
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 947,044 – 947,150 |
| Length | 106 |
| Max. P | 0.500000 |

| Location | 947,044 – 947,150 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.81 |
| Mean single sequence MFE | -29.17 |
| Consensus MFE | -12.50 |
| Energy contribution | -12.67 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.43 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
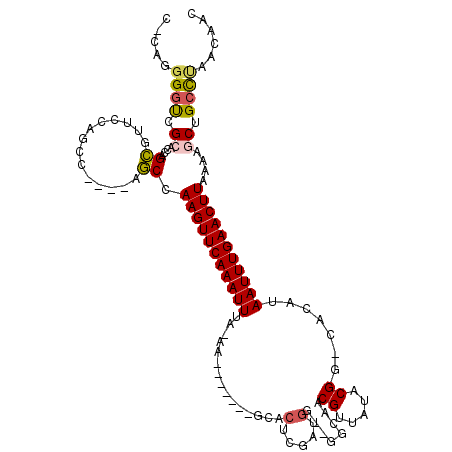
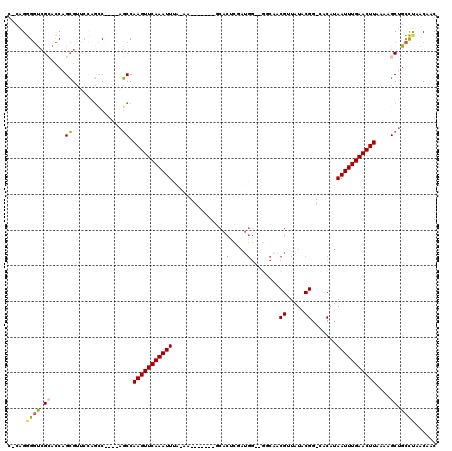
>3R_DroMel_CAF1 947044 106 - 27905053 C-CAGGGGUCGCACCAGCGUUCCAGCC----AGCCAAGUUCAAAUUUA-AA-------GCACUCGAUGGCAGGCAACGUUAUACGG-CACAUAAUUUGAACUUAAAAGCAGCCUAACAAC .-.(((((.(((....)))..)).((.----....(((((((((((..-..-------((.....(((((.(....))))))...)-)....)))))))))))....))..)))...... ( -26.70) >DroSec_CAF1 36101 110 - 1 C-CAGGGGUCGCACCAGCGUUCCAGCCAGCCAGCCAAGUUCAAAUUUA-AA-------GCACUCGAUGGCAGGCAACGUUAUACGG-CACAUAAUUUGAACUUAAAAGCUGCCUAACAAC .-..((((.(((....)))))))((.((((.....(((((((((((..-..-------((.....(((((.(....))))))...)-)....)))))))))))....)))).))...... ( -30.50) >DroSim_CAF1 42114 110 - 1 C-CAGGGGUCGCACCAGCGUUCCAGCCAGCCAGCCAAGUUCAAAUUUA-AA-------GCACUCGAUGGCAGGCAACGUUAUACGG-CACAUAAUUUGAACUUAAAAGCUGCCUAACAAC .-..((((.(((....)))))))((.((((.....(((((((((((..-..-------((.....(((((.(....))))))...)-)....)))))))))))....)))).))...... ( -30.50) >DroEre_CAF1 36887 102 - 1 CCCAGGGGUCGCACCUGCGUUCCAGCC----AGCCAAGUUCAAAUUCA-AA-------CCACAUCAUGG----CCACGUUAUGCGG-CACAUAAUUUGAACUU-AAAGCUGCCUAACAAC ....((((.(((....)))))))((.(----(((.(((((((((((..-..-------((.(((.(((.----...))).))).))-.....)))))))))))-...)))).))...... ( -29.90) >DroAna_CAF1 33717 95 - 1 C-AAGGGGUCGGGCCAGCGG------C----AGCCAAGUUCAAAUUUUUAA-------CCGCACU------GCGAACGGUAUACGG-CACAUAAUUUGAACUUAAAAGCUGCCCAACAAC .-..(..((.((((.(((((------.----..))(((((((((((.....-------(((.(((------(....))))...)))-.....)))))))))))....))))))).))..) ( -29.50) >DroPer_CAF1 123555 111 - 1 C-CAGAGCCAGAGUCAGAGUCAGAGGC----AUCCAAGUUCAAAUUUUUAACCGCACCGCACUCUAUGG----CAACGGUAUACGGGAACAUAAUUUGAACUUAAAAGCUGUUCGGCAAC .-....(((.(((.(((.((.....))----....(((((((((((.....(((.((((..(.....).----...))))...)))......))))))))))).....)))))))))... ( -27.90) >consensus C_CAGGGGUCGCACCAGCGUUCCAGCC____AGCCAAGUUCAAAUUUA_AA_______GCACUCGAUGG__GGCAACGUUAUACGG_CACAUAAUUUGAACUUAAAAGCUGCCUAACAAC .....((((.((....((..............)).(((((((((((...............(.....)........((.....)).......)))))))))))....)).))))...... (-12.50 = -12.67 + 0.17)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:38:31 2006