
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 6,957,901 – 6,957,995 |
| Length | 94 |
| Max. P | 0.922370 |

| Location | 6,957,901 – 6,957,995 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 95.64 |
| Mean single sequence MFE | -30.18 |
| Consensus MFE | -25.38 |
| Energy contribution | -25.74 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.737605 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6957901 94 + 27905053 ACAACGAAUGCAUGAAUUCACCGGUCUCAAGGUAAAGAACCGUGGUCUAAGUAGCCCGGAAAAGCACAGCGGUGGUAUUUACGGUGGUUUUGCC .........(((.....((((((...(((.(((.....))).)))..((((((.((((...........))).).))))))))))))...))). ( -21.40) >DroSec_CAF1 7093 94 + 1 ACAACGAAUGCAUGAAUUCACCGGUCUCAAGGUAAAGAACCGGGGUCUAAGUAGCCCGGAAAACCACCGCGGUGGUAUUUGCGGUGGUUUUGCC .....((((......))))(((........)))......(((((.(......).)))))(((((((((((((......)))))))))))))... ( -34.30) >DroSim_CAF1 7126 94 + 1 ACAACGAAUGCAUGAAUUCACCGGUCUCAAGGUAAAGAACCGGGGUCUAAGUAGCCUGGAAAACCACCGCGGUGGUAUUUGCGGUGGUUUUGCC .....((((......)))).(((((.((........)))))))(((.......))).(((((((((((((((......))))))))))))).)) ( -33.20) >DroEre_CAF1 5409 94 + 1 ACAACGAAUGCAUGAAUUCACCGGUCUCAAGGUAAAGAACCGGAGUCUAAGUAGCCCGGAAAACCACCGCGGUGGUAUUUGCGGUGGUUUUGCC .....((((......)))).(((((.((........)))))))..............(((((((((((((((......))))))))))))).)) ( -32.00) >DroYak_CAF1 8051 94 + 1 ACAACGAAUGCAUGAAUUCACCGGUCUCAAGGUAAAGAGCCGUGGUCUAAGUAGUCCGGAAAACCACUGCGGUGGUAUUGGCGGUGGUUUUGCC .....((((......))))..(((.(((........)))))).((.((....)).))(((((((((((((.(......).))))))))))).)) ( -30.00) >consensus ACAACGAAUGCAUGAAUUCACCGGUCUCAAGGUAAAGAACCGGGGUCUAAGUAGCCCGGAAAACCACCGCGGUGGUAUUUGCGGUGGUUUUGCC .....((((......)))).((((.(((..(((.....)))(....)...).)).))))(((((((((((((......)))))))))))))... (-25.38 = -25.74 + 0.36)


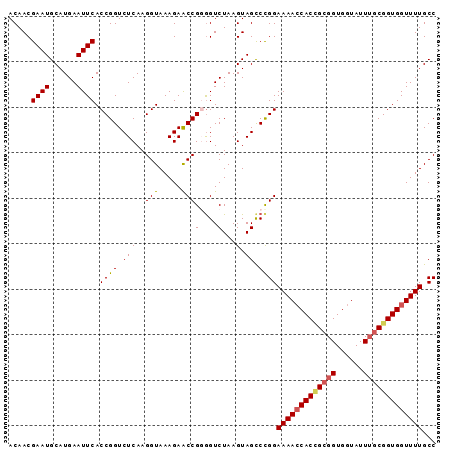
| Location | 6,957,901 – 6,957,995 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 95.64 |
| Mean single sequence MFE | -27.24 |
| Consensus MFE | -24.56 |
| Energy contribution | -25.24 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.922370 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6957901 94 - 27905053 GGCAAAACCACCGUAAAUACCACCGCUGUGCUUUUCCGGGCUACUUAGACCACGGUUCUUUACCUUGAGACCGGUGAAUUCAUGCAUUCGUUGU .(((....(((((...........((((((.(((...((....)).))).))))))((((......)))).)))))......)))......... ( -17.20) >DroSec_CAF1 7093 94 - 1 GGCAAAACCACCGCAAAUACCACCGCGGUGGUUUUCCGGGCUACUUAGACCCCGGUUCUUUACCUUGAGACCGGUGAAUUCAUGCAUUCGUUGU ((.(((((((((((..........)))))))))))))(((((....)).)))((((.(((......)))))))..((((......))))..... ( -32.20) >DroSim_CAF1 7126 94 - 1 GGCAAAACCACCGCAAAUACCACCGCGGUGGUUUUCCAGGCUACUUAGACCCCGGUUCUUUACCUUGAGACCGGUGAAUUCAUGCAUUCGUUGU ((.(((((((((((..........))))))))))))).((((....)).))(((((.(((......)))))))).((((......))))..... ( -31.40) >DroEre_CAF1 5409 94 - 1 GGCAAAACCACCGCAAAUACCACCGCGGUGGUUUUCCGGGCUACUUAGACUCCGGUUCUUUACCUUGAGACCGGUGAAUUCAUGCAUUCGUUGU ((.(((((((((((..........)))))))))))))..((..........(((((.(((......)))))))).........))......... ( -31.61) >DroYak_CAF1 8051 94 - 1 GGCAAAACCACCGCCAAUACCACCGCAGUGGUUUUCCGGACUACUUAGACCACGGCUCUUUACCUUGAGACCGGUGAAUUCAUGCAUUCGUUGU (((.........))).....((((...(((((((...(.....)..)))))))(((((........))).)))))).................. ( -23.80) >consensus GGCAAAACCACCGCAAAUACCACCGCGGUGGUUUUCCGGGCUACUUAGACCCCGGUUCUUUACCUUGAGACCGGUGAAUUCAUGCAUUCGUUGU ((.(((((((((((..........))))))))))))).(((..........((((.((((......)))))))).((((......))))))).. (-24.56 = -25.24 + 0.68)

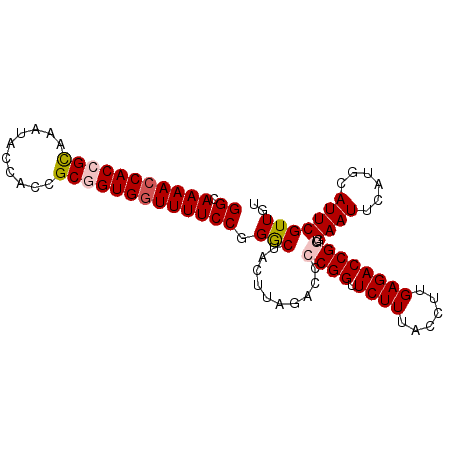
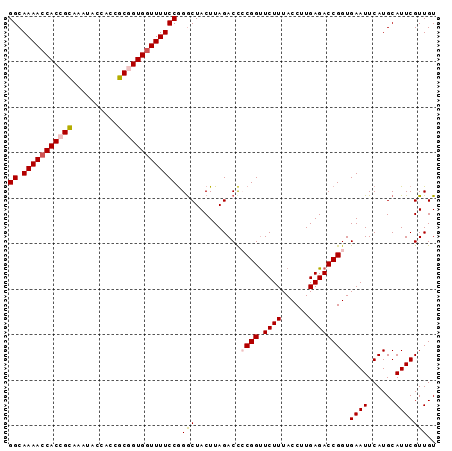
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:39:38 2006