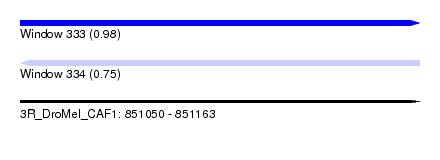

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 851,050 – 851,163 |
| Length | 113 |
| Max. P | 0.982805 |

| Location | 851,050 – 851,163 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.41 |
| Mean single sequence MFE | -45.03 |
| Consensus MFE | -34.50 |
| Energy contribution | -35.56 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.93 |
| SVM RNA-class probability | 0.982805 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 851050 113 + 27905053 UGGGCUGGGGGCUUAGGACUUGG-------GACUUGGGCCUUGGGAUUUUGGAGUGAAUGGGAUAAGCUCACGCACUGCCACGCCCCCGGCCCCGAAAAUGUUAUUUUAACUACAUUUCG .(((((((((((......((..(-------(........))..))....(((((((..((((.....))))..)))).))).)))))))))))(((((.(((..........)))))))) ( -46.70) >DroSec_CAF1 26723 113 + 1 CGGGCUGGGGGCUUAGGACUUGG-------GACUUGGGCCUUGGGACUUCGGAGUGAAUGGGCUAAGCUCACGCACUGCCACGCCCCCGGCCCCAAAAAUGUUAUUUUAACUACAUUUCG .(((((((((((..((..((.((-------(.(....)))).))..))..((((((..(((((...)))))..)))).))..)))))))))))...((((((..........)))))).. ( -48.80) >DroSim_CAF1 25944 120 + 1 CGGGCUGGGGGCUUAGGACUUGGGACUUGGGACUUGGGCCUUGGGACUUCGGAGUGAAUGGGCUAAGCUCACGCACUGCCACGCCCCCGGCCCCAAAAAUGUUAUUUUAACUACAUUUCG .(((((((((((..((..((.(((.((..(...)..))))).))..))..((((((..(((((...)))))..)))).))..)))))))))))...((((((..........)))))).. ( -51.40) >DroEre_CAF1 27511 91 + 1 UGGGCUGGG----------------------------GCAUUGGGAC-UCGAAGUGAAUGGGCUAAGCUCACGCACUGCCACGCCCCCGGCCCCAAAAAUGUUAUUUUAACUACAUUUCG .((((((((----------------------------(.....((..-....((((..(((((...)))))..)))).))....)))))))))...((((((..........)))))).. ( -33.20) >consensus CGGGCUGGGGGCUUAGGACUUGG_______GACUUGGGCCUUGGGACUUCGGAGUGAAUGGGCUAAGCUCACGCACUGCCACGCCCCCGGCCCCAAAAAUGUUAUUUUAACUACAUUUCG .((((((((((((((((...............))))))))).(((.....(.((((..((((.....))))..)))).)....))))))))))...((((((..........)))))).. (-34.50 = -35.56 + 1.06)

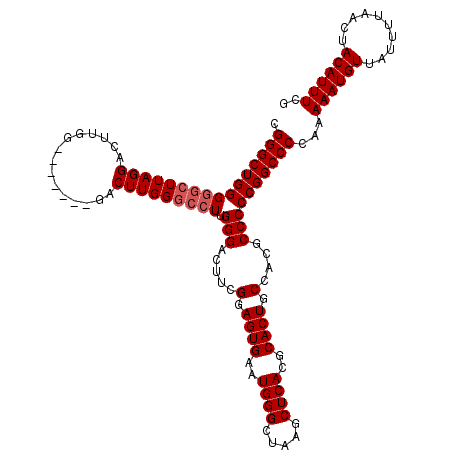
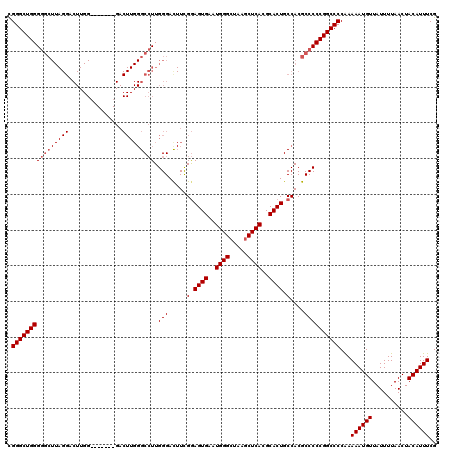
| Location | 851,050 – 851,163 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.41 |
| Mean single sequence MFE | -37.25 |
| Consensus MFE | -30.77 |
| Energy contribution | -31.90 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.746082 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 851050 113 - 27905053 CGAAAUGUAGUUAAAAUAACAUUUUCGGGGCCGGGGGCGUGGCAGUGCGUGAGCUUAUCCCAUUCACUCCAAAAUCCCAAGGCCCAAGUC-------CCAAGUCCUAAGCCCCCAGCCCA ((((((((.((....)).))).)))))((((.((((((.(((.((((.(((.........))).))))))).........(((....)))-------...........)))))).)))). ( -37.90) >DroSec_CAF1 26723 113 - 1 CGAAAUGUAGUUAAAAUAACAUUUUUGGGGCCGGGGGCGUGGCAGUGCGUGAGCUUAGCCCAUUCACUCCGAAGUCCCAAGGCCCAAGUC-------CCAAGUCCUAAGCCCCCAGCCCG .(((((((.((....)).)))))))..((((.((((((.(((.((((.(((.((...)).))).)))))))..(((....))).......-------...........)))))).)))). ( -38.40) >DroSim_CAF1 25944 120 - 1 CGAAAUGUAGUUAAAAUAACAUUUUUGGGGCCGGGGGCGUGGCAGUGCGUGAGCUUAGCCCAUUCACUCCGAAGUCCCAAGGCCCAAGUCCCAAGUCCCAAGUCCUAAGCCCCCAGCCCG .(((((((.((....)).)))))))..((((.((((((.(((.((((.(((.((...)).))).)))))))..(((....))).........................)))))).)))). ( -38.40) >DroEre_CAF1 27511 91 - 1 CGAAAUGUAGUUAAAAUAACAUUUUUGGGGCCGGGGGCGUGGCAGUGCGUGAGCUUAGCCCAUUCACUUCGA-GUCCCAAUGC----------------------------CCCAGCCCA .(((((((.((....)).)))))))..((((..((((((((((...((....))...))).((((.....))-))....))))----------------------------))).)))). ( -34.30) >consensus CGAAAUGUAGUUAAAAUAACAUUUUUGGGGCCGGGGGCGUGGCAGUGCGUGAGCUUAGCCCAUUCACUCCGAAGUCCCAAGGCCCAAGUC_______CCAAGUCCUAAGCCCCCAGCCCA .(((((((.((....)).)))))))..((((.((((((.(((.((((.(((.((...)).))).)))))))..(((....))).........................)))))).)))). (-30.77 = -31.90 + 1.13)

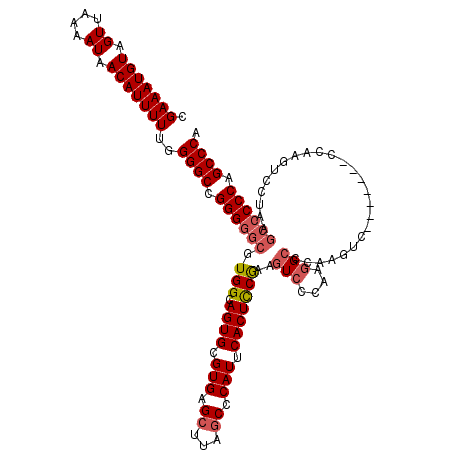
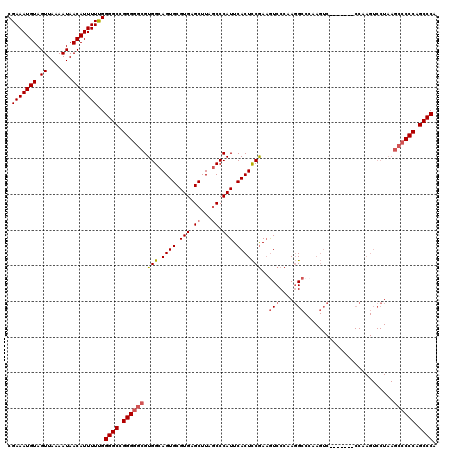
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:36:56 2006