
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 6,939,722 – 6,939,960 |
| Length | 238 |
| Max. P | 0.981806 |

| Location | 6,939,722 – 6,939,830 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 98.77 |
| Mean single sequence MFE | -37.67 |
| Consensus MFE | -37.67 |
| Energy contribution | -37.67 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.63 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.981806 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6939722 108 + 27905053 CAUAAUUAAAGGGGAAAACCUCAAAAGAGCCUUGAACUCAUUCAACUGGCUGUGGCUGCAUUGCGUGCAGUUUGGCCCGAAAGUUGGCCAAGUUGCCCUGGCCGCACA ..........((((....)))).....(((((((((....)))))..))))((((((((...))(.(((..(((((((....)..))))))..))).).))))))... ( -37.20) >DroSim_CAF1 89710 108 + 1 CAUAAUUAAAGGGGAAAACCUCAAAAAAGCCUUGAACUCAUUCAACUGGCUGUGGCUGCAUUGCGUGCAGUUUGGCCCGAAAGUUGGCCAAGUUGCUCUGGCCGCACA ..........((((....)))).....(((((((((....)))))..))))((((((((...))(.(((..(((((((....)..))))))..))).).))))))... ( -37.90) >DroYak_CAF1 90346 108 + 1 CAUAAUUAAAGGGGAAAACCUCAAAAAAGCCUUGAACUCAUUCAACUGGCUGUGGCUGCAUUGCGUGCAGUUUGGCCCGAAAGUUGGCCAAGUUGCUCUGGCCGCACA ..........((((....)))).....(((((((((....)))))..))))((((((((...))(.(((..(((((((....)..))))))..))).).))))))... ( -37.90) >consensus CAUAAUUAAAGGGGAAAACCUCAAAAAAGCCUUGAACUCAUUCAACUGGCUGUGGCUGCAUUGCGUGCAGUUUGGCCCGAAAGUUGGCCAAGUUGCUCUGGCCGCACA ..........((((....)))).....(((((((((....)))))..))))((((((((...))(.(((..(((((((....)..))))))..))).).))))))... (-37.67 = -37.67 + -0.00)

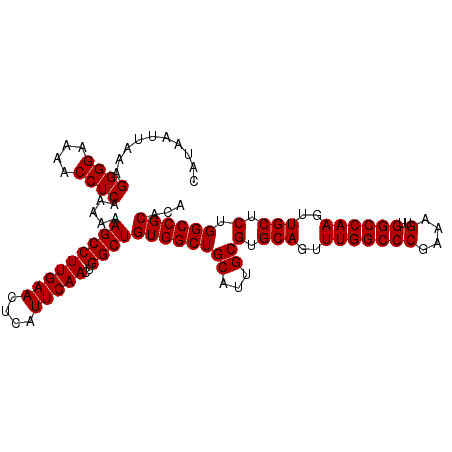

| Location | 6,939,752 – 6,939,858 |
|---|---|
| Length | 106 |
| Sequences | 3 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 94.65 |
| Mean single sequence MFE | -40.77 |
| Consensus MFE | -36.54 |
| Energy contribution | -36.54 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.01 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.50 |
| SVM RNA-class probability | 0.959406 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6939752 106 + 27905053 CUUGAACUCAUUCAACUGGCUGUGGCUGCAUUGCGUGCAGUUUGGCCCGAAAGUUGGCCAAGUUGCCCUGGCCGCACAAUUGUUGCACGCACACAGUCUACAGACC .(((((....)))))..(((((((((((...(((((((((.(((((((....)..))))))((.((....)).)).......)))))))))..))).)))))).)) ( -42.40) >DroSim_CAF1 89740 99 + 1 CUUGAACUCAUUCAACUGGCUGUGGCUGCAUUGCGUGCAGUUUGGCCCGAAAGUUGGCCAAGUUGCUCUGGCCGCACAAUUGUUGCACGCACACAGUC-------C .(((((....)))))..(((((((.......(((((((((.(((((((....)..))))))((.((....)).)).......))))))))))))))))-------. ( -36.31) >DroYak_CAF1 90376 106 + 1 CUUGAACUCAUUCAACUGGCUGUGGCUGCAUUGCGUGCAGUUUGGCCCGAAAGUUGGCCAAGUUGCUCUGGCCGCACAAUUGUUGCACGCACACAGUCCACAGACC .(((((....)))))..(((((((((((...(((((((((.(((((((....)..))))))((.((....)).)).......)))))))))..))).)))))).)) ( -43.60) >consensus CUUGAACUCAUUCAACUGGCUGUGGCUGCAUUGCGUGCAGUUUGGCCCGAAAGUUGGCCAAGUUGCUCUGGCCGCACAAUUGUUGCACGCACACAGUC_ACAGACC .(((((....)))))..(((((((.......(((((((((.(((((((....)..))))))((.((....)).)).......))))))))))))))))........ (-36.54 = -36.54 + -0.00)


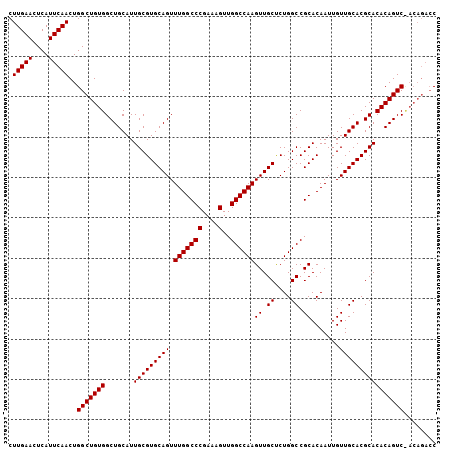
| Location | 6,939,752 – 6,939,858 |
|---|---|
| Length | 106 |
| Sequences | 3 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 94.65 |
| Mean single sequence MFE | -42.63 |
| Consensus MFE | -34.07 |
| Energy contribution | -35.73 |
| Covariance contribution | 1.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.21 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.826141 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6939752 106 - 27905053 GGUCUGUAGACUGUGUGCGUGCAACAAUUGUGCGGCCAGGGCAACUUGGCCAACUUUCGGGCCAAACUGCACGCAAUGCAGCCACAGCCAGUUGAAUGAGUUCAAG ((.((((.(.(((((((((((((....(((.(((((((((....)))))))..(.....))))))..))))))).))))))).))))))..(((((....))))). ( -45.70) >DroSim_CAF1 89740 99 - 1 G-------GACUGUGUGCGUGCAACAAUUGUGCGGCCAGAGCAACUUGGCCAACUUUCGGGCCAAACUGCACGCAAUGCAGCCACAGCCAGUUGAAUGAGUUCAAG (-------(.(((((((((((((.......(((.......)))..((((((........))))))..))))))).))))))))........(((((....))))). ( -35.40) >DroYak_CAF1 90376 106 - 1 GGUCUGUGGACUGUGUGCGUGCAACAAUUGUGCGGCCAGAGCAACUUGGCCAACUUUCGGGCCAAACUGCACGCAAUGCAGCCACAGCCAGUUGAAUGAGUUCAAG ((.((((((.(((((((((((((.......(((.......)))..((((((........))))))..))))))).))))))))))))))..(((((....))))). ( -46.80) >consensus GGUCUGU_GACUGUGUGCGUGCAACAAUUGUGCGGCCAGAGCAACUUGGCCAACUUUCGGGCCAAACUGCACGCAAUGCAGCCACAGCCAGUUGAAUGAGUUCAAG ((.((((.(.(((((((((((((.......(((.......)))..((((((........))))))..))))))).))))))).))))))..(((((....))))). (-34.07 = -35.73 + 1.67)

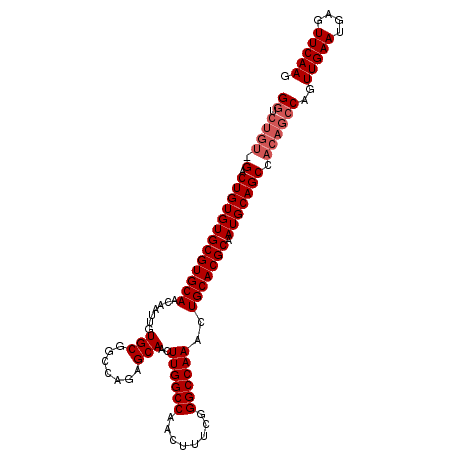

| Location | 6,939,792 – 6,939,891 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 72.91 |
| Mean single sequence MFE | -35.57 |
| Consensus MFE | -14.91 |
| Energy contribution | -15.85 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.42 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.542023 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6939792 99 - 27905053 -GCCCAUAGGCAU------UUUUCCACUCCACUUGACUGUGGUCUGUAGA------------CUGUGUGCGUGCAACAAUUGUGCGGCCAGGGCAACUU--GGCCAACUUUCGGGCCAAA -((((....((((------...((.((.((((......))))...)).))------------....))))(..((.....))..)(((((((....)))--)))).......)))).... ( -39.20) >DroSim_CAF1 89780 92 - 1 -GCCCAUAGGCAU------UUUUCCACUCCACUUGACUGUG-------GA------------CUGUGUGCGUGCAACAAUUGUGCGGCCAGAGCAACUU--GGCCAACUUUCGGGCCAAA -((((....((((------.....((.(((((......)))-------))------------.)).))))(..((.....))..)((((((......))--)))).......)))).... ( -32.90) >DroWil_CAF1 102976 115 - 1 UGCCACUUGCCACUUGCUACUUGCCACUCGAAAGCAAUUUGGUUAGCCGAUUUUGUGUGUUUGUGUAUGCGUGCAACAAAU--GCGGCU---GCAACUUGUGGCCAACUUUUUGCCCAAG .(((((..((((.(((((..(((.....))).)))))..))))((((((..(((((.(((.(((....))).)))))))).--.)))))---)......)))))................ ( -30.80) >DroYak_CAF1 90416 99 - 1 -GCCCAUCGGCAU------UUUUCCACUCCACUUGACUGUGGUCUGUGGA------------CUGUGUGCGUGCAACAAUUGUGCGGCCAGAGCAACUU--GGCCAACUUUCGGGCCAAA -((((....((((------...(((((.((((......))))...)))))------------....))))(..((.....))..)((((((......))--)))).......)))).... ( -39.40) >consensus _GCCCAUAGGCAU______UUUUCCACUCCACUUGACUGUGGUCUGU_GA____________CUGUGUGCGUGCAACAAUUGUGCGGCCAGAGCAACUU__GGCCAACUUUCGGGCCAAA .((.....(((.................((((......))))...........................((((((.....)))))))))...)).......((((........))))... (-14.91 = -15.85 + 0.94)



| Location | 6,939,858 – 6,939,960 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 80.43 |
| Mean single sequence MFE | -24.71 |
| Consensus MFE | -16.54 |
| Energy contribution | -16.43 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.754499 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6939858 102 + 27905053 ACAGUCAAGUGGAGUGGAAAAAUGCCUAUGGGCGUGUCC------CCACU--GGAAGGCCAAAUGCACUAGGAAAUUGUAUAGGAAAAAUCUUGGAAAAUUAUAGAACCA .......(((((...(((...(((((....))))).)))------)))))--((...((.....)).(((...((((...(((((....)))))...)))).)))..)). ( -23.30) >DroSim_CAF1 89839 91 + 1 ACAGUCAAGUGGAGUGGAAAAAUGCCUAUGGGCGUGUCC------CCACU--GGAAGGCCAA-----------AAAUAUAUAGGAAAAAUCUUGGAAAAUUAUAGAACAA .(..((.(((((...(((...(((((....))))).)))------)))))--.))..)....-----------.......(((((....)))))................ ( -19.80) >DroYak_CAF1 90482 110 + 1 ACAGUCAAGUGGAGUGGAAAAAUGCCGAUGGGCGUGUCCAUCCUUCCAGUGGGGAAGGCCAAAUGCACUAGGAAAAUUUGUAAGAAAUGUUUUGGAAACUUAUAAAACCA ..(((((..(((.(((((...(((((....))))).)))))((((((.....)))))))))..)).))).((....((((((((..............)))))))).)). ( -31.04) >consensus ACAGUCAAGUGGAGUGGAAAAAUGCCUAUGGGCGUGUCC______CCACU__GGAAGGCCAAAUGCACUAGGAAAAUAUAUAGGAAAAAUCUUGGAAAAUUAUAGAACCA .(..((.(((((...(((...(((((....))))).)))......)))))...))..)......................(((((....)))))................ (-16.54 = -16.43 + -0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:39:28 2006