
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 996,622 – 996,727 |
| Length | 105 |
| Max. P | 0.757399 |

| Location | 996,622 – 996,727 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 82.06 |
| Mean single sequence MFE | -33.04 |
| Consensus MFE | -23.01 |
| Energy contribution | -24.58 |
| Covariance contribution | 1.57 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.715444 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
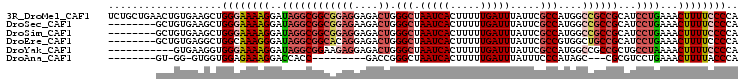
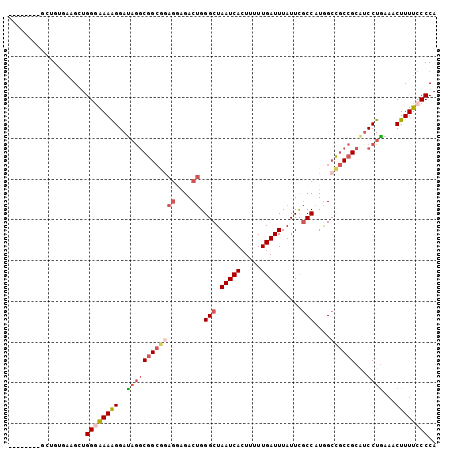
>3R_DroMel_CAF1 996622 105 + 27905053 UCUGCUGAACUGUGAAGCUGGGAAAAGGAUAGGCGGCGGAGGAGACUGGGCUAAUCACUUUUUGAUUUAUUCGCCAUGGCCGCCGCAUCCUGAAACUUUUCCCCA ...(((.........))).((((((((..((((((((((((....)).(((.(((((.....))))).....)))....)))))))...)))...)))))))).. ( -37.20) >DroSec_CAF1 41468 97 + 1 --------GCUGUGAAGCUGGGAAAAGGAUAGGCGGCGGAGAAGACUGGGCUAAUCACUUUUUGAUUUAUUCGCCAUGGCCGCCGCAUCCUGAAACUUUUCCCCA --------(((....))).((((((((..((((((((((.........(((.(((((.....))))).....)))....)))))))...)))...)))))))).. ( -34.02) >DroSim_CAF1 43858 97 + 1 --------GCUGUGAAGCUGGGAAAAGGAUAGGCGGCGGAGGAGACUGGGCUAAUCACUUUUUGAUUUAUUCGCCAUGGCCGCCGCAUCCUGAAACUUUUCCCCA --------(((....))).((((((((..((((((((((((....)).(((.(((((.....))))).....)))....)))))))...)))...)))))))).. ( -38.00) >DroEre_CAF1 43734 97 + 1 --------GCUGUGAGGCUGGCAAAGGGAUAGGCGGCACAGGAGACUGGGCUAAUCACUUUUUGAUUUAUUCGCCGUGGCUGCCGCAUCCUGAAACUUUUCCCCA --------(((........)))...(((((.((((((.(((....)))(((.(((((.....))))).....)))...))))))..))))).............. ( -32.30) >DroYak_CAF1 46058 94 + 1 -----------GUGAAGGUGGGAAAAGGAUAGGCGGAAGAGGAGACUGGGCUAAUCACUUUUUGAUUUAUUCGCCAUGGCCGCCGCUGCCUAAAACUUUUCCCCA -----------......(.((((((((..(((((((...((....)).(((.(((((.....))))).....((....)).))).)))))))...))))))))). ( -33.90) >DroAna_CAF1 41367 83 + 1 --------GU-GG-GUGGUGGAGAAAGGACCACC---------GACCGGGCUAAUCACUUUUUGAUUUAUUUCCCAUAGC---CGCGUCCUGAAACUUUUACCCA --------.(-((-(((((((........)))))---------(((..(((((((((.....))))..........))))---)..)))...........))))) ( -22.80) >consensus ________GCUGUGAAGCUGGGAAAAGGAUAGGCGGCGGAGGAGACUGGGCUAAUCACUUUUUGAUUUAUUCGCCAUGGCCGCCGCAUCCUGAAACUUUUCCCCA ...................((((((((..((((((((((((....)).(((.(((((.....))))).....)))....))))))...))))...)))))))).. (-23.01 = -24.58 + 1.57)


| Location | 996,622 – 996,727 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 82.06 |
| Mean single sequence MFE | -30.17 |
| Consensus MFE | -21.96 |
| Energy contribution | -23.31 |
| Covariance contribution | 1.35 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.757399 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
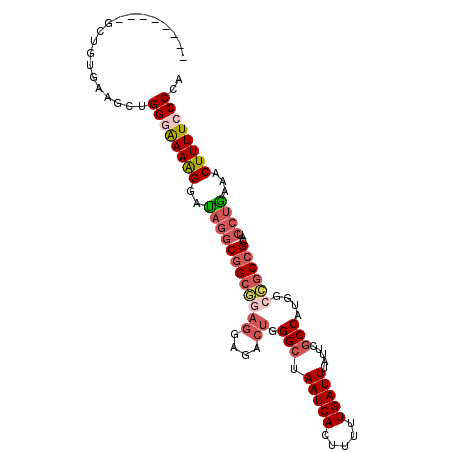
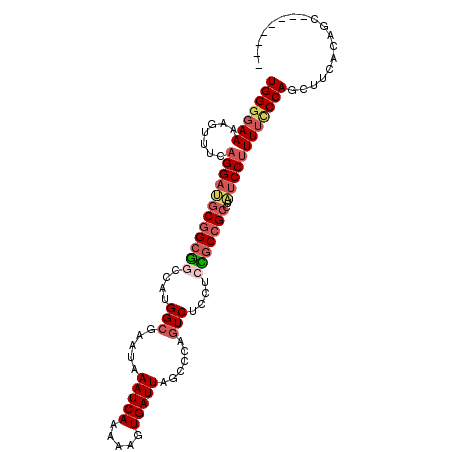
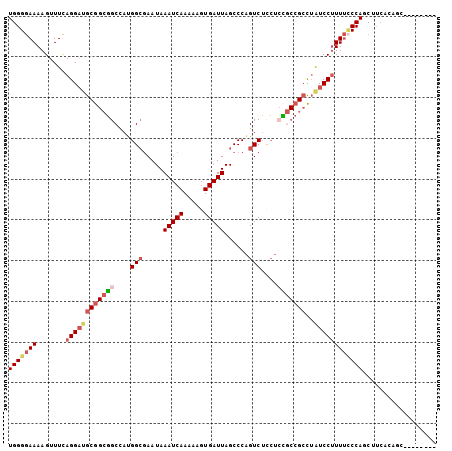
>3R_DroMel_CAF1 996622 105 - 27905053 UGGGGAAAAGUUUCAGGAUGCGGCGGCCAUGGCGAAUAAAUCAAAAAGUGAUUAGCCCAGUCUCCUCCGCCGCCUAUCCUUUUCCCAGCUUCACAGUUCAGCAGA (((((((.......((((((.((((((...((.((...(((((.....))))).......)).))...)))))))))))))))))))(((.........)))... ( -33.01) >DroSec_CAF1 41468 97 - 1 UGGGGAAAAGUUUCAGGAUGCGGCGGCCAUGGCGAAUAAAUCAAAAAGUGAUUAGCCCAGUCUUCUCCGCCGCCUAUCCUUUUCCCAGCUUCACAGC-------- (((((((.......((((((((((((....(((.....(((((.....)))))......)))....)))))))..))))))))))))(((....)))-------- ( -32.01) >DroSim_CAF1 43858 97 - 1 UGGGGAAAAGUUUCAGGAUGCGGCGGCCAUGGCGAAUAAAUCAAAAAGUGAUUAGCCCAGUCUCCUCCGCCGCCUAUCCUUUUCCCAGCUUCACAGC-------- (((((((.......((((((.((((((...((.((...(((((.....))))).......)).))...)))))))))))))))))))(((....)))-------- ( -32.11) >DroEre_CAF1 43734 97 - 1 UGGGGAAAAGUUUCAGGAUGCGGCAGCCACGGCGAAUAAAUCAAAAAGUGAUUAGCCCAGUCUCCUGUGCCGCCUAUCCCUUUGCCAGCCUCACAGC-------- (((((....((....((((((((((.....(((.....(((((.....))))).)))(((....)))))))))..))))....))...)))))....-------- ( -27.50) >DroYak_CAF1 46058 94 - 1 UGGGGAAAAGUUUUAGGCAGCGGCGGCCAUGGCGAAUAAAUCAAAAAGUGAUUAGCCCAGUCUCCUCUUCCGCCUAUCCUUUUCCCACCUUCAC----------- ..((((((((...(((((((.((.(((...(((.....(((((.....))))).)))..))).)).))...)))))..))))))))........----------- ( -31.00) >DroAna_CAF1 41367 83 - 1 UGGGUAAAAGUUUCAGGACGCG---GCUAUGGGAAAUAAAUCAAAAAGUGAUUAGCCCGGUC---------GGUGGUCCUUUCUCCACCAC-CC-AC-------- (((((..........(..((.(---((((..........((((.....)))))))))))..)---------(((((........)))))))-))-).-------- ( -25.40) >consensus UGGGGAAAAGUUUCAGGAUGCGGCGGCCAUGGCGAAUAAAUCAAAAAGUGAUUAGCCCAGUCUCCUCCGCCGCCUAUCCUUUUCCCAGCUUCACAGC________ (((((((.......((((((((((((....(((.....(((((.....)))))......)))....)))))))..)))))))))))).................. (-21.96 = -23.31 + 1.35)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:38:48 2006