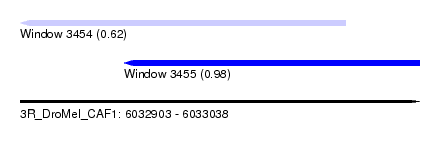
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 6,032,903 – 6,033,038 |
| Length | 135 |
| Max. P | 0.978744 |

| Location | 6,032,903 – 6,033,013 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 78.92 |
| Mean single sequence MFE | -26.78 |
| Consensus MFE | -14.82 |
| Energy contribution | -15.26 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.623106 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
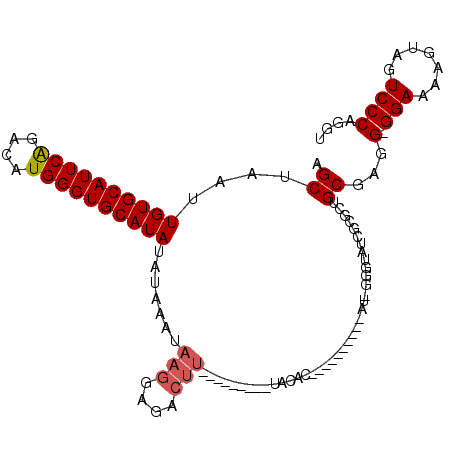
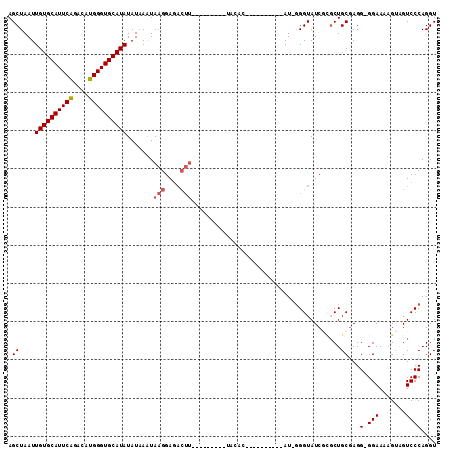
>3R_DroMel_CAF1 6032903 110 - 27905053 AGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUUAGUAAGGAGACUUUUUUAUGUAUACACACACACACAGAU-GGGUAUCGCUCUGCGAGG-GGAAAAGUAGUCCCAGGU .((((((((((((((((....)))))))))).))))))..(((.((((((((.....(((...((........)-).)))((((...))))..-))))))))..)))..... ( -28.80) >DroSec_CAF1 38932 86 - 1 AGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUU------------------------AU-GGGUAUCGCGCUGCGAGG-GGAAAAGUCGUCCCAGGU .......((((((((((....))))))))))(((..(((((....)))------------------------))-..)))((((...)))).(-(((.......)))).... ( -27.10) >DroSim_CAF1 39250 87 - 1 AGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUU------------------------AU-GGGUAUCGCGCUGCAAGGGGGAAAAGUCGUCCCAGGU (((....((((((((((....))))))))))(((..(((((....)))------------------------))-..)))....)))....((((((....)).)))).... ( -26.90) >DroEre_CAF1 39316 84 - 1 AGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUACGGA--------------UACAC----------AU-GGGUAUC--GCUGCGAGG-GGAAAAGGAGUCCCAGGU (((....((((((((((....))))))))))..........((--------------(((..----------..-..)))))--))).....(-(((.......)))).... ( -23.70) >DroYak_CAF1 40888 100 - 1 AGCUAAUUGUGCAUUCGGACAUGGGUGCAUAUAUAAAUAAGGAGACUUAUCUGCAUAUACAC----------AUUUGGUAUC-UGCUGCGAGG-GGAAACGGAGUCCCAGGU .((....((((((((((....)))))))))).....(((((....)))))..(((.((((.(----------....))))).-))).))...(-(((.......)))).... ( -27.40) >consensus AGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUU_________UACAC__________AU_GGGUAUCGCGCUGCGAGG_GGAAAAGUAGUCCCAGGU .((....((((((((((....)))))))))).......(((....))).......................................))...(.(((.......)))).... (-14.82 = -15.26 + 0.44)

| Location | 6,032,938 – 6,033,038 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 76.92 |
| Mean single sequence MFE | -22.45 |
| Consensus MFE | -12.98 |
| Energy contribution | -13.23 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.58 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978744 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
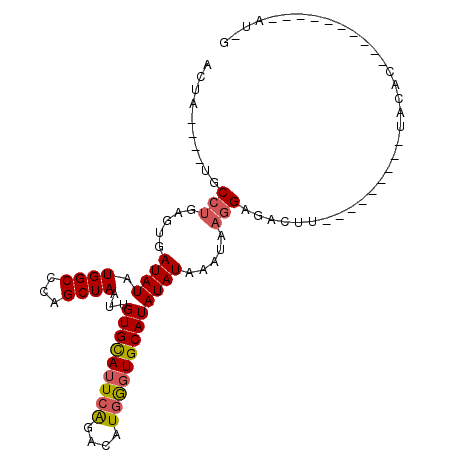
>3R_DroMel_CAF1 6032938 100 - 27905053 ACUA----UGCCUGAGUGAUAUAUGGCCCAGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUUAGUAAGGAGACUUUUUUAUGUAUACACACACACACAGAU-G .((.----((..((.(((((((((((....((((((((((((((((....)))))))))).))))))(((....)))..)))))))).))).))..)).))..-. ( -29.20) >DroSec_CAF1 38967 76 - 1 ACUA----UGCCUGAGUGAUAUAUGGCCCAGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUU------------------------AU-G .(((----((.((((((((((..((((...))))..))).))))))).)))))............(((((....)))------------------------))-. ( -23.20) >DroSim_CAF1 39286 76 - 1 ACUA----UGCCUGAGUGAUAUAUGGCCCAGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUU------------------------AU-G .(((----((.((((((((((..((((...))))..))).))))))).)))))............(((((....)))------------------------))-. ( -23.20) >DroEre_CAF1 39349 76 - 1 ACUA----UGCCUGAGUGAUAUAUGGCCCAGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUACGGA--------------UACAC----------AU-G .(((----((.((((((((((..((((...))))..))).))))))).)))))...................--------------.....----------..-. ( -17.50) >DroYak_CAF1 40922 91 - 1 ACUA----UGCCUGAGUGAUAUAUGGCCAAGCUAAUUGUGCAUUCGGACAUGGGUGCAUAUAUAAAUAAGGAGACUUAUCUGCAUAUACAC----------AUUU .(((----((.((((((((((..((((...))))..))).))))))).)))))(((.((((.((.(((((....))))).)).)))).)))----------.... ( -28.00) >DroPer_CAF1 36720 82 - 1 ACUACAUUUGCCUGCCUGAUAUAUGGCCACGCUAAUUGUGUAUUCAGAUAUCACUGCAUAUAUAAAUAAAGAC------------AUAGAC----------AU-G .(((.....((..(((........)))...))...((((((((.(((......))).))))))))........------------.)))..----------..-. ( -13.60) >consensus ACUA____UGCCUGAGUGAUAUAUGGCCCAGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUU_________UACAC__________AU_G ..........(((.....((((.((((...))))...(((((((((....))))))))))))).....))).................................. (-12.98 = -13.23 + 0.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:27:44 2006