAGCUUCCUCUCCAUCCCGCAUCCUGUGGCCCCACACCUUUCC (((((.((((((.((((((((((......((((....))))(((....))))).))))))))..)))))).))...((((......))))...((((.....)))).))).......... ( -40.40) >DroYak_CAF1 3900 117 - 1 --GGAGGAACGACUGUCGUGUGUAAAAUUGAUAACUUUGUUUGUGUAAACAGCGACGCGACAUAUCGCCCUCCAUGAAAGCUUCCUCUCCAUCCCGCAUCCUGUG-CUCCAUACCUUUGC --(((((..(((.((((((((((......((((....))))(((....))))).))))))))..))).)))))......................((((...)))-)............. ( -24.20) >consensus GAUGGGGGGCGACUGUCGUGUGUAAAAUUGAUAACUUUGUUUGUGUAAACAGCGACGCGACAUAUCGCCCUCCAUGGGAGUUUCCUCUCCAUUCCGCAUCCUGUGCCCCUCCACCUUUAC ...(((((((((.((((((((((......((((....))))(((....))))).))))))))..)))))))))..(((((....))))).....(((.....)))............... (-30.22 = -30.90 + 0.68)

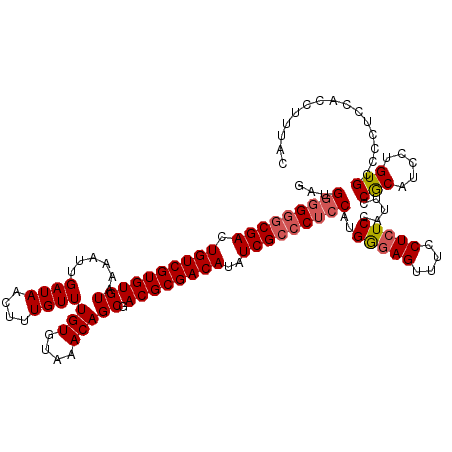

| Location | 7,323,763 – 7,323,875 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.88 |
| Mean single sequence MFE | -25.26 |
| Consensus MFE | -22.78 |
| Energy contribution | -22.30 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.87 |
| Structure conservation index | 0.90 |
| SVM decision value | 3.10 |
| SVM RNA-class probability | 0.998441 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7323763 112 + 27905053 CUCCCAUGGAGGGCGAUAUGUCGCGUCGCUGUUUACACAAACAAAGUUAUCAAUUUUACACACGACAGUCGCCCCCCAUCCACCAU-----CCCGCACC---ACACAUCCACAUCCCCAU .....((((.(((((((.(((((.((...(((((....)))))..((..........)))).)))))))))))).)))).......-----........---.................. ( -25.70) >DroSec_CAF1 96 106 + 1 CUCCCAUGGAGGGCGAUAUGUCGCGUCGCUGUUUACACAAACAAAGUUAUCAAUUUUACACACGACAGUCGCCCCCCAUCAACCAU-----CCCGCAUC---------CCACAUCCCCAU .....((((.(((((((.(((((.((...(((((....)))))..((..........)))).)))))))))))).)))).......-----........---------............ ( -25.70) >DroSim_CAF1 3853 106 + 1 CUCCCAUGGAGGGCGAUAUGUCGCGUCGCUGUUUACACAAACAAAGUUAUCAAUUUUACACACGACAGUCGCCCCCCAUUCACCAU-----CCCGCAUC---------CGACAUCCCCAU .....((((.(((((((.(((((.((...(((((....)))))..((..........)))).)))))))))))).)))).......-----........---------............ ( -25.70) >DroEre_CAF1 3836 119 + 1 CUCCCUUGGAGGGCGAUAUGUCGCGUCGCUGUUUACACAAACAAAGUUAUCAAUUUUACACACGACAGUCGCCCACCCCC-CGCACCCGCACCCGCACCCACACCCAUCCGCACACCCAU .......((((((((((.(((((.((...(((((....)))))..((..........)))).))))))))))))......-.((....)).................))).......... ( -26.20) >DroYak_CAF1 3939 112 + 1 CUUUCAUGGAGGGCGAUAUGUCGCGUCGCUGUUUACACAAACAAAGUUAUCAAUUUUACACACGACAGUCGUUCCUCC---CUCAU-----CCCGCAGCCAUCCGCAUACCCAUACCCAU .......((((((((((.(((((.((...(((((....)))))..((..........)))).))))))))).))))))---.....-----...((........)).............. ( -23.00) >consensus CUCCCAUGGAGGGCGAUAUGUCGCGUCGCUGUUUACACAAACAAAGUUAUCAAUUUUACACACGACAGUCGCCCCCCAUC_ACCAU_____CCCGCACC____C_CAUCCACAUCCCCAU .......((.(((((((.(((((.((...(((((....)))))..((..........)))).)))))))))))).))........................................... (-22.78 = -22.30 + -0.48)


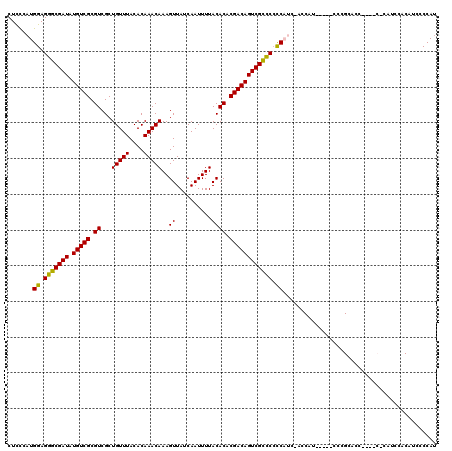
| Location | 7,323,875 – 7,323,984 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.08 |
| Mean single sequence MFE | -37.74 |
| Consensus MFE | -23.12 |
| Energy contribution | -24.50 |
| Covariance contribution | 1.38 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.561201 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7323875 109 - 27905053 UGCCAUUCGCUGCUCC--UACCUGGCUGCUCCU-UCU-GCAUCCCCUCCCC------UGCCUUAUCCCAACCACUUUG-GAAGCCCUUCCGCUGGGAAGUGCUGUGCUGGGGAAACGGUG .......(((((....--.....((.(((....-...-))).))..(((((------.((......((((.....)))-).(((.(((((....))))).)))..)).)))))..))))) ( -32.80) >DroSec_CAF1 202 107 - 1 UACCAUUCGCUGCUCCUCUACCUUGCGGCUCCU-CUU-GCUC-----CCCC------AGCCUUAUCCCUACCACUUUGGGAAGCCCUUCCGCUGGGAAGUGCUGUGCUGGGGAAACGGUG ((((....(((((...........)))))....-...-....-----((((------(((....((((.........))))(((.(((((....))))).)))..)))))))....)))) ( -41.10) >DroSim_CAF1 3959 107 - 1 UACCAUUCGCUGCUCCUCUACCUUGCGGCUCCU-CUU-GCUC-----CCCC------AGCCUUAUCCCUGCCACUUUGGGAAGCCCUUCCGCUGGGAAGUGCUGUGCUGGGGAAACGGUG ((((....(((((...........)))))....-...-....-----((((------(((....((((.........))))(((.(((((....))))).)))..)))))))....)))) ( -41.10) >DroYak_CAF1 4051 116 - 1 UGCCAUUCGCUGCUCC--UUCCUUUCUGCUCCCUCCUGGCUG-CUCCUUCCUGGGCGCUCCUUAUCCCAACCACUUUGGGAAGCCCUUCCGCUG-GAAGUGCCGUGCUGGGAGAACGGGU .(((....((......--.........))...((((..((.(-(..(.....)((((((((...((((((.....))))))(((......))))-).))))))))))..))))....))) ( -35.96) >consensus UACCAUUCGCUGCUCC__UACCUUGCGGCUCCU_CCU_GCUC_____CCCC______AGCCUUAUCCCAACCACUUUGGGAAGCCCUUCCGCUGGGAAGUGCUGUGCUGGGGAAACGGUG .(((....(((((...........)))))..................((((.............((((((.....))))))(((.(((((....))))).))).....))))....))). (-23.12 = -24.50 + 1.38)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:43:27 2006