
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,091,611 – 7,091,750 |
| Length | 139 |
| Max. P | 0.979687 |

| Location | 7,091,611 – 7,091,717 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 85.62 |
| Mean single sequence MFE | -21.74 |
| Consensus MFE | -18.22 |
| Energy contribution | -18.26 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.541243 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7091611 106 + 27905053 CCAUCUAGCUGCCAUGUUCGGUUCAAUUGUUUGCCAUUCUUGGCCAUUGUCUGGACAUCUCCAGUGUCCAUUCUGAGGCCCCCCCCCAUCUCCAGUGUCCUCCAUC .......((((..(((...(((..........)))......((((...(..(((((((.....)))))))..)...))))......)))...)))).......... ( -22.60) >DroSec_CAF1 24692 95 + 1 CCAUGUAGCUGCCAUGUUCGGUUCAAUUGUUUGCCAUUCUUGGCCAUUGUCUGGACAUCUCCAGUGUCCAUUCCGAGGCCC---CCCAUCUCCAGUGU-------- .......((((..(((...(((..........)))......((((.(((..(((((((.....)))))))...))))))).---..)))...))))..-------- ( -22.70) >DroSim_CAF1 27449 100 + 1 CCAUGUAGCUGCCAUGUUCGGUUCAAUUGUUUGCCAUUCUUGGCCAUUGUCUGGACAUCUCCAGUGUCCAUUCUGAGGCCC---CCCAUCUCCAGUGUCG---CUC ....((.((((..(((...(((..........)))......((((...(..(((((((.....)))))))..)...)))).---..)))...))))...)---).. ( -22.80) >DroEre_CAF1 25416 90 + 1 CCAUGUAGCUGCCAUGUUCGGUUCAAUUGUUUGCCAUUCUUGGCCAUUGUCUGGACGUCUCCAGU-UCCAUUCUAAAGC-U---CC---CUCC--------CCAUC ..(((.(((((..((((((((..((((.....((((....)))).)))).))))))))...))))-).)))........-.---..---....--------..... ( -20.10) >DroYak_CAF1 26326 94 + 1 CCAUGUAGCUGCCAUGUUCGGUUCAAUUGUUUGCCAUUCUUGGCCAUUGUCUGGACAUCUCCAGU-UCCAUUCUGAAACCU---CCCCUCCCC--------CCAUC ..(((.(((((..((((((((..((((.....((((....)))).)))).))))))))...))))-).)))..........---.........--------..... ( -20.50) >consensus CCAUGUAGCUGCCAUGUUCGGUUCAAUUGUUUGCCAUUCUUGGCCAUUGUCUGGACAUCUCCAGUGUCCAUUCUGAGGCCC___CCCAUCUCCAGUGU___CCAUC ..(((..((((..((((((((..((((.....((((....)))).)))).))))))))...))))...)))................................... (-18.22 = -18.26 + 0.04)


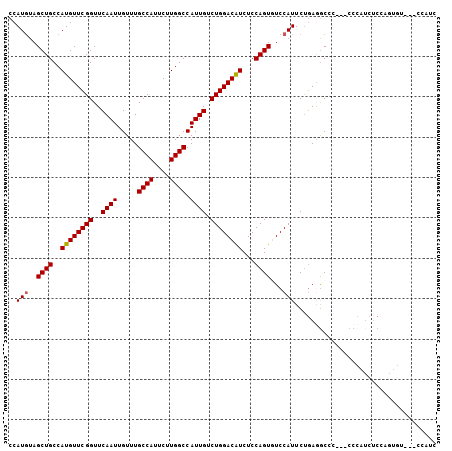
| Location | 7,091,642 – 7,091,750 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 85.13 |
| Mean single sequence MFE | -21.26 |
| Consensus MFE | -15.70 |
| Energy contribution | -16.95 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.79 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.84 |
| SVM RNA-class probability | 0.979687 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
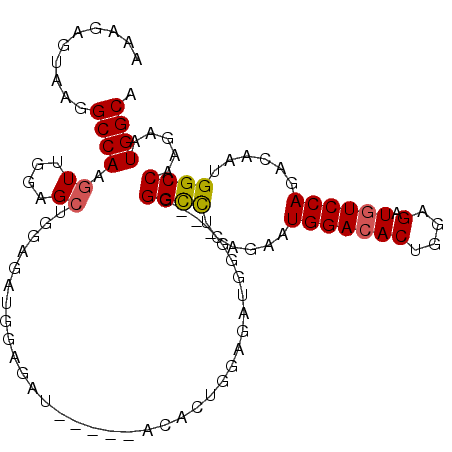
>3R_DroMel_CAF1 7091642 108 + 27905053 UGCCAUUCUUGGCCAUUGUCUGGACAUCUCCAGUGUCCAUUCUGAGGCCCCCCCCCAUCUCCAGUGUCCUCCAUCUCCAUCUCCAGCUCCAACUUGGCCUUACUCUUU .((((....))))....(..(((((((.....)))))))..)(((((((.........((..((((...........)).))..)).........)))))))...... ( -21.27) >DroSec_CAF1 24723 96 + 1 UGCCAUUCUUGGCCAUUGUCUGGACAUCUCCAGUGUCCAUUCCGAGGCCC---CCCAUCUCCAGUGU---------CCAUCUCCUGCUCCAACUUGGCCUUACUCUUU .((((....)))).......(((((((.....)))))))....((((((.---........(((.(.---------......)))).........))))))....... ( -23.47) >DroSim_CAF1 27480 102 + 1 UGCCAUUCUUGGCCAUUGUCUGGACAUCUCCAGUGUCCAUUCUGAGGCCC---CCCAUCUCCAGUGUCG---CUCUCCAUCUCCAGCUCCAACUUGGCCUUACUCUUU .((((....))))....(..(((((((.....)))))))..)(((((((.---..(((.....)))..(---((..........)))........)))))))...... ( -23.90) >DroYak_CAF1 26357 96 + 1 UGCCAUUCUUGGCCAUUGUCUGGACAUCUCCAGU-UCCAUUCUGAAACCU---CCCCUCCCC--------CCAUCUUCAACUCCAGCUCCAAGUUGGCCGUACUCUUU .((((..(((((...(((.(((((....)))))(-((......)))....---.........--------.............)))..))))).)))).......... ( -16.40) >consensus UGCCAUUCUUGGCCAUUGUCUGGACAUCUCCAGUGUCCAUUCUGAGGCCC___CCCAUCUCCAGUGU_____AUCUCCAUCUCCAGCUCCAACUUGGCCUUACUCUUU .((((....))))....(..(((((((.....)))))))..)(((((((..............................................)))))))...... (-15.70 = -16.95 + 1.25)



| Location | 7,091,642 – 7,091,750 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 85.13 |
| Mean single sequence MFE | -25.62 |
| Consensus MFE | -18.88 |
| Energy contribution | -19.00 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.561147 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
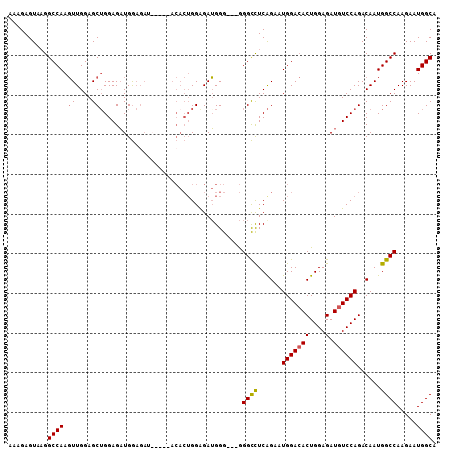
>3R_DroMel_CAF1 7091642 108 - 27905053 AAAGAGUAAGGCCAAGUUGGAGCUGGAGAUGGAGAUGGAGGACACUGGAGAUGGGGGGGGGCCUCAGAAUGGACACUGGAGAUGUCCAGACAAUGGCCAAGAAUGGCA .........(((((..(((...(((((.((.....(((.(..((.......(((((.....)))))...))..).)))...)).))))).)))))))).......... ( -26.60) >DroSec_CAF1 24723 96 - 1 AAAGAGUAAGGCCAAGUUGGAGCAGGAGAUGG---------ACACUGGAGAUGGG---GGGCCUCGGAAUGGACACUGGAGAUGUCCAGACAAUGGCCAAGAAUGGCA ..........((((........(((.......---------...)))........---.((((...(..(((((((....).))))))..)...)))).....)))). ( -25.30) >DroSim_CAF1 27480 102 - 1 AAAGAGUAAGGCCAAGUUGGAGCUGGAGAUGGAGAG---CGACACUGGAGAUGGG---GGGCCUCAGAAUGGACACUGGAGAUGUCCAGACAAUGGCCAAGAAUGGCA ..........(((((((.(..(((..........))---)..)))).........---.((((......(((((((....).))))))......)))).....)))). ( -27.70) >DroYak_CAF1 26357 96 - 1 AAAGAGUACGGCCAACUUGGAGCUGGAGUUGAAGAUGG--------GGGGAGGGG---AGGUUUCAGAAUGGA-ACUGGAGAUGUCCAGACAAUGGCCAAGAAUGGCA ..........((((.((((((((....))).....((.--------..(((.(..---.((((((.....)))-))).....).)))...))....)))))..)))). ( -22.90) >consensus AAAGAGUAAGGCCAAGUUGGAGCUGGAGAUGGAGAU_____ACACUGGAGAUGGG___GGGCCUCAGAAUGGACACUGGAGAUGUCCAGACAAUGGCCAAGAAUGGCA ..........((((.((....))....................................((((......(((((((....).))))))......)))).....)))). (-18.88 = -19.00 + 0.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:47 2006