
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 910,718 – 910,811 |
| Length | 93 |
| Max. P | 0.979143 |

| Location | 910,718 – 910,811 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 82.52 |
| Mean single sequence MFE | -54.13 |
| Consensus MFE | -45.24 |
| Energy contribution | -45.13 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.58 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.979143 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
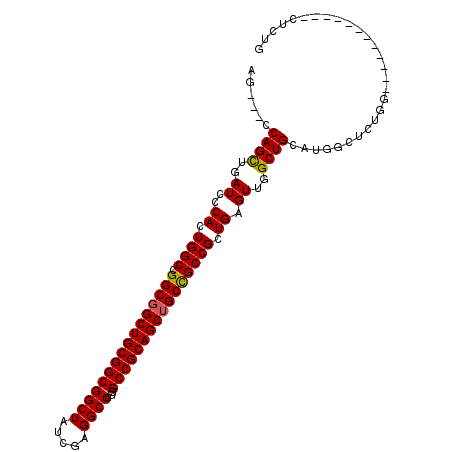
>3R_DroMel_CAF1 910718 93 + 27905053 AG---CCAGUUGAUCCCACUGGCCGGCGGCUGCGGCGGCUAUCGAGGCCUGGGCCGCAGCUGCCGCCGCUGAGUUGGCUGCAUGGCUCUGG------------CUCUG ((---((((...........((((((((((((((((((((.....))))...))))))))))))((.(((.....))).))..))))))))------------))... ( -51.40) >DroSim_CAF1 4907 87 + 1 AG---CCAGCUGAUCCCACUGGCCGGCGGCUGCGGCGGCUAUCGAGGCCUGUGCCGCAGCUGCUGCCGCUGAGUGGGCUGCAUGGCUCC------------------G ((---(((((....((((((((((((((((((((((((((.....))))...)))))))))))))..))).))))))..)).)))))..------------------. ( -53.80) >DroYak_CAF1 12751 108 + 1 AGCCGCCAGCUGAUCCCACUGGCCGGCGGCUGCGGCGGCUAUCGAGGCCUGGGCCGCAGCAGCCGCCGCUGAGUUGGCUGCAUGGCUCUGGCUCUGACUCUGGCUCUG (((((((.(((.........))).)))))))(((((((((.....(((....))).....))))))))).((((..(...((.(((....))).))...)..)))).. ( -57.20) >consensus AG___CCAGCUGAUCCCACUGGCCGGCGGCUGCGGCGGCUAUCGAGGCCUGGGCCGCAGCUGCCGCCGCUGAGUUGGCUGCAUGGCUCUGG____________CUCUG ......((((..((..((.((((.((((((((((((((((.....))))...)))))))))))))))).)).))..))))............................ (-45.24 = -45.13 + -0.11)


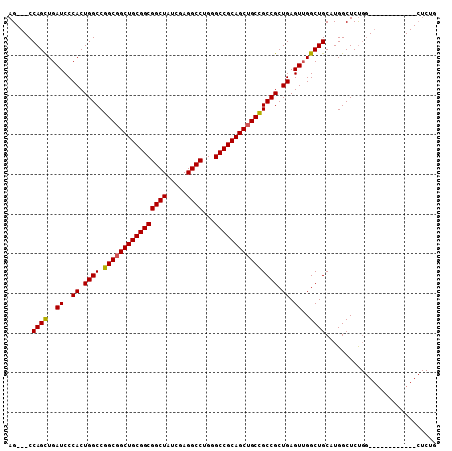
| Location | 910,718 – 910,811 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 82.52 |
| Mean single sequence MFE | -48.47 |
| Consensus MFE | -37.65 |
| Energy contribution | -37.77 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.892280 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
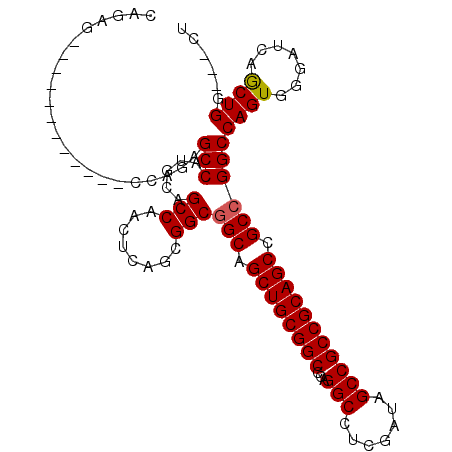
>3R_DroMel_CAF1 910718 93 - 27905053 CAGAG------------CCAGAGCCAUGCAGCCAACUCAGCGGCGGCAGCUGCGGCCCAGGCCUCGAUAGCCGCCGCAGCCGCCGGCCAGUGGGAUCAACUGG---CU ...((------------((((.((......))...((((..((((((.((((((((...(((.......))))))))))).))).)))..)))).....))))---)) ( -45.80) >DroSim_CAF1 4907 87 - 1 C------------------GGAGCCAUGCAGCCCACUCAGCGGCAGCAGCUGCGGCACAGGCCUCGAUAGCCGCCGCAGCCGCCGGCCAGUGGGAUCAGCUGG---CU .------------------..(((((.((..((((((....(((.((.((((((((...(((.......))))))))))).))..)))))))))....)))))---)) ( -46.30) >DroYak_CAF1 12751 108 - 1 CAGAGCCAGAGUCAGAGCCAGAGCCAUGCAGCCAACUCAGCGGCGGCUGCUGCGGCCCAGGCCUCGAUAGCCGCCGCAGCCGCCGGCCAGUGGGAUCAGCUGGCGGCU ...((((.((((..(...((......))...)..))))..((((((((((.(((((.............))))).))))))))))((((((.......)))))))))) ( -53.32) >consensus CAGAG____________CCAGAGCCAUGCAGCCAACUCAGCGGCGGCAGCUGCGGCCCAGGCCUCGAUAGCCGCCGCAGCCGCCGGCCAGUGGGAUCAGCUGG___CU ......................(((.....(((........)))(((.((((((((...(((.......))))))))))).))))))((((.......))))...... (-37.65 = -37.77 + 0.11)

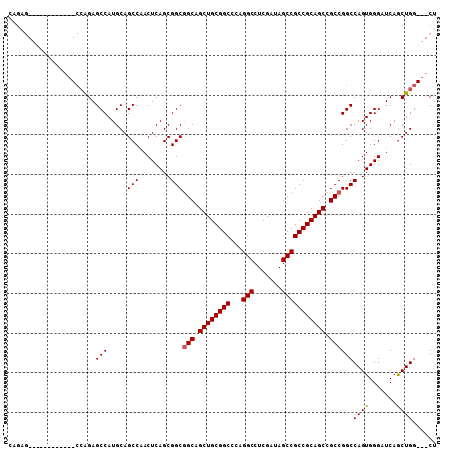
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:38:11 2006