
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 5,492,943 – 5,493,063 |
| Length | 120 |
| Max. P | 0.824300 |

| Location | 5,492,943 – 5,493,063 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.39 |
| Mean single sequence MFE | -35.15 |
| Consensus MFE | -31.21 |
| Energy contribution | -31.10 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.809065 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5492943 120 + 27905053 UCCAAUAUGUUCUUGAGCGAGAAACCCUCCGAGUCGAACUCGAAGAUGUAGCGAGUAACAUCCAUGAUCCGUUGCCAGUGGCGUGGCUUCAUGGCUUUAUUCGCCAUCAACUCCAGUAGC ........(((((.(((.(((.....)))((((.....))))..((((..((((((((...((((((.((((.(((...)))))))..))))))..))))))))))))..))).)).))) ( -34.20) >DroVir_CAF1 33989 120 + 1 UCCAGUAUGUUCUUGAGCGAGAAACCCUCCGAGUCGAACUCGAAGAUGUAGCGAGUAACGUCCAUGAUCCGCUGCCAAUGGCGUGGCUUCAUGGCUUUAUUCGCCAUCAACUCCAAUAAC ........(((....)))(((........((((.....))))..((((..((((((((.(.((((((.((((.(((...)))))))..))))))).))))))))))))..)))....... ( -38.90) >DroGri_CAF1 27853 120 + 1 UCCAGUAUGUUCUUGAGCGAGAAACCCUCCGAGUCGAACUCGAAGAUGUAGCGAGUAACAUCCAUGAUCCGUUGCCAAUGGCGUGGCUUCAUGGCUUUAUUCGCCAUCAACUCCAGUAAU ...........((.(((.(((.....)))((((.....))))..((((..((((((((...((((((.((((.(((...)))))))..))))))..))))))))))))..))).)).... ( -33.20) >DroWil_CAF1 26000 120 + 1 UCCAAAAUGUUCUUGAGCGAGAAACCCUCGGAAUCAAAUUCAAAGAUGUAGCGAGUAACGUCCAUGAUCCGUUGCCAAUGGCGUGGCUUCAUGGCUUUAUUCGCCAUCAACUCCAGGAGC ........(((((((((((((.....))))..............((((..((((((((.(.((((((.((((.(((...)))))))..))))))).))))))))))))..)).))))))) ( -38.10) >DroMoj_CAF1 33637 120 + 1 UCCAGUAUGUUCUUAAGCGAGAAACCCUCCGAGUCGAACUCGAAGAUGUAGCGAGUAACAUCCAUGAUCCGUUGCCAAUGGCGUGGCUUCAUGGCUUUAUUCGCCAUCAACUCCAAUAGU ................(.(((........((((.....))))..((((..((((((((...((((((.((((.(((...)))))))..))))))..))))))))))))..))))...... ( -32.50) >DroAna_CAF1 26165 120 + 1 UCCAAUAUGUUCUUGAGCGAGAAACCCUCCGAGUCGAACUCGAAGAUGUAGCGAGUAACAUCCAUGAUUCGUUGCCAAUGACGUGGCUUCAUGGCUUUAUUCGCCAUCAACUCCAGUAGC ........(((((.(((.(((.....)))((((.....))))..((((..((((((((...((((((......((((......)))).))))))..))))))))))))..))).)).))) ( -34.00) >consensus UCCAAUAUGUUCUUGAGCGAGAAACCCUCCGAGUCGAACUCGAAGAUGUAGCGAGUAACAUCCAUGAUCCGUUGCCAAUGGCGUGGCUUCAUGGCUUUAUUCGCCAUCAACUCCAGUAGC ................(.(((........((((.....))))..((((..((((((((...((((((......((((......)))).))))))..))))))))))))..))))...... (-31.21 = -31.10 + -0.11)

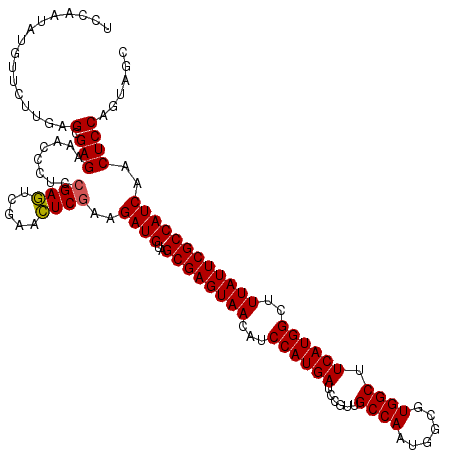

| Location | 5,492,943 – 5,493,063 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.39 |
| Mean single sequence MFE | -36.92 |
| Consensus MFE | -34.30 |
| Energy contribution | -33.83 |
| Covariance contribution | -0.47 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.824300 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5492943 120 - 27905053 GCUACUGGAGUUGAUGGCGAAUAAAGCCAUGAAGCCACGCCACUGGCAACGGAUCAUGGAUGUUACUCGCUACAUCUUCGAGUUCGACUCGGAGGGUUUCUCGCUCAAGAACAUAUUGGA ....((.((((((.((((((.(((..((((((..((..(((...)))...)).))))))...))).))))))))((((((((.....)))))))).......)))).))........... ( -37.50) >DroVir_CAF1 33989 120 - 1 GUUAUUGGAGUUGAUGGCGAAUAAAGCCAUGAAGCCACGCCAUUGGCAGCGGAUCAUGGACGUUACUCGCUACAUCUUCGAGUUCGACUCGGAGGGUUUCUCGCUCAAGAACAUACUGGA (((....((((((.((((((.(((..((((((..((.((((...))).).)).))))))...))).))))))))((((((((.....)))))))).......))))...)))........ ( -39.80) >DroGri_CAF1 27853 120 - 1 AUUACUGGAGUUGAUGGCGAAUAAAGCCAUGAAGCCACGCCAUUGGCAACGGAUCAUGGAUGUUACUCGCUACAUCUUCGAGUUCGACUCGGAGGGUUUCUCGCUCAAGAACAUACUGGA ....((.((((((.((((((.(((..((((((..((..(((...)))...)).))))))...))).))))))))((((((((.....)))))))).......)))).))........... ( -37.50) >DroWil_CAF1 26000 120 - 1 GCUCCUGGAGUUGAUGGCGAAUAAAGCCAUGAAGCCACGCCAUUGGCAACGGAUCAUGGACGUUACUCGCUACAUCUUUGAAUUUGAUUCCGAGGGUUUCUCGCUCAAGAACAUUUUGGA ..(((.(((((..(((((((.(((..((((((..((..(((...)))...)).))))))...))).))))))((....))...)..)))))(((.(.....).)))...........))) ( -35.10) >DroMoj_CAF1 33637 120 - 1 ACUAUUGGAGUUGAUGGCGAAUAAAGCCAUGAAGCCACGCCAUUGGCAACGGAUCAUGGAUGUUACUCGCUACAUCUUCGAGUUCGACUCGGAGGGUUUCUCGCUUAAGAACAUACUGGA .(((((((((.((.((((((.(((..((((((..((..(((...)))...)).))))))...))).)))))))).))))))((((......(((.....)))......))))....))). ( -35.50) >DroAna_CAF1 26165 120 - 1 GCUACUGGAGUUGAUGGCGAAUAAAGCCAUGAAGCCACGUCAUUGGCAACGAAUCAUGGAUGUUACUCGCUACAUCUUCGAGUUCGACUCGGAGGGUUUCUCGCUCAAGAACAUAUUGGA ....((.((((((.((((((.(((..((((((.((((......))))......))))))...))).))))))))((((((((.....)))))))).......)))).))........... ( -36.10) >consensus GCUACUGGAGUUGAUGGCGAAUAAAGCCAUGAAGCCACGCCAUUGGCAACGGAUCAUGGAUGUUACUCGCUACAUCUUCGAGUUCGACUCGGAGGGUUUCUCGCUCAAGAACAUACUGGA ....((.((((((.((((((.(((..((((((.((((......))))......))))))...))).))))))))((((((((.....)))))))).......)))).))........... (-34.30 = -33.83 + -0.47)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:23:34 2006