
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 6,797,875 – 6,797,970 |
| Length | 95 |
| Max. P | 0.834975 |

| Location | 6,797,875 – 6,797,970 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 90.02 |
| Mean single sequence MFE | -28.46 |
| Consensus MFE | -19.18 |
| Energy contribution | -20.38 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.579065 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6797875 95 + 27905053 ACUCUUAUACUCUGGCCCUUGCCAGCCGUUCUGGCUUUCGAAAUCGAAUCGGCGGCAAGUGUUGGCCCACUUACAAGGCUCACACACACACACAC ............((((....))))((((..((((..((((....))))))))))))..((((.((((.........)))).)))).......... ( -26.10) >DroSec_CAF1 56500 87 + 1 ACUCUUAUACGCUGGCCCUUGCCAGCCGUUCUGGCAUUCGAAAUCGAAUCGGCCAACAGCGUUGGCCCACUUACAGGGCUCACAC--------AA ..........((((((....)))))).(((((((.(((((....))))))(((((((...)))))))......))))))......--------.. ( -32.20) >DroSim_CAF1 61633 87 + 1 ACUCUUAUACGCUGGCCCUUGCCAGCCGUUCUGGCUUUCGAAAUCGAAUCGUCGACCAGCGUUGGCCCACUUACAGGGCUCACAC--------AC ..........(((((.....(((((.....))))).((((....)))).......)))))((.(((((.......))))).))..--------.. ( -28.90) >DroEre_CAF1 41777 87 + 1 ACUCUUAUACUCUGGCCCUUGCCAGCCGUUCUAGCUUUCGAAAUCGAAUCGGCGGCAAGCGUUGGCCCACUUACAGGGCUCACAC--------AC .................((((((.((((........((((....)))).)))))))))).((.(((((.......))))).))..--------.. ( -27.90) >DroYak_CAF1 54625 87 + 1 ACUCUUAUACGCUGGCCCUUGGCAGCCGUUCUGGCUUUCGAACUCGAAUCGGCGACAAGCGUUGGCCCACUUACAGGGCUCACAC--------AC .........((((((((...)))((((.....))))((((....)))).)))))....(.((.(((((.......))))).)).)--------.. ( -27.20) >consensus ACUCUUAUACGCUGGCCCUUGCCAGCCGUUCUGGCUUUCGAAAUCGAAUCGGCGACAAGCGUUGGCCCACUUACAGGGCUCACAC________AC ..........((((((....)))))).......(((((((....))))..))).......((.(((((.......))))).))............ (-19.18 = -20.38 + 1.20)



| Location | 6,797,875 – 6,797,970 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 90.02 |
| Mean single sequence MFE | -33.10 |
| Consensus MFE | -25.10 |
| Energy contribution | -25.02 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.834975 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
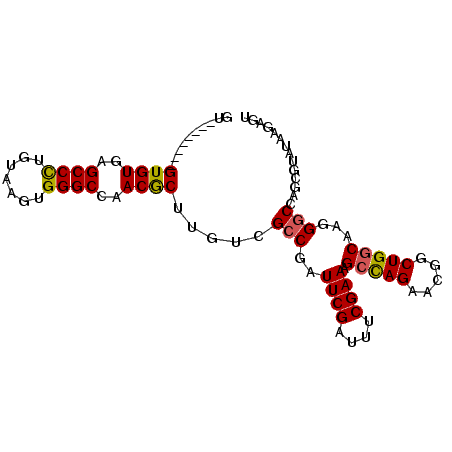
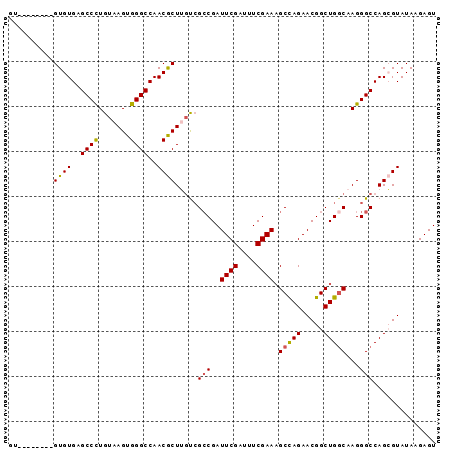
>3R_DroMel_CAF1 6797875 95 - 27905053 GUGUGUGUGUGUGUGAGCCUUGUAAGUGGGCCAACACUUGCCGCCGAUUCGAUUUCGAAAGCCAGAACGGCUGGCAAGGGCCAGAGUAUAAGAGU ................((((((((....((((....((((((((((.(((.....(....)...))))))).))))))))))....)))))).)) ( -32.70) >DroSec_CAF1 56500 87 - 1 UU--------GUGUGAGCCCUGUAAGUGGGCCAACGCUGUUGGCCGAUUCGAUUUCGAAUGCCAGAACGGCUGGCAAGGGCCAGCGUAUAAGAGU ..--------((((..((((.......))))..)))).(((((((.(((((....)))))(((((.....)))))...))))))).......... ( -38.70) >DroSim_CAF1 61633 87 - 1 GU--------GUGUGAGCCCUGUAAGUGGGCCAACGCUGGUCGACGAUUCGAUUUCGAAAGCCAGAACGGCUGGCAAGGGCCAGCGUAUAAGAGU ..--------...((.((((.......))))))(((((((((.....((((....)))).(((((.....)))))...)))))))))........ ( -36.60) >DroEre_CAF1 41777 87 - 1 GU--------GUGUGAGCCCUGUAAGUGGGCCAACGCUUGCCGCCGAUUCGAUUUCGAAAGCUAGAACGGCUGGCAAGGGCCAGAGUAUAAGAGU ..--------.(.((.((((((((((((......))))))).(((..((((....))))((((.....))))))).))))))).).......... ( -30.10) >DroYak_CAF1 54625 87 - 1 GU--------GUGUGAGCCCUGUAAGUGGGCCAACGCUUGUCGCCGAUUCGAGUUCGAAAGCCAGAACGGCUGCCAAGGGCCAGCGUAUAAGAGU ((--------((((..((((((((((((......)))))).)((((.(((..((......))..))))))).....)))))..))))))...... ( -27.40) >consensus GU________GUGUGAGCCCUGUAAGUGGGCCAACGCUUGUCGCCGAUUCGAUUUCGAAAGCCAGAACGGCUGGCAAGGGCCAGCGUAUAAGAGU ..........((((..((((.......))))..)))).....(((..((((....)))).(((((.....)))))...))).............. (-25.10 = -25.02 + -0.08)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:36:58 2006