
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 6,749,537 – 6,749,737 |
| Length | 200 |
| Max. P | 0.685080 |

| Location | 6,749,537 – 6,749,657 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.47 |
| Mean single sequence MFE | -23.10 |
| Consensus MFE | -19.61 |
| Energy contribution | -20.37 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.685080 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
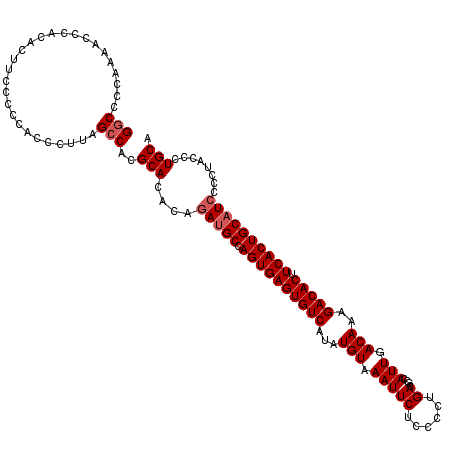
>3R_DroMel_CAF1 6749537 120 - 27905053 GGCCCCAAAAACCACACUUCCGCGUCCCUUAGCCACGCACACAGACGCCAGUGAGUGUCAUAUGUAAAUUCUCCCCUGACGCCAUUGACAAAGACACUUCACUGCAUCCCCAACCCUGCA .((..................(((((.....((...)).....)))))(((((((((((...(((.(((((......))....))).)))..))))).)))))).............)). ( -22.80) >DroSec_CAF1 13778 120 - 1 GGCCCCAAAACCCACACUCCCCCCACCCUUAGCCACGCACACAGAUGCCAGUGAGUGUCAUAUGUAAAUUCUCCCCUGACGCCAUUGACAAAGACACUUCACUGCAUCCCCUAGCCUGCA (((............................((...)).....(((((.((((((((((...(((.(((((......))....))).)))..))))).)))))))))).....))).... ( -24.20) >DroSim_CAF1 18106 120 - 1 GGCCCCAAAACCCACACUUCCCCCACCCUUAGCCACGCACACAGAUGCCAGUGAGUGUCAUAUGUAAAUUCUCCCCUGACGCCAUUGACAAAGACACUUCACUGCAUCCCCUAGCCUGCA (((............................((...)).....(((((.((((((((((...(((.(((((......))....))).)))..))))).)))))))))).....))).... ( -24.20) >DroYak_CAF1 9957 119 - 1 GACCCCAAAACCCGUACUUCCC-CUCCCUUGGCCACGCACACAAAUGCCAGUGAGUGUCAUAUGUAAAUUCUCCCCUGACGCCAUUGACAAAGACACUUCACUGCAUCCCCUGCCCUGCA ......................-........((...(((.....((((.((((((((((...(((.(((((......))....))).)))..))))).)))))))))....)))...)). ( -21.20) >consensus GGCCCCAAAACCCACACUUCCCCCACCCUUAGCCACGCACACAGAUGCCAGUGAGUGUCAUAUGUAAAUUCUCCCCUGACGCCAUUGACAAAGACACUUCACUGCAUCCCCUACCCUGCA (((............................)))..(((....(((((.((((((((((...(((.(((((......))....))).)))..))))).))))))))))........))). (-19.61 = -20.37 + 0.75)



| Location | 6,749,617 – 6,749,737 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.34 |
| Mean single sequence MFE | -30.20 |
| Consensus MFE | -19.38 |
| Energy contribution | -19.16 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.27 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.553551 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6749617 120 - 27905053 AAAAAUAACACAAAGGCUUGUUCGUGUGAGUGUAUGUGUGUAUGUGUGUGCACAAAAGAGCACUAAAUACAUACAUUCCAGGCCCCAAAAACCACACUUCCGCGUCCCUUAGCCACGCAC ..............((((.(..((((..(((((((((((((((....((((........))))...))))))))))....((.........)))))))..))))..)...))))...... ( -36.60) >DroSec_CAF1 13858 110 - 1 AAAAAUAACACAAAGGCUUGUUCGUGUGAGUGUGU----------GUGUGCACAAAAGAGCACUAAAUACACACAUUCCAGGCCCCAAAACCCACACUCCCCCCACCCUUAGCCACGCAC ..............((((.....(((.((((((((----------((((((........)))).....))))))))))..((........))...........)))....))))...... ( -25.40) >DroSim_CAF1 18186 112 - 1 AAAAAUAACACAAAGGCUUGUUCGUGUGAGUGUGU--------GUGUGUGCACAAAAGAGCACUAAAUACACACAUUCCAGGCCCCAAAACCCACACUUCCCCCACCCUUAGCCACGCAC ..............((((.....(((.((((((((--------((((((((........))))...))))))))))))..((........))...........)))....))))...... ( -28.60) >consensus AAAAAUAACACAAAGGCUUGUUCGUGUGAGUGUGU________GUGUGUGCACAAAAGAGCACUAAAUACACACAUUCCAGGCCCCAAAACCCACACUUCCCCCACCCUUAGCCACGCAC ..............((((((...(((((.((((..............((((........))))...)))).)))))..)))))).................................... (-19.38 = -19.16 + -0.22)

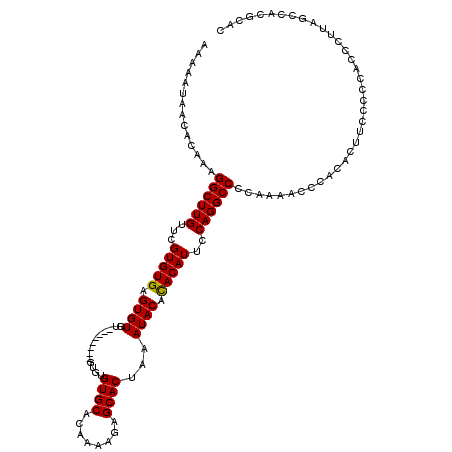

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:34:38 2006