
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 6,202,572 – 6,202,730 |
| Length | 158 |
| Max. P | 0.974413 |

| Location | 6,202,572 – 6,202,664 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 75.65 |
| Mean single sequence MFE | -20.80 |
| Consensus MFE | -12.22 |
| Energy contribution | -12.92 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.621767 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6202572 92 - 27905053 CUAUGUUCCUUCCGAAAUUUGGAGA-----------AGUGUACAGGACCAACGCCAACCUCCUGUGCACUUCCGCUUUGCUGCCUCGCCACUCGUCCUGCUUC ............(((....(((.((-----------(((((((((((............))))))))))))).((......))....))).)))......... ( -24.00) >DroSec_CAF1 22992 81 - 1 CUAUGUUCCUUACAAAAUUUGGAAA-----------AGUGUACAGGACC------AACCUCCUGUGCACUUCCGCUUUGCUGC-----CACUCGUCCUGCUUC ...(((.....)))......(((..-----------(((((((((((..------....))))))))))))))((...((...-----.....))...))... ( -21.70) >DroSim_CAF1 23425 92 - 1 CUAUGUUCCUUCCAAAAUUUGGAAA-----------AGUGUACAGGACCAACGCCAACCUCCUGUGCACUUCCGCUUUGCUGCCUCGCCACUCGUCCUGCUUC .........(((((.....)))))(-----------(((((((((((............))))))))))))..((......)).................... ( -21.20) >DroEre_CAF1 23590 85 - 1 -------UAUUUAGAAAUUUAGAGA-----------AGUGUACCGGCCCAAUGGCAACCUCCUGUGCACUUCCGCUUUGCUGCCUCGCCACUCGUCCUGCUUC -------......((......(.((-----------(((((((.((......(....)..)).))))))))))((......)).))................. ( -15.90) >DroYak_CAF1 20156 86 - 1 ------GCCUUUAGAAAUGUAGAGA-----------AGUGUACAGGACCAACGACAACCUCCUGUGCACUUCCGCUUUGCUGCCUCGCCGCUCGUCCUGCUUC ------............((((.((-----------(((((((((((............)))))))))))))......((.((...)).)).....))))... ( -25.30) >DroAna_CAF1 29551 95 - 1 AAGAAUUGUUUUCAAAACUUACAAAAUACGCAGUUCAGUGUACAG--------CCAUCCUCCCGUGCAUUUCCGCUUUGCUGCCUCGCAACGUGUCCUGCUUC ..(((.....)))................((((...(((((((..--------..........)))))))..(((.((((......)))).)))..))))... ( -16.70) >consensus CUAUGUUCCUUUCAAAAUUUAGAAA___________AGUGUACAGGACCAACGCCAACCUCCUGUGCACUUCCGCUUUGCUGCCUCGCCACUCGUCCUGCUUC ....................................(((((((((((............)))))))))))...((......)).................... (-12.22 = -12.92 + 0.70)


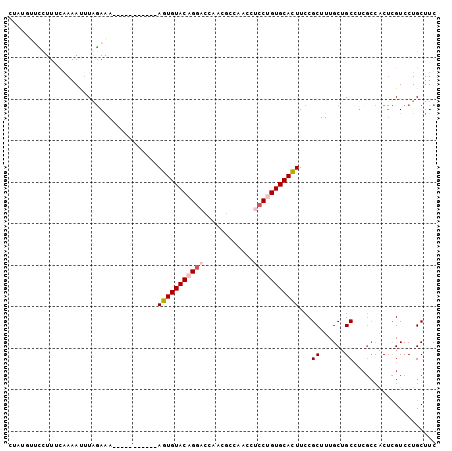
| Location | 6,202,601 – 6,202,704 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 71.10 |
| Mean single sequence MFE | -22.16 |
| Consensus MFE | -14.25 |
| Energy contribution | -13.88 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.64 |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.908608 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6202601 103 - 27905053 AUGCACAUCAAUUCAUUUGAAAUUAGGUUUCGUAAGCUUUCUAUGUUCCUUCCGAAAUUUGGAGA-----------AGUGUACAGGACCAACGCCAACCUCCUGUGCACUUCCG .......((((.....))))..((((((((((.(((............))).)))))))))).((-----------(((((((((((............))))))))))))).. ( -28.40) >DroSec_CAF1 23016 97 - 1 AUGCGCAUCAAUCCAUUUGAAAUUAGGCUUCGUAAGCUAUCUAUGUUCCUUACAAAAUUUGGAAA-----------AGUGUACAGGACC------AACCUCCUGUGCACUUCCG ...........((((..(......(((...((((.......))))..)))......)..)))).(-----------(((((((((((..------....))))))))))))... ( -25.60) >DroSim_CAF1 23454 103 - 1 AUGCGCAUCAAUCCAUUUGAAAUUAGGCUUCGUAAGCUAUCUAUGUUCCUUCCAAAAUUUGGAAA-----------AGUGUACAGGACCAACGCCAACCUCCUGUGCACUUCCG ....(((((((.....)))).....((((.....)))).....)))...(((((.....)))))(-----------(((((((((((............))))))))))))... ( -24.70) >DroAna_CAF1 29580 103 - 1 ---CGAAUCAAAACUUUUUAAAUUGAAGAACAAAUACAAGAAGAAUUGUUUUCAAAACUUACAAAAUACGCAGUUCAGUGUACAG--------CCAUCCUCCCGUGCAUUUCCG ---..........((((.......))))..............(((((((....................)))))))(((((((..--------..........))))))).... ( -9.95) >consensus AUGCGCAUCAAUCCAUUUGAAAUUAGGCUUCGUAAGCUAUCUAUGUUCCUUCCAAAAUUUGGAAA___________AGUGUACAGGACC____CCAACCUCCUGUGCACUUCCG .............................................((((...........))))............(((((((((((............))))))))))).... (-14.25 = -13.88 + -0.38)



| Location | 6,202,635 – 6,202,730 |
|---|---|
| Length | 95 |
| Sequences | 3 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 93.68 |
| Mean single sequence MFE | -21.44 |
| Consensus MFE | -20.55 |
| Energy contribution | -20.44 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.96 |
| SVM decision value | 1.73 |
| SVM RNA-class probability | 0.974413 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 6202635 95 - 27905053 UUCACUUUUCCUUUCAAAACCAAGGAAUGCACAUCAAUUCAUUUGAAAUUAGGUUUCGUAAGCUUUCUAUGUUCCUUCCGAAAUUUGGAGAAGUG ..(((((((((..........(((((((.....((((.....))))...((((...(....)...)))).))))))).........))))))))) ( -21.41) >DroSec_CAF1 23044 95 - 1 UUAACUUUUCCUUUCAAAACCAAGGAAUGCGCAUCAAUCCAUUUGAAAUUAGGCUUCGUAAGCUAUCUAUGUUCCUUACAAAAUUUGGAAAAGUG ...((((((((..........(((((((.....((((.....)))).....((((.....))))......))))))).........)))))))). ( -20.61) >DroSim_CAF1 23488 95 - 1 UUCACUUUUCCUUUUAAAACCAAGGAAUGCGCAUCAAUCCAUUUGAAAUUAGGCUUCGUAAGCUAUCUAUGUUCCUUCCAAAAUUUGGAAAAGUG ..(((((((((..........(((((((.....((((.....)))).....((((.....))))......))))))).........))))))))) ( -22.31) >consensus UUCACUUUUCCUUUCAAAACCAAGGAAUGCGCAUCAAUCCAUUUGAAAUUAGGCUUCGUAAGCUAUCUAUGUUCCUUCCAAAAUUUGGAAAAGUG ..(((((((((..........(((((((.....((((.....)))).....((((.....))))......))))))).........))))))))) (-20.55 = -20.44 + -0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:28:48 2006