
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 7,054,468 – 7,054,646 |
| Length | 178 |
| Max. P | 0.946032 |

| Location | 7,054,468 – 7,054,566 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 85.39 |
| Mean single sequence MFE | -43.62 |
| Consensus MFE | -34.67 |
| Energy contribution | -35.45 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.88 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.887002 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7054468 98 + 27905053 AUAGUGAAAGGAAUCGG---CAGCUGCUUCAAGUGUGGGUGUGGCCAGUGUGCUGCCCACACCGCUGGCCGAGUGAAAGACCUCGAAGGAGGUCUGCAGUG ...((........((((---((((.((.....))...(((((((.(((....))).)))))))))).))))).....(((((((....))))))))).... ( -42.80) >DroSec_CAF1 4079 98 + 1 AUAGUGAAAGGAAUCGG---CAGAUGCUUCCAGUGUGGGUGUGGCCAGUGUGCUGCCCACACCGCUGGCCGAGUGAAAGACCUCGAAGGAGGUCUGCAGUG ...((........((((---(.((....))((((...(((((((.(((....))).)))))))))))))))).....(((((((....))))))))).... ( -42.00) >DroSim_CAF1 5452 98 + 1 AUAGUGAAAGGAAUCGG---CAGCUGCUUCCAGUGUGGGUGUGGCCAGUGUGCUGCCCACACCGCUGGCCGAGUGAAAGACCUCGAAGGAGGUCUGCAGUG .................---..(((((..(((((...(((((((.(((....))).)))))))))))).........(((((((....)))))))))))). ( -43.10) >DroEre_CAF1 1939 98 + 1 GUAGUGGAAAGAGUCGG---CAGCUGCUUCCAGCGUGGGUGUGGCCAGUGUGCUGCACACACCGCUGGCCGAAUGGAAGACCUCGAAUGAGGUCUGCAGUG ...((.((.....)).)---).(((((..((((((((.(((..((......))..))).)).)))))).........(((((((....)))))))))))). ( -41.60) >DroYak_CAF1 1938 98 + 1 GUAGUGGAAGGAGUCUG---CACCUGCUUCCAGUGUAGGUGUGGCCAGCGUGCUGCCUACACCGCUGGCCGAAUGAAAGACCUCGAAGGAGGUCUGCAGUG ..............(((---(((((((.......)))))).(((((((((((.......)).)))))))))......(((((((....))))))))))).. ( -43.80) >DroAna_CAF1 1966 101 + 1 GUAGGCGAAGGAUUCGCCUCCAGCCGCUUCCAUGCCAGGCGUAGCCAGGGCAGUGCCCACAUUGGUGGCGGAGUGAAAGACCUCGAAAGAGGUCUGCAACG (.((((((.....)))))).)..(((((.(((.....((((..((....))..)))).....))).))))).((...(((((((....)))))))...)). ( -48.40) >consensus AUAGUGAAAGGAAUCGG___CAGCUGCUUCCAGUGUGGGUGUGGCCAGUGUGCUGCCCACACCGCUGGCCGAGUGAAAGACCUCGAAGGAGGUCUGCAGUG ......................(((((..(((((...(((((((.(((....))).)))))))))))).........(((((((....)))))))))))). (-34.67 = -35.45 + 0.78)



| Location | 7,054,468 – 7,054,566 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 85.39 |
| Mean single sequence MFE | -38.82 |
| Consensus MFE | -31.15 |
| Energy contribution | -31.98 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.16 |
| Mean z-score | -3.40 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.36 |
| SVM RNA-class probability | 0.946032 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7054468 98 - 27905053 CACUGCAGACCUCCUUCGAGGUCUUUCACUCGGCCAGCGGUGUGGGCAGCACACUGGCCACACCCACACUUGAAGCAGCUG---CCGAUUCCUUUCACUAU ......(((((((....))))))).....(((((.((((((((((.(((....))).)))))))((....)).....))))---))))............. ( -38.70) >DroSec_CAF1 4079 98 - 1 CACUGCAGACCUCCUUCGAGGUCUUUCACUCGGCCAGCGGUGUGGGCAGCACACUGGCCACACCCACACUGGAAGCAUCUG---CCGAUUCCUUUCACUAU ......(((((((....))))))).....(((((..(.(((((((.(((....))).))))))))....((....))...)---))))............. ( -35.60) >DroSim_CAF1 5452 98 - 1 CACUGCAGACCUCCUUCGAGGUCUUUCACUCGGCCAGCGGUGUGGGCAGCACACUGGCCACACCCACACUGGAAGCAGCUG---CCGAUUCCUUUCACUAU ......(((((((....))))))).....(((((.((((((((((.(((....))).))))))).....((....))))))---))))............. ( -40.40) >DroEre_CAF1 1939 98 - 1 CACUGCAGACCUCAUUCGAGGUCUUCCAUUCGGCCAGCGGUGUGUGCAGCACACUGGCCACACCCACGCUGGAAGCAGCUG---CCGACUCUUUCCACUAC ......(((((((....))))))).....(((((..(.((((((..(((....)))..)))))))..((((....)))).)---))))............. ( -37.10) >DroYak_CAF1 1938 98 - 1 CACUGCAGACCUCCUUCGAGGUCUUUCAUUCGGCCAGCGGUGUAGGCAGCACGCUGGCCACACCUACACUGGAAGCAGGUG---CAGACUCCUUCCACUAC ..(((((((((((....))))))........((((((((.(((.....))))))))))).........(((....))).))---))).............. ( -39.10) >DroAna_CAF1 1966 101 - 1 CGUUGCAGACCUCUUUCGAGGUCUUUCACUCCGCCACCAAUGUGGGCACUGCCCUGGCUACGCCUGGCAUGGAAGCGGCUGGAGGCGAAUCCUUCGCCUAC .((((((((((((....))))))).....((((((((....)).)))..((((..(((...))).)))).))).)))))...((((((.....)))))).. ( -42.00) >consensus CACUGCAGACCUCCUUCGAGGUCUUUCACUCGGCCAGCGGUGUGGGCAGCACACUGGCCACACCCACACUGGAAGCAGCUG___CCGAUUCCUUCCACUAC ..(((((((((((....))))))).........((((.(((((((.(((....))).)))))))....))))..))))....................... (-31.15 = -31.98 + 0.84)



| Location | 7,054,500 – 7,054,606 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 89.37 |
| Mean single sequence MFE | -43.35 |
| Consensus MFE | -32.00 |
| Energy contribution | -32.62 |
| Covariance contribution | 0.61 |
| Combinations/Pair | 1.18 |
| Mean z-score | -3.04 |
| Structure conservation index | 0.74 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.514067 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
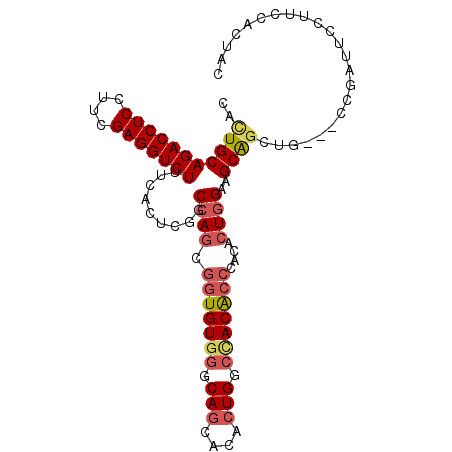
>3R_DroMel_CAF1 7054500 106 - 27905053 UCACUGCAGCUGCUGGAUGAGAAGGAGCAGGAGAUUAAGACACUGCAGACCUCCUUCGAGGUCUUUCACUCGGCCAGCGGUGUGGGCAGCACACUGGCCACACCCA ........(((((((..(((((....((((............)))).((((((....)))))))))))..))).))))(((((((.(((....))).))))))).. ( -48.10) >DroSec_CAF1 4111 106 - 1 UCACUGCAGCUGCUGGACGAGAAGGAGCAGGAGAUCAAGACACUGCAGACCUCCUUCGAGGUCUUUCACUCGGCCAGCGGUGUGGGCAGCACACUGGCCACACCCA ...........(((((.((((.....((((............))))(((((((....)))))))....)))).)))))(((((((.(((....))).))))))).. ( -49.10) >DroSim_CAF1 5484 106 - 1 UCACUGCAGCUGCUGGACGAGAAGGAGCAGGAGAUCAAGACACUGCAGACCUCCUUCGAGGUCUUUCACUCGGCCAGCGGUGUGGGCAGCACACUGGCCACACCCA ...........(((((.((((.....((((............))))(((((((....)))))))....)))).)))))(((((((.(((....))).))))))).. ( -49.10) >DroEre_CAF1 1971 106 - 1 UCACUGCAGCUUCUUGAUGAGAAAGAGCAGGAGAUCAAGACACUGCAGACCUCAUUCGAGGUCUUCCAUUCGGCCAGCGGUGUGUGCAGCACACUGGCCACACCCA ....(((((..(((((((...............)))))))..)))))((((((....))))))........((((((..((((.....)))).))))))....... ( -39.56) >DroYak_CAF1 1970 106 - 1 UCACUGCAGCUUCUUGAUGAGAAGGAGCAGGAGAUCAAGACACUGCAGACCUCCUUCGAGGUCUUUCAUUCGGCCAGCGGUGUAGGCAGCACGCUGGCCACACCUA ....(((((..(((((((...............)))))))..)))))((((((....))))))........((((((((.(((.....)))))))))))....... ( -42.26) >DroAna_CAF1 2001 106 - 1 UCCCUCCAGCUUCUUGACGAAAAAGAGCAGGAGAUCAAAACGUUGCAGACCUCUUUCGAGGUCUUUCACUCCGCCACCAAUGUGGGCACUGCCCUGGCUACGCCUG ...((((.(((.(((.......)))))).))))...........(((((((((....)))))))........((((.......((((...))))))))...))... ( -32.01) >consensus UCACUGCAGCUGCUGGACGAGAAGGAGCAGGAGAUCAAGACACUGCAGACCUCCUUCGAGGUCUUUCACUCGGCCAGCGGUGUGGGCAGCACACUGGCCACACCCA ........(((......((((.....((((............))))(((((((....)))))))....))))...)))(((((((.(((....))).))))))).. (-32.00 = -32.62 + 0.61)


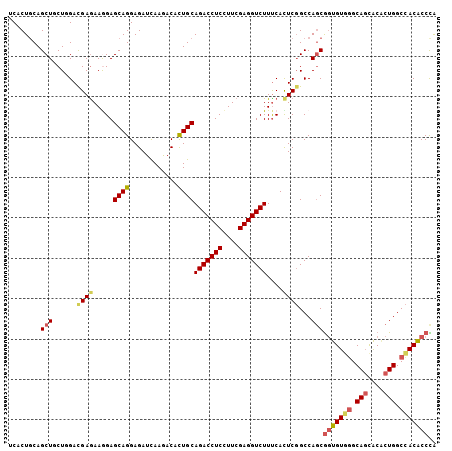
| Location | 7,054,530 – 7,054,646 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 82.68 |
| Mean single sequence MFE | -44.75 |
| Consensus MFE | -26.40 |
| Energy contribution | -27.10 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.33 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.728883 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7054530 116 + 27905053 ---UGGCCGAGUGAAAGACCUCGAAGGAGGUCUGCAGUGUCUUAAUCUCCUGCUCCUUCUCAUCCAGCAGCUGCAGUGACCUCUGGCGCUGCUUCUGCAGUAGCUGUUCCAGCUCGUUC ---(((..(((.((.(((((((....)))))))((((.(.........)))))))...)))..)))((((..((((((.((...))))))))..)))).(.(((((...)))))).... ( -44.20) >DroVir_CAF1 2014 116 + 1 ---UGGGCUUGUGAAACACCUCGAAGGAGGUUUGCAGCGUCUUAAUCUCCUGCUCCUUCUCGUCGAGCAACUGUAAUGAACGCUGCCGCUGCUUUUGGAGCAGCUGCUCCAGCUCGUUA ---.((((..(((....(((((....)))))..(((((((..........(((((.........)))))..........))))))))))......((((((....)))))))))).... ( -42.05) >DroPse_CAF1 1991 119 + 1 CACCGGCUGUAUGAAAUACCUCAAAUGAGGUCUGCAGCGUCUUGAUCUCCUGCUCCUUCUCGUCGAGAAGCUGCAGGGAUCGCUGUCGCUGCUUCUGCAGCAGCUGCUCCAGCUCGUUG ((.((((((((.(((..(((((....)))))..((((((.(.(((((.(((((..((((((...))))))..))))))))))..).)))))))))))))))(((((...))))))).)) ( -55.10) >DroGri_CAF1 2235 116 + 1 ---UGGGCUUGUGAAACACUUCGAAGGAGGUUUGCAAUGUUUUAAUCUCCUGCUCCUUCUCGUCGAGCAGCUGCAGCGAUCGUUGCCGCUGCUUCUGCAGCAACUGUUCCAGCUCAUUC ---((((((.(.(((.((..((((((((((...(((..............))).))))))..))))((((..((((((........))))))..))))......))))))))))))... ( -38.94) >DroMoj_CAF1 1988 116 + 1 ---UUGGCUUGUGGAACACCUCGAAGGAGGUCUGCAACGUUUUAAUCUCCUGCUCCUUCUCGUCGAGCAGUUGGAGUGAACGCUGCCGCUGUUUUUGCAGCAACUGCUCCAGCUCGUUC ---..((((.(..(...(((((....)))))..(((.(((((...(((.((((((.........))))))..)))..))))).))).(((((....)))))..)..)...))))..... ( -39.60) >DroAna_CAF1 2031 116 + 1 ---UGGCGGAGUGAAAGACCUCGAAAGAGGUCUGCAACGUUUUGAUCUCCUGCUCUUUUUCGUCAAGAAGCUGGAGGGAUCGCUGGCGCUGCUUCUGCAGCAACUGCUCCAGUUCGUUG ---.((((((.((..(((((((....)))))))(((..(((.(((((.(((.(.((((((....))))))..).))))))))..)))(((((....)))))...)))..)).)))))). ( -48.60) >consensus ___UGGCCGUGUGAAACACCUCGAAGGAGGUCUGCAACGUCUUAAUCUCCUGCUCCUUCUCGUCGAGCAGCUGCAGUGAUCGCUGCCGCUGCUUCUGCAGCAACUGCUCCAGCUCGUUC .....(((.........(((((....)))))((((((............((((((.........))))))))))))........)))(((((....)))))(((((...)))))..... (-26.40 = -27.10 + 0.70)


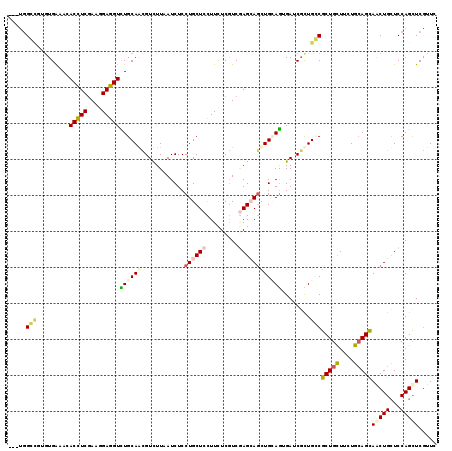
| Location | 7,054,530 – 7,054,646 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 82.68 |
| Mean single sequence MFE | -43.21 |
| Consensus MFE | -23.80 |
| Energy contribution | -23.89 |
| Covariance contribution | 0.09 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.73 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.643058 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 7054530 116 - 27905053 GAACGAGCUGGAACAGCUACUGCAGAAGCAGCGCCAGAGGUCACUGCAGCUGCUGGAUGAGAAGGAGCAGGAGAUUAAGACACUGCAGACCUCCUUCGAGGUCUUUCACUCGGCCA--- (..((((.((((.((((..((((((..((..(....)..))..))))))..))))...........((((............))))(((((((....)))))))))))))))..).--- ( -42.10) >DroVir_CAF1 2014 116 - 1 UAACGAGCUGGAGCAGCUGCUCCAAAAGCAGCGGCAGCGUUCAUUACAGUUGCUCGACGAGAAGGAGCAGGAGAUUAAGACGCUGCAAACCUCCUUCGAGGUGUUUCACAAGCCCA--- ....(.((((((((.((((((.....)))))).((((((((........((((((.........))))))........))))))))..(((((....))))))))))...))).).--- ( -41.49) >DroPse_CAF1 1991 119 - 1 CAACGAGCUGGAGCAGCUGCUGCAGAAGCAGCGACAGCGAUCCCUGCAGCUUCUCGACGAGAAGGAGCAGGAGAUCAAGACGCUGCAGACCUCAUUUGAGGUAUUUCAUACAGCCGGUG ...((.((((..(((((((((((....)))))).....(((((((((..((((((...))))))..))))).)))).....)))))..(((((....)))))........))))))... ( -55.30) >DroGri_CAF1 2235 116 - 1 GAAUGAGCUGGAACAGUUGCUGCAGAAGCAGCGGCAACGAUCGCUGCAGCUGCUCGACGAGAAGGAGCAGGAGAUUAAAACAUUGCAAACCUCCUUCGAAGUGUUUCACAAGCCCA--- ...((.(((((((((((((..((((..(((((((......)))))))..)))).))))..(((((((...(..((......))..)....)))))))....))))))...))).))--- ( -39.00) >DroMoj_CAF1 1988 116 - 1 GAACGAGCUGGAGCAGUUGCUGCAAAAACAGCGGCAGCGUUCACUCCAACUGCUCGACGAGAAGGAGCAGGAGAUUAAAACGUUGCAGACCUCCUUCGAGGUGUUCCACAAGCCAA--- ....(.((((((((...(((((......)))))((((((((..((((...(((((.........))))))))).....))))))))..(((((....))))))))))...))))..--- ( -40.60) >DroAna_CAF1 2031 116 - 1 CAACGAACUGGAGCAGUUGCUGCAGAAGCAGCGCCAGCGAUCCCUCCAGCUUCUUGACGAAAAAGAGCAGGAGAUCAAAACGUUGCAGACCUCUUUCGAGGUCUUUCACUCCGCCA--- (((((..((((....((((((.....)))))).)))).((((..(((.(((.(((.......)))))).)))))))....))))).(((((((....)))))))............--- ( -40.80) >consensus GAACGAGCUGGAGCAGCUGCUGCAGAAGCAGCGGCAGCGAUCACUGCAGCUGCUCGACGAGAAGGAGCAGGAGAUUAAAACGCUGCAGACCUCCUUCGAGGUGUUUCACAAGCCCA___ ........(((((..((((((.....))))))(.(((((.......(..((((((.........))))))..).......))))))..(((((....))))).)))))........... (-23.80 = -23.89 + 0.09)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:27 2006