
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 6,754,234 – 6,754,423 |
| Length | 189 |
| Max. P | 0.999306 |

| Location | 6,754,234 – 6,754,343 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.29 |
| Mean single sequence MFE | -44.68 |
| Consensus MFE | -37.88 |
| Energy contribution | -37.96 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.90 |
| Structure conservation index | 0.85 |
| SVM decision value | 3.50 |
| SVM RNA-class probability | 0.999306 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6754234 109 + 27905053 CGACAUUCUAUAAAUUAAUCAGACGGAGUGCUCUUAGCUCCGCCAGGAGCCAGAAGCCCGGAGCAGG-----------CAGCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCA ...................(((..(((((((((((.((((((((.((.((.....)))).).)).))-----------.))).))))))))))))))((((((.....))))))...... ( -42.90) >DroSec_CAF1 18399 109 + 1 CGACAUUCUAUAAAUUAAUCAGACGGAGUGCUCUUAGCUCCGCCAGGAGCCAGAAGCCCGGAGCAGC-----------CAGCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCA ...................(((..(((((((((((.(((((....))))).....((..((.....)-----------).)).))))))))))))))((((((.....))))))...... ( -41.30) >DroSim_CAF1 22800 109 + 1 CGACAUUCUAUAAAUUAAUCAGACGGAGUGCUCUUAGCUCCGCCAGGAGCCAGAAGCCCGGAGCAGC-----------CAGCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCA ...................(((..(((((((((((.(((((....))))).....((..((.....)-----------).)).))))))))))))))((((((.....))))))...... ( -41.30) >DroEre_CAF1 2787 120 + 1 CGACAUUCUAUAAAUUAAUCAGACGGAGUGCUCUUAGCUCCGCUAGGAGCCAGAAGCCCGGAGAAGCCAGCGAGGAACCAGCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCA .......................(((.(((((((((((...))))))))).....)))))(((..((((((((((..((((.((((....))))))))....))))..))))))..))). ( -47.40) >DroYak_CAF1 14723 120 + 1 CGACAUUCUAUAAAUUAAUCAGACGGAGUGCUCUUAGCUCCGCCAGGAGCAAGAAGCCCGGAGUAUCCAGCGAGGAGCCAGCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCA .......................(((.((..((((.(((((....))))))))).)))))((((..((((((.(((((..((.(((((...))))).))..).))))))))))..)))). ( -50.50) >consensus CGACAUUCUAUAAAUUAAUCAGACGGAGUGCUCUUAGCUCCGCCAGGAGCCAGAAGCCCGGAGCAGC___________CAGCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCA ...................(((..(((((((((((.(((((....))))).....((.....))...................))))))))))))))((((((.....))))))...... (-37.88 = -37.96 + 0.08)


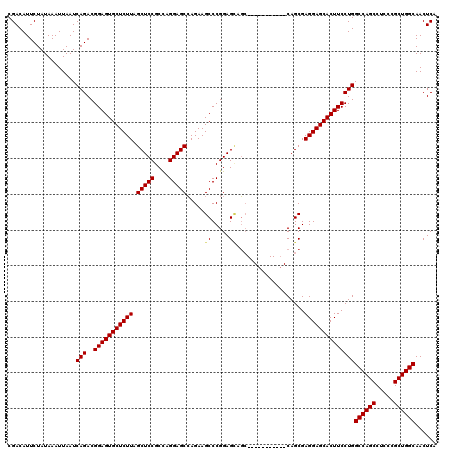
| Location | 6,754,234 – 6,754,343 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.29 |
| Mean single sequence MFE | -49.76 |
| Consensus MFE | -36.86 |
| Energy contribution | -37.06 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.53 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.878339 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6754234 109 - 27905053 UGAGUUGCCAGCGGGAGGCUGGCCAGGAAGUGCUCCUCGCUG-----------CCUGCUCCGGGCUUCUGGCUCCUGGCGGAGCUAAGAGCACUCCGUCUGAUUAAUUUAUAGAAUGUCG ((((((((((((.....))))))((((.(((((((......(-----------((((...)))))...(((((((....))))))).)))))))...))))...)))))).......... ( -43.20) >DroSec_CAF1 18399 109 - 1 UGAGUUGCCAGCGGGAGGCUGGCCAGGAAGUGCUCCUCGCUG-----------GCUGCUCCGGGCUUCUGGCUCCUGGCGGAGCUAAGAGCACUCCGUCUGAUUAAUUUAUAGAAUGUCG .((((.((((((((((((((........))).)))).)))))-----------)).))))((((((..(((((((....)))))))..)))...)))....................... ( -48.00) >DroSim_CAF1 22800 109 - 1 UGAGUUGCCAGCGGGAGGCUGGCCAGGAAGUGCUCCUCGCUG-----------GCUGCUCCGGGCUUCUGGCUCCUGGCGGAGCUAAGAGCACUCCGUCUGAUUAAUUUAUAGAAUGUCG .((((.((((((((((((((........))).)))).)))))-----------)).))))((((((..(((((((....)))))))..)))...)))....................... ( -48.00) >DroEre_CAF1 2787 120 - 1 UGAGUUGCCAGCGGGAGGCUGGCCAGGAAGUGCUCCUCGCUGGUUCCUCGCUGGCUUCUCCGGGCUUCUGGCUCCUAGCGGAGCUAAGAGCACUCCGUCUGAUUAAUUUAUAGAAUGUCG .(((..(((((((((..((..((.((((.....)))).))..))..)))))))))..)))((((((..(((((((....)))))))..)))...)))....................... ( -56.90) >DroYak_CAF1 14723 120 - 1 UGAGUUGCCAGCGGGAGGCUGGCCAGGAAGUGCUCCUCGCUGGCUCCUCGCUGGAUACUCCGGGCUUCUUGCUCCUGGCGGAGCUAAGAGCACUCCGUCUGAUUAAUUUAUAGAAUGUCG .((((..((((((((..((..((.((((.....)))).))..))..))))))))..))))((((((.((((((((....))))).))))))...)))....................... ( -52.70) >consensus UGAGUUGCCAGCGGGAGGCUGGCCAGGAAGUGCUCCUCGCUG___________GCUGCUCCGGGCUUCUGGCUCCUGGCGGAGCUAAGAGCACUCCGUCUGAUUAAUUUAUAGAAUGUCG ((((((((((((.....))))))((((.(((((((((((..(...............)..))))....(((((((....))))))).)))))))...))))...)))))).......... (-36.86 = -37.06 + 0.20)


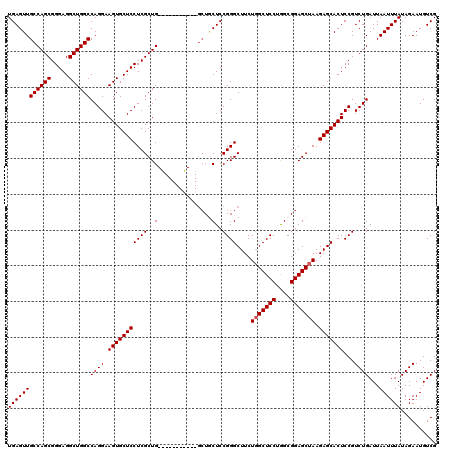
| Location | 6,754,303 – 6,754,423 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 99.50 |
| Mean single sequence MFE | -35.36 |
| Consensus MFE | -34.88 |
| Energy contribution | -34.64 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.02 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.983175 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6754303 120 + 27905053 GCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCAAUCGCGAUAAAGCCCCAGGUGAAUUAUAGUUAUUUGAUUUUGUUUUGGUGAUUUGUUUGCUUGAGUUUCAGUUCUUUAAA ..(((((((........((((((.....))))))(((((((..((((((((.(.(((((.((((((........))))))...))))).).)))).)))))))))))...)))))))... ( -35.60) >DroSec_CAF1 18468 120 + 1 GCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCAAUCGCGAUAAAGCCCCAGGUGAAUUAUAGUUAUUUGAUUUUGUUUUGGUGAUUUGUUUGCUUGAGUUUCAGUUCUUUAAA ..(((((((........((((((.....))))))(((((((..((((((((.(.(((((.((((((........))))))...))))).).)))).)))))))))))...)))))))... ( -35.60) >DroSim_CAF1 22869 120 + 1 GCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCAAUCGCGAUAAAGCCUCAGGUGAAUUAUAGUUAUUUGAUUUUGUUUUGGUGAUUUGUUUGCUUGAGUUUCAGUUCUUUAAA ..(((((((........((((((.....)))))).........((......))((((((..(((...((((((..((......))..)))))).)))..)))))).....)))))))... ( -35.00) >DroEre_CAF1 2867 120 + 1 GCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCAAUCGCGAUAAAGCCCCAGGUGAAUUAUAGUUAUUUGAUUUUGUUUUGGUGAUUUGUUUGCUUGAGUUUCAGUUCUUUAAA ..(((((((........((((((.....))))))(((((((..((((((((.(.(((((.((((((........))))))...))))).).)))).)))))))))))...)))))))... ( -35.60) >DroYak_CAF1 14803 120 + 1 GCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCAAUCGCGAUAAAGCCUCAGGUGAAUUAUAGUUAUUUGAUUUUGUUUUGGUGAUUUGUUUGCUUGAGUUUCAGUUCUUUAAA ..(((((((........((((((.....)))))).........((......))((((((..(((...((((((..((......))..)))))).)))..)))))).....)))))))... ( -35.00) >consensus GCGAGGAGCACUUCCUGGCCAGCCUCCCGCUGGCAACUCAAUCGCGAUAAAGCCCCAGGUGAAUUAUAGUUAUUUGAUUUUGUUUUGGUGAUUUGUUUGCUUGAGUUUCAGUUCUUUAAA ..(((((((........((((((.....))))))(((((((..((((((((.(.(((((.((((((........))))))...))))).).)))).)))))))))))...)))))))... (-34.88 = -34.64 + -0.24)

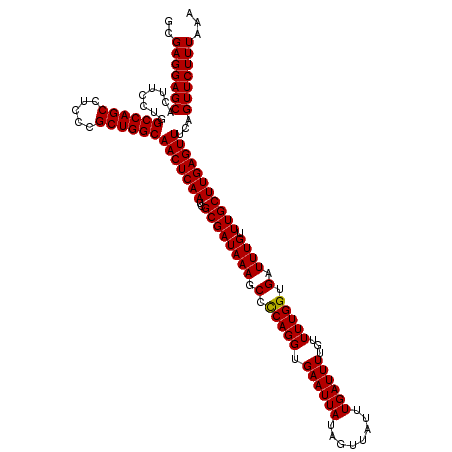

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:34:56 2006