
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 5,398,700 – 5,398,912 |
| Length | 212 |
| Max. P | 0.992174 |

| Location | 5,398,700 – 5,398,794 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 69.03 |
| Mean single sequence MFE | -27.27 |
| Consensus MFE | -15.90 |
| Energy contribution | -15.80 |
| Covariance contribution | -0.10 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.609624 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5398700 94 - 27905053 CUUGAUAACAACCGCAUACGGUUGUGUUCGUGAGCGGAUUGCCUUCUG-----------UUGUGGAACCCUAUGAAAAAAAGUUUCCCAAAA---------AAAUCAAAAACUU------ .(((((.(((((((....)))))))((((...((((((......))))-----------))...))))........................---------..)))))......------ ( -20.20) >DroSim_CAF1 8429 100 - 1 CUUGAUA---ACCGCAUGCGGUUGUGUUCGUGAGCGGAUUGCCUUCUG-----------UUAUGGAACCCUAUGAAAAAAGGUUCCCCAAAAUCGCGAUAAUUAUCAAAAACUG------ .((((((---((((....)))).....((((((..((...(((((...-----------((((((....))))))...)))))...))....))))))....))))))......------ ( -27.70) >DroYak_CAF1 8761 116 - 1 CUUGAUA---ACCGCAUGCGGUUGUGUUUGUGAGCGGAUUGCUUUCUGGUUGCUUUCAGCUGUGGAGGCCUAAACAGAAAGAUUCCCGAAUAUCGCUUGA-AAAUCCGAAAGUUAAAGGA ....(((---((((....)))))))....((((.(((....(((((((...(((((((....))))))).....)))))))....)))....))))....-...(((..........))) ( -33.90) >consensus CUUGAUA___ACCGCAUGCGGUUGUGUUCGUGAGCGGAUUGCCUUCUG___________UUGUGGAACCCUAUGAAAAAAGGUUCCCCAAAAUCGC___A_AAAUCAAAAACUU______ .(((((.....((((.(((((......))))).))))..........................((((((...........)))))).................)))))............ (-15.90 = -15.80 + -0.10)



| Location | 5,398,754 – 5,398,874 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.11 |
| Mean single sequence MFE | -32.19 |
| Consensus MFE | -24.67 |
| Energy contribution | -24.29 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.93 |
| Structure conservation index | 0.77 |
| SVM decision value | 2.31 |
| SVM RNA-class probability | 0.992174 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5398754 120 + 27905053 AAUCCGCUCACGAACACAACCGUAUGCGGUUGUUAUCAAGCGCAGGUUGCUCGAAAUAAUCAAAUAAUUUCAUUUAAUGGCACCUGCGCUCAGAUUUGUCUACCGUCGUUUCGGAUUUAU ((((((...((((..(((((((....))))))).(((.(((((((((.(((..((((..............))))...))))))))))))..)))..........))))..))))))... ( -38.14) >DroSim_CAF1 8492 117 + 1 AAUCCGCUCACGAACACAACCGCAUGCGGU---UAUCAAGCGCAGGUUGCUCGAAAUAAUCAAAUAAUUUCAUUUAAUGGCACCUGCGCUCAGAUUUGUCUACCGUCGUUUCGGAUUUAU ((((((...((((....(((((....))))---)(((.(((((((((.(((..((((..............))))...))))))))))))..)))..........))))..))))))... ( -34.64) >DroYak_CAF1 8840 117 + 1 AAUCCGCUCACAAACACAACCGCAUGCGGU---UAUCAAGCGCAGGUUGCUCGAAAUAAUCAAAUAAUUUCAUUUAAUGGCACCUGCGCACAGAUUUGUCUACCGUCGUUUCGGAUUUAU ((((((.....(((((((((((....))))---).....((((((((.(((..((((..............))))...)))))))))))...............)).))))))))))... ( -31.54) >DroAna_CAF1 8407 99 + 1 ------------GACACA--C---GGCAGU---UAUCAAGCGCAGGUUGCUCAAAAUAAUCAAAUAAUUUCGUUUAAUAGCACCUGCGCACAGAUUUGUAUACUACUGCUU-GGAUUGAC ------------....((--.---((((((---......((((((((.(((..((((..............))))...)))))))))))((......)).....)))))))-)....... ( -24.44) >consensus AAUCCGCUCACGAACACAACCGCAUGCGGU___UAUCAAGCGCAGGUUGCUCGAAAUAAUCAAAUAAUUUCAUUUAAUGGCACCUGCGCACAGAUUUGUCUACCGUCGUUUCGGAUUUAU ...................(((...(((((....(((..((((((((.(((..((((..............))))...)))))))))))...)))......))))).....)))...... (-24.67 = -24.29 + -0.38)



| Location | 5,398,754 – 5,398,874 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.11 |
| Mean single sequence MFE | -36.93 |
| Consensus MFE | -25.58 |
| Energy contribution | -27.57 |
| Covariance contribution | 2.00 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.17 |
| Structure conservation index | 0.69 |
| SVM decision value | 2.16 |
| SVM RNA-class probability | 0.989267 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5398754 120 - 27905053 AUAAAUCCGAAACGACGGUAGACAAAUCUGAGCGCAGGUGCCAUUAAAUGAAAUUAUUUGAUUAUUUCGAGCAACCUGCGCUUGAUAACAACCGCAUACGGUUGUGUUCGUGAGCGGAUU ...((((((..((((..........(((.(((((((((((((((((((((....))))))))......).)).))))))))))))).(((((((....)))))))..))))...)))))) ( -43.00) >DroSim_CAF1 8492 117 - 1 AUAAAUCCGAAACGACGGUAGACAAAUCUGAGCGCAGGUGCCAUUAAAUGAAAUUAUUUGAUUAUUUCGAGCAACCUGCGCUUGAUA---ACCGCAUGCGGUUGUGUUCGUGAGCGGAUU ...((((((..((((..............(((((((((((((((((((((....))))))))......).)).)))))))))).(((---((((....)))))))..))))...)))))) ( -41.30) >DroYak_CAF1 8840 117 - 1 AUAAAUCCGAAACGACGGUAGACAAAUCUGUGCGCAGGUGCCAUUAAAUGAAAUUAUUUGAUUAUUUCGAGCAACCUGCGCUUGAUA---ACCGCAUGCGGUUGUGUUUGUGAGCGGAUU ...((((((..((((................(((((((((((((((((((....))))))))......).)).))))))))...(((---((((....)))))))..))))...)))))) ( -36.60) >DroAna_CAF1 8407 99 - 1 GUCAAUCC-AAGCAGUAGUAUACAAAUCUGUGCGCAGGUGCUAUUAAACGAAAUUAUUUGAUUAUUUUGAGCAACCUGCGCUUGAUA---ACUGCC---G--UGUGUC------------ ........-..(((((.(....)..(((...(((((((((((..(((.((((....)))).))).....))).))))))))..))).---))))).---.--......------------ ( -26.80) >consensus AUAAAUCCGAAACGACGGUAGACAAAUCUGAGCGCAGGUGCCAUUAAAUGAAAUUAUUUGAUUAUUUCGAGCAACCUGCGCUUGAUA___ACCGCAUGCGGUUGUGUUCGUGAGCGGAUU ...((((((..((((.(((......(((.(((((((((((((((((((((....))))))))......).)).)))))))))))))....)))(((......)))..))))...)))))) (-25.58 = -27.57 + 2.00)



| Location | 5,398,794 – 5,398,912 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.46 |
| Mean single sequence MFE | -31.64 |
| Consensus MFE | -25.16 |
| Energy contribution | -24.40 |
| Covariance contribution | -0.75 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.541254 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5398794 118 + 27905053 CGCAGGUUGCUCGAAAUAAUCAAAUAAUUUCAUUUAAUGGCACCUGCGCUCAGAUUUGUCUACCGUCGUUUCGGAUUUAUGGUG--AACUAUAGUAUGUGUGCGAUAAUGAAAAUCAGCG .((((((.(((..((((..............))))...)))))))))(((..(((((.((....(((((..((.((((((((..--..)))))).)).)).)))))...)))))))))). ( -29.94) >DroSim_CAF1 8529 118 + 1 CGCAGGUUGCUCGAAAUAAUCAAAUAAUUUCAUUUAAUGGCACCUGCGCUCAGAUUUGUCUACCGUCGUUUCGGAUUUAUGGUG--AACUAUAGUAUGUGUGCGAUAAUGAAAAUCAGCG .((((((.(((..((((..............))))...)))))))))(((..(((((.((....(((((..((.((((((((..--..)))))).)).)).)))))...)))))))))). ( -29.94) >DroYak_CAF1 8877 118 + 1 CGCAGGUUGCUCGAAAUAAUCAAAUAAUUUCAUUUAAUGGCACCUGCGCACAGAUUUGUCUACCGUCGUUUCGGAUUUAUGGUG--AACUAUAGUAUGUGUGCGAUAAUGAAAAUCAGCG .((((((.(((..((((..............))))...)))))))))(((((.((.......(((......)))..((((((..--..)))))).)).)))))(((.......))).... ( -28.84) >DroAna_CAF1 8427 119 + 1 CGCAGGUUGCUCAAAAUAAUCAAAUAAUUUCGUUUAAUAGCACCUGCGCACAGAUUUGUAUACUACUGCUU-GGAUUGACAGUGCGGGCGGUAGUAUGUGUGCGAUAAUGAAAAUCAGCG .((((((.(((..((((..............))))...)))))))))(((((......(((((((((((((-(.((.....)).)))))))))))))))))))(((.......))).... ( -37.84) >consensus CGCAGGUUGCUCGAAAUAAUCAAAUAAUUUCAUUUAAUGGCACCUGCGCACAGAUUUGUCUACCGUCGUUUCGGAUUUAUGGUG__AACUAUAGUAUGUGUGCGAUAAUGAAAAUCAGCG (((((((.(((..((((..............))))...))))))))))....(((((.......(((((.......(((((((....))))))).......))))).....))))).... (-25.16 = -24.40 + -0.75)

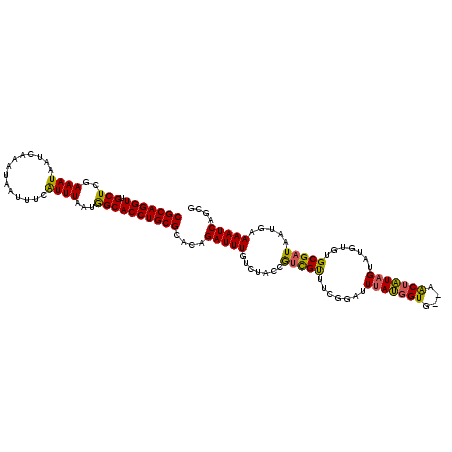

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:21:46 2006