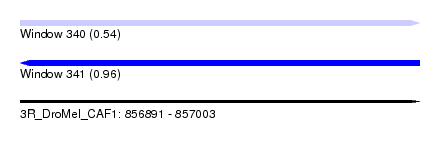

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 856,891 – 857,003 |
| Length | 112 |
| Max. P | 0.962945 |

| Location | 856,891 – 857,003 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.19 |
| Mean single sequence MFE | -42.70 |
| Consensus MFE | -30.58 |
| Energy contribution | -32.83 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.542699 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
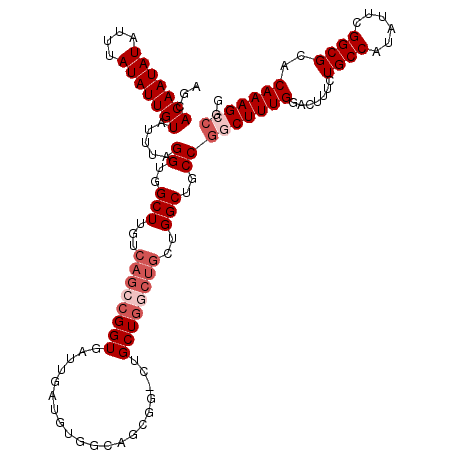
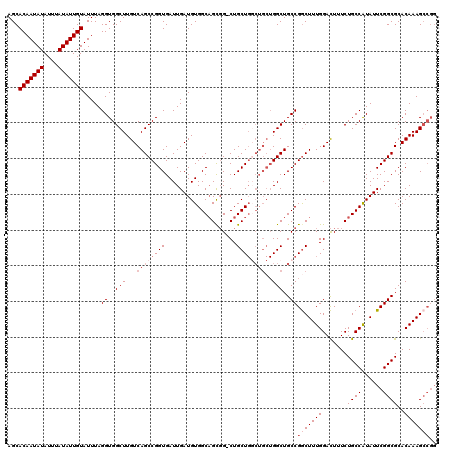
>3R_DroMel_CAF1 856891 112 + 27905053 AGCACAAUAUAUUUAUAUUGUAUUUAGGUGGCUUGUCAGCCGGUGAUUGAUGUGGCAGCGG-CUGCU-------GGCUGCCGGCUUUGGACUUUCUGCCAUAUUCGGCGCACAAAGCCGG ...(((((((....))))))).....((..(((.(.((((((.((..........)).)))-)))).-------)))..))(((((((.......((((......))))..))))))).. ( -43.30) >DroSec_CAF1 31701 115 + 1 AGCACAAUAUAUUUAUAUUGUAUUUAGGUGGCUUGUCAGCCGGUGAUUGAUGUGGCAGCGG-CUGCUGGCUGCUGGCUGCCGGCUUUGGACUUUCUGCCAUAUUCGGCGCACAAAG---- .(((((((((....)))))))......(((((..(((((((((((........(.((((((-(.....))))))).)))))))))...))).....))))).......))......---- ( -40.50) >DroSim_CAF1 30859 119 + 1 AGCACAAUAUAUUUAUAUUGUAUUUAGGUGGCUUGUCAGCCGGUGAUUGAUGUGGCAGUGG-CUGCUGGCUGCUGGCUGCCGGCUUUGGACUUUCUGCCAUAUUCGGCGCACAAAGCCGG ((((((((((....))))))).....((..(((.(.((((((.((..........)).)))-)))).)))..)).))).(((((((((.......((((......))))..))))))))) ( -45.80) >DroEre_CAF1 31927 120 + 1 AGCACAAUAUAUUUAUAUUGUAUUUAGGUGGCUUGUCAGCCGGUGAUUGAUGUGGCUGCUGCCUGCUGCCUGCUGGCUGCCGACUUUGGAUUUUCUGCCGUAUUCGGCGCACAAAGCCGG ((((((((((....)))))))....(((..((..(.((((.(((.((....)).))))))))..))..))))))(((((((((...(((........)))...)))))......)))).. ( -41.20) >consensus AGCACAAUAUAUUUAUAUUGUAUUUAGGUGGCUUGUCAGCCGGUGAUUGAUGUGGCAGCGG_CUGCUGGCUGCUGGCUGCCGGCUUUGGACUUUCUGCCAUAUUCGGCGCACAAAGCCGG ...(((((((....))))))).....((..(((...((((((((....................))))))))..)))..))(((((((.......((((......))))..))))))).. (-30.58 = -32.83 + 2.25)

| Location | 856,891 – 857,003 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.19 |
| Mean single sequence MFE | -36.47 |
| Consensus MFE | -29.90 |
| Energy contribution | -30.84 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.95 |
| Structure conservation index | 0.82 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.962945 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
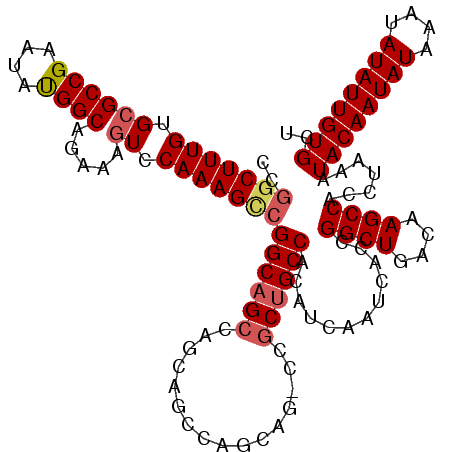
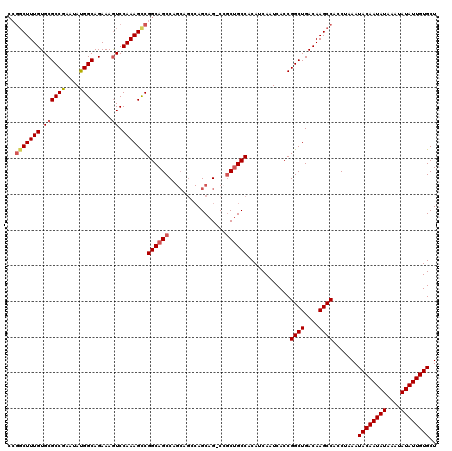
>3R_DroMel_CAF1 856891 112 - 27905053 CCGGCUUUGUGCGCCGAAUAUGGCAGAAAGUCCAAAGCCGGCAGCC-------AGCAG-CCGCUGCCACAUCAAUCACCGGCUGACAAGCCACCUAAAUACAAUAUAAAUAUAUUGUGCU ..(((((((..((((......))).....)..)))))))((((((.-------.....-..))))))............((((....)))).......((((((((....)))))))).. ( -38.10) >DroSec_CAF1 31701 115 - 1 ----CUUUGUGCGCCGAAUAUGGCAGAAAGUCCAAAGCCGGCAGCCAGCAGCCAGCAG-CCGCUGCCACAUCAAUCACCGGCUGACAAGCCACCUAAAUACAAUAUAAAUAUAUUGUGCU ----(((((..((((......))).....)..)))))..((((((..((........)-).))))))............((((....)))).......((((((((....)))))))).. ( -31.60) >DroSim_CAF1 30859 119 - 1 CCGGCUUUGUGCGCCGAAUAUGGCAGAAAGUCCAAAGCCGGCAGCCAGCAGCCAGCAG-CCACUGCCACAUCAAUCACCGGCUGACAAGCCACCUAAAUACAAUAUAAAUAUAUUGUGCU ..(((((((..((((......))).....)..)))))))(((((...((........)-)..)))))............((((....)))).......((((((((....)))))))).. ( -36.40) >DroEre_CAF1 31927 120 - 1 CCGGCUUUGUGCGCCGAAUACGGCAGAAAAUCCAAAGUCGGCAGCCAGCAGGCAGCAGGCAGCAGCCACAUCAAUCACCGGCUGACAAGCCACCUAAAUACAAUAUAAAUAUAUUGUGCU (((((((((...((((....)))).(.....))))))))))..(((....)))(((.(((..(((((............)))))....)))........(((((((....)))))))))) ( -39.80) >consensus CCGGCUUUGUGCGCCGAAUAUGGCAGAAAGUCCAAAGCCGGCAGCCAGCAGCCAGCAG_CCGCUGCCACAUCAAUCACCGGCUGACAAGCCACCUAAAUACAAUAUAAAUAUAUUGUGCU ..(((((((.((((((....)))).....)).)))))))((((((................))))))............((((....)))).......((((((((....)))))))).. (-29.90 = -30.84 + 0.94)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:37:03 2006