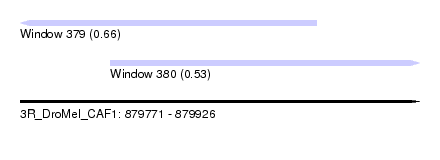

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 879,771 – 879,926 |
| Length | 155 |
| Max. P | 0.660067 |

| Location | 879,771 – 879,886 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.09 |
| Mean single sequence MFE | -38.14 |
| Consensus MFE | -29.79 |
| Energy contribution | -29.85 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.660067 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
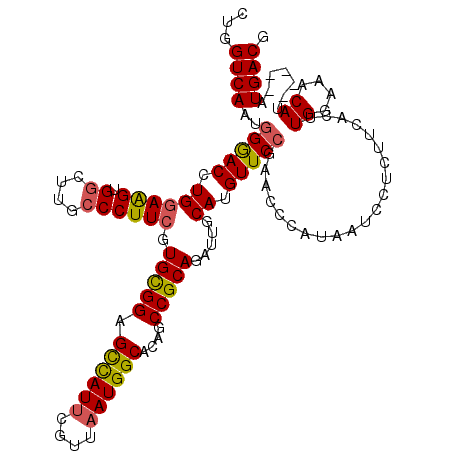
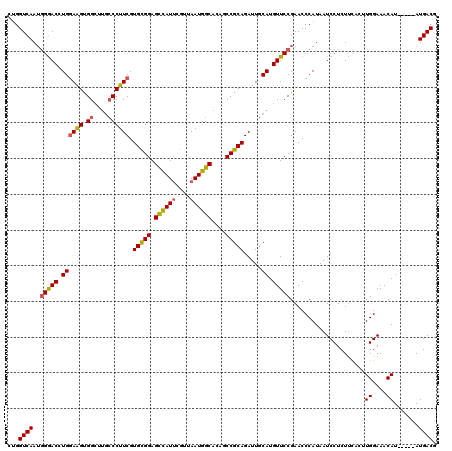
>3R_DroMel_CAF1 879771 115 - 27905053 CUGGUCAAAGGAACCUGGAGGCGGCGUGCCCUUCGUGCGGAGCCAUUCGUUAAUGGCAGAUCCGCAGAUUGCAUGUUCCGAACCCAUAAUCCUCUUCACUUGGAAACAU-----AUGACG ...((((.((((...(((...((((((((...((.((((((((((((....))))))...))))))))..)))))..)))...)))...)))).......((....)).-----.)))). ( -40.10) >DroSec_CAF1 47780 115 - 1 CUGGUCAAUGGGACCUGGAAGUGGCUUGCCCUUCGUGCGGAGCCAUUUGUUAAUGGCACAGCCGCAGAUUGCAUGUUCCGAACCCAUAAUCCUCUUCACUUGGAAACAU-----AUGACG ..((...(((((...((((....((.(((...((.(((((.((((((....))))))....)))))))..))).))))))..)))))...))...(((..((....)).-----.))).. ( -37.30) >DroSim_CAF1 46980 115 - 1 CUGGUCAAUGGGACCUGCAAGUGGCUUGCCCUUCGUGCGGAGCCAUUCGCUAAUGGCACAGCCGCAGAUUGCAUGUUCCGAACCCAUAAUCCUCUUCACUUGGAAACAU-----AUGACG ..(((...((((((.(((((((((((((((......(((((....)))))....)))).))))))...))))).)))))).)))...........(((..((....)).-----.))).. ( -40.70) >DroEre_CAF1 46884 116 - 1 CAGGUCAAUAGGACCUGGAAGUGG----ACCUUCGUGUGGAGUUAUCCGUUAAUGGCAGAGCCGCAGAUUGCAUGUUCCGAACCCAUAAUCCUCUCUACUUGGAAACAUGUAGUAUGACG ((((((.....))))))...((((----..((((((.((((....))))...)))).))..))))..((((((((((((((..................)))).))))))))))...... ( -34.47) >consensus CUGGUCAAUGGGACCUGGAAGUGGCUUGCCCUUCGUGCGGAGCCAUUCGUUAAUGGCACAGCCGCAGAUUGCAUGUUCCGAACCCAUAAUCCUCUUCACUUGGAAACAU_____AUGACG ...((((..(((((.((((((.((....)))))).(((((.((((((....))))))....))))).....)).))))).....................((....)).......)))). (-29.79 = -29.85 + 0.06)

| Location | 879,806 – 879,926 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.29 |
| Mean single sequence MFE | -34.65 |
| Consensus MFE | -22.84 |
| Energy contribution | -24.40 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.531852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
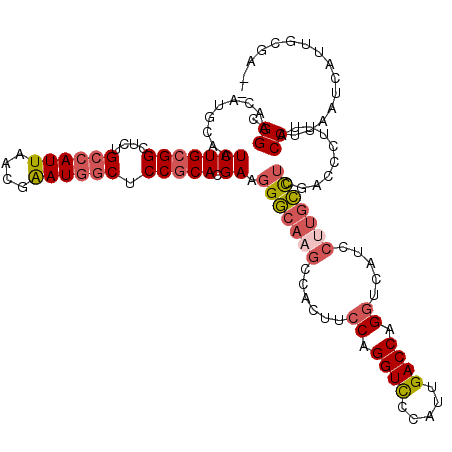
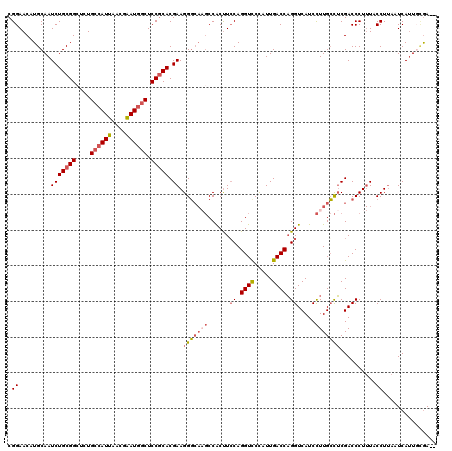
>3R_DroMel_CAF1 879806 120 + 27905053 CGGAACAUGCAAUCUGCGGAUCUGCCAUUAACGAAUGGCUCCGCACGAAGGGCACGCCGCCUCCAGGUUCCUUUGACCAGGUCAUCCUUGCCUCGACCCUUUACCUUAAUCAUUUUAAUC .(((((.((...((((((((...((((((....)))))))))))).)).((((.....)))).)).)))))........((((...........))))...................... ( -32.80) >DroSec_CAF1 47815 118 + 1 CGGAACAUGCAAUCUGCGGCUGUGCCAUUAACAAAUGGCUCCGCACGAAGGGCAAGCCACUUCCAGGUCCCAUUGACCAGGUCAUCCUUGCCUCGACCCUUUACCUUAAUCAUUGCGA-- .......((((((.(((((....((((((....)))))).)))))(((..((((((......((.((((.....)))).)).....)))))))))................)))))).-- ( -37.00) >DroSim_CAF1 47015 118 + 1 CGGAACAUGCAAUCUGCGGCUGUGCCAUUAGCGAAUGGCUCCGCACGAAGGGCAAGCCACUUGCAGGUCCCAUUGACCAGGUCAUCCUUGCCUCGACCCUUUACCUUAAUCAUUGCGA-- .......((((((.(((((....((((((....)))))).)))))..(((((((((...))))).((((.....)))).((((...........)))).....))))....)))))).-- ( -36.30) >DroEre_CAF1 46924 112 + 1 CGGAACAUGCAAUCUGCGGCUCUGCCAUUAACGGAUAACUCCACACGAAGGU----CCACUUCCAGGUCCUAUUGACCUGGUCAUCUUUGGUGCGACCCU-UACCUGAGUCAGUU-GA-- ...(((..((.....))(((((..........(((....))).......(((----(((((.(((((((.....)))))))........)))).))))..-.....))))).)))-..-- ( -32.50) >consensus CGGAACAUGCAAUCUGCGGCUCUGCCAUUAACGAAUGGCUCCGCACGAAGGGCAAGCCACUUCCAGGUCCCAUUGACCAGGUCAUCCUUGCCUCGACCCUUUACCUUAAUCAUUGCGA__ .((.........(((((((....((((((....)))))).))))).)).(((((((......((.((((.....)))).)).....)))))))..........))............... (-22.84 = -24.40 + 1.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:37:43 2006