
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,153,467 – 7,153,571 |
| Length | 104 |
| Max. P | 0.663516 |

| Location | 7,153,467 – 7,153,571 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.42 |
| Mean single sequence MFE | -33.87 |
| Consensus MFE | -19.36 |
| Energy contribution | -20.03 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.34 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.663516 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7153467 104 + 27905053 GGGUGUUCCAUACUUUUUGGCGUUGUUUGAUUUUUAGGCGCAGCA------AAAUGCGUUUUGCUCUUGAUUGAAAGUGUGAAGAGUGUGGGGCA---GCAUGCCAGCGUGUG------- ...((((((((((((((((.((...((..(((...(((((((...------...))))))).......)))..))..))))))))))))))))))---(((((....))))).------- ( -34.10) >DroVir_CAF1 134408 111 + 1 UCGUGUUGCAUACUUUUUGGCGCUAUUUGAUUUUUAGGCGCAGCAAACACAAAAUGCGUUCGCCGCUUGAUUGGGAUAGCUGUUUGUGUGAGGCAUGUGCAUGUCUCUGCC--------- .(((((..(((((.....((((((((((.(...(((((((..((.(((.(.....).))).))))))))).).))))))).))).)))))..))))).(((......))).--------- ( -32.00) >DroSec_CAF1 86376 104 + 1 GGGUGUUCCAUACUUUUUGGCGUUGUUUGAUUUUUAGGCGCAGCA------AAAUGCGUUUUGCUCUUGAUUGGAAGUGAGAAAAGUGUGGGGCG---GCAUGCCAGCGUGUG------- ...(((((((((((((((.((....((..(((...(((((((...------...))))))).......)))..)).))..)))))))))))))))---(((((....))))).------- ( -35.00) >DroSim_CAF1 98959 104 + 1 GGGUGUUCCAUACUUUUUGGCGUUGUUUGAUUUUUAGGCGCAGCA------AAAUGCGUUUUGCUCUUGAUUGGAAGUGUGAAGAGUGUGGGGCG---GCAUGCCAGCGUGUG------- ...((((((((((((((((.((...((..(((...(((((((...------...))))))).......)))..))..))))))))))))))))))---(((((....))))).------- ( -33.90) >DroYak_CAF1 89930 111 + 1 GGGUGUUGCAUACUUUUUGGCGUUGUUUGAUUUUUAGGCGCAGCA------AAAUGCGUUUUGCUCUUGAUUGGAAGUGUGAAGAGUGUGGGGCG---GCAUGCCAGUGUGUGGCAGCGC ..(((((((((((...((((((((((((.((....(((((((...------...))))))).((((((.((.......)).)))))))).)))))---))..))))).)))).))))))) ( -39.00) >DroAna_CAF1 95038 100 + 1 --ACGCCGCAUACUUUUCGGCGUUGUUUGAUUUUUAGGCGCAGCA------AAAUGCGUUUUGCUCUUGAUUGGAUGCCCGAAGGAUGAGAGGAG---GC---CCAGGCUC------CAC --((((((.(.....).))))))..(((.((((((.((.((((((------(((....))))))(((.....)))))))).)))))).)))((((---.(---....))))------).. ( -29.20) >consensus GGGUGUUCCAUACUUUUUGGCGUUGUUUGAUUUUUAGGCGCAGCA______AAAUGCGUUUUGCUCUUGAUUGGAAGUGUGAAGAGUGUGGGGCG___GCAUGCCAGCGUGUG_______ ........(((((((((((((....((..(((...(((((((............))))))).......)))..)).)).))))))))))).((((......))))............... (-19.36 = -20.03 + 0.67)

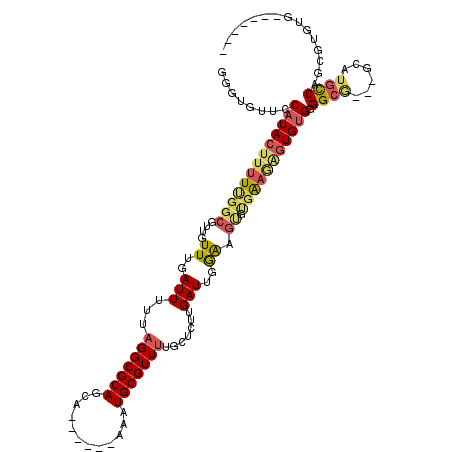
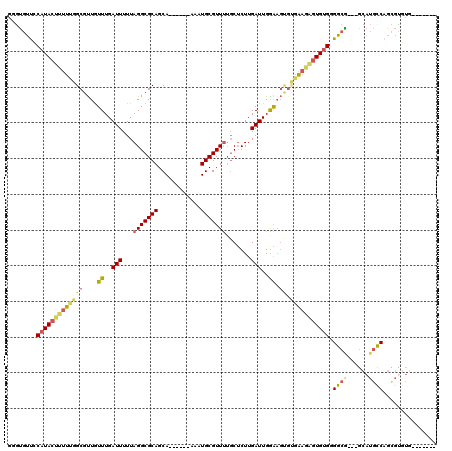
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:41:42 2006