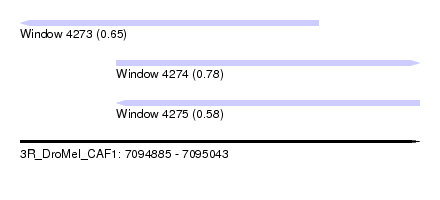

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,094,885 – 7,095,043 |
| Length | 158 |
| Max. P | 0.784222 |

| Location | 7,094,885 – 7,095,003 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.58 |
| Mean single sequence MFE | -30.34 |
| Consensus MFE | -22.32 |
| Energy contribution | -21.84 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.652722 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7094885 118 - 27905053 CUCCUGGAUAACUUAAUUAUCCGCAGAACCACCACCUGGUGGUUGAAAGCUGAACCCUCAAACCAUUUCUCCGAACUUGUUCAGGAAGCCAAACAACAGAGGGCGCUACUAUUAAAAC-- .....((((((.....))))))((..(((((((....)))))))..........(((((..........((((((....))).))).(.....)....))))).))............-- ( -28.90) >DroSec_CAF1 27978 120 - 1 CUCCUGGAUAACUUAAUUAUCCGCAGAACCACCACCUGGUGGUUAAAAGCUGAACCCUCAAUCCAUUUAUCCGUACUUGUUCAGGAAACCAAACAACAGAGGGCGCUACUAUUAAAACGC .....((((((.....))))))((..(((((((....)))))))...(((....(((((((.....))........(((((..(....)..)))))..))))).)))...........)) ( -29.60) >DroSim_CAF1 30778 120 - 1 CUCCUGGAUAACUUAAUUAUCCGCAGAACCACCACCUGGUGGUUGAAAGCUGAACCCUUAAACCAUUUAUCCGAACUUGUUGAGGAAAUCAAACAACAGAGGGCGCUACUAAUAAAACGC .....((((((.....))))))((..(((((((....)))))))...(((....(((((....(........)....(((((..(....)...)))))))))).)))...........)) ( -26.10) >DroEre_CAF1 28763 120 - 1 CUUCUGGAUAACUUAAUUAUCCGGAGUGCCACCACCUGGUGGUUGAAAGCUGAACCCUUAAAGCAUUUAUCCGAACUUGCUUAGGAAGCCAGGCAACAGAGAGCGCCACUAUUAAGAAGC (((((((((((.....))))))(..((((((((....))))((((...(((....(((..(((((............)))))))).)))....)))).....))))..).....))))). ( -32.70) >DroYak_CAF1 29666 120 - 1 CUUCUGGAUAACUUAAUUAUCCGGAGUGCCACCACCUGGUGGUUGAAAGAUGAACCCUUAAACCAUUUAUCCGAACUGGCUCGGGCAGCCAAGCAACAGAGGGCGCCACUUAUAAAAAGC ..(((((((((.....)))))))))(((.(.((..(((.((((((..(((((...........))))).(((((......))))))))))).....)))..)).).)))........... ( -34.40) >consensus CUCCUGGAUAACUUAAUUAUCCGCAGAACCACCACCUGGUGGUUGAAAGCUGAACCCUUAAACCAUUUAUCCGAACUUGUUCAGGAAGCCAAACAACAGAGGGCGCUACUAUUAAAACGC .....((((((.....))))))....(((((((....)))))))...((..(..(((((....(........)...(((((..(....)..)))))..)))))..)..)).......... (-22.32 = -21.84 + -0.48)


| Location | 7,094,923 – 7,095,043 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.75 |
| Mean single sequence MFE | -34.82 |
| Consensus MFE | -25.92 |
| Energy contribution | -25.96 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.784222 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7094923 120 + 27905053 ACAAGUUCGGAGAAAUGGUUUGAGGGUUCAGCUUUCAACCACCAGGUGGUGGUUCUGCGGAUAAUUAAGUUAUCCAGGAGUGAGAGAGCAGUAUUCUCUUCGCGAAAUGGAUAAAAUGAU ....(((((.((((.....(((((((.....)))))))(((((....))))))))).))))).((((..(((((((...(((((((((.....))))).))))....)))))))..)))) ( -41.20) >DroSec_CAF1 28018 120 + 1 ACAAGUACGGAUAAAUGGAUUGAGGGUUCAGCUUUUAACCACCAGGUGGUGGUUCUGCGGAUAAUUAAGUUAUCCAGGAGUGAGAGAGCAGUAUUCUCUUUGCGAAAUGGAUAAAAUGAA ...................((((..((((.((....(((((((....)))))))..))))))..)))).(((((((...(..((((((.....))))).)..)....)))))))...... ( -31.30) >DroSim_CAF1 30818 120 + 1 ACAAGUUCGGAUAAAUGGUUUAAGGGUUCAGCUUUCAACCACCAGGUGGUGGUUCUGCGGAUAAUUAAGUUAUCCAGGAGUGAGAGAGCAGUAUUCUCUUGGCGAUAUGGAUAAAAUGAA ....(((((.....((.(((.(((((..(.((((((.((((((....))))))(((..(((((((...))))))).)))....)))))).)....)))))))).)).)))))........ ( -32.10) >DroEre_CAF1 28803 120 + 1 GCAAGUUCGGAUAAAUGCUUUAAGGGUUCAGCUUUCAACCACCAGGUGGUGGCACUCCGGAUAAUUAAGUUAUCCAGAAGCGAGAGCGCACUGUUCUCUUGGCGAAGAGGAUAAAAUGAA ((......((((...(......)..)))).((((((..(((((....)))))...((.(((((((...))))))).))...))))))))..(((((((((....)))))))))....... ( -35.10) >DroYak_CAF1 29706 120 + 1 GCCAGUUCGGAUAAAUGGUUUAAGGGUUCAUCUUUCAACCACCAGGUGGUGGCACUCCGGAUAAUUAAGUUAUCCAGAAGUGAGAGAGCACUGUUCUCUUGGCGAAAUGGAUAAAAUAAA .(((.((((..........(((((((..((.(((((..(((((....)))))(((((.(((((((...))))))).).)))).)))))...))..))))))))))).))).......... ( -34.40) >consensus ACAAGUUCGGAUAAAUGGUUUAAGGGUUCAGCUUUCAACCACCAGGUGGUGGUUCUGCGGAUAAUUAAGUUAUCCAGGAGUGAGAGAGCAGUAUUCUCUUGGCGAAAUGGAUAAAAUGAA ....(((((............((((......))))...(((((....))))).....)))))..(((..(((((((...(((((((((.....))))))).))....)))))))..))). (-25.92 = -25.96 + 0.04)



| Location | 7,094,923 – 7,095,043 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.75 |
| Mean single sequence MFE | -26.91 |
| Consensus MFE | -18.98 |
| Energy contribution | -19.14 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.578602 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7094923 120 - 27905053 AUCAUUUUAUCCAUUUCGCGAAGAGAAUACUGCUCUCUCACUCCUGGAUAACUUAAUUAUCCGCAGAACCACCACCUGGUGGUUGAAAGCUGAACCCUCAAACCAUUUCUCCGAACUUGU ..............((((.((.((((.....(((.((........((((((.....))))))....(((((((....))))))))).)))(((....))).....))))))))))..... ( -26.20) >DroSec_CAF1 28018 120 - 1 UUCAUUUUAUCCAUUUCGCAAAGAGAAUACUGCUCUCUCACUCCUGGAUAACUUAAUUAUCCGCAGAACCACCACCUGGUGGUUAAAAGCUGAACCCUCAAUCCAUUUAUCCGUACUUGU ((((((((.........((..(((((.......))))).......((((((.....))))))))..(((((((....))))))).)))).)))).......................... ( -23.60) >DroSim_CAF1 30818 120 - 1 UUCAUUUUAUCCAUAUCGCCAAGAGAAUACUGCUCUCUCACUCCUGGAUAACUUAAUUAUCCGCAGAACCACCACCUGGUGGUUGAAAGCUGAACCCUUAAACCAUUUAUCCGAACUUGU (((...(((((((........(((((.......)))))......)))))))........((.((..(((((((....)))))))....)).))...................)))..... ( -22.84) >DroEre_CAF1 28803 120 - 1 UUCAUUUUAUCCUCUUCGCCAAGAGAACAGUGCGCUCUCGCUUCUGGAUAACUUAAUUAUCCGGAGUGCCACCACCUGGUGGUUGAAAGCUGAACCCUUAAAGCAUUUAUCCGAACUUGC (((........(((((....)))))...((((((((.(((((.((((((((.....)))))))))))((((((....)))))).)).)))(((....)))..))))).....)))..... ( -31.30) >DroYak_CAF1 29706 120 - 1 UUUAUUUUAUCCAUUUCGCCAAGAGAACAGUGCUCUCUCACUUCUGGAUAACUUAAUUAUCCGGAGUGCCACCACCUGGUGGUUGAAAGAUGAACCCUUAAACCAUUUAUCCGAACUGGC ..........(((.((((...(((((.......)))))((((.((((((((.....))))))))))))(((((....)))))..((.(((((...........))))).)))))).))). ( -30.60) >consensus UUCAUUUUAUCCAUUUCGCCAAGAGAAUACUGCUCUCUCACUCCUGGAUAACUUAAUUAUCCGCAGAACCACCACCUGGUGGUUGAAAGCUGAACCCUUAAACCAUUUAUCCGAACUUGU .................((((.((((.......))))(((((...((((((.....))))))....(((((((....)))))))...)).))).......................)))) (-18.98 = -19.14 + 0.16)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:52 2006