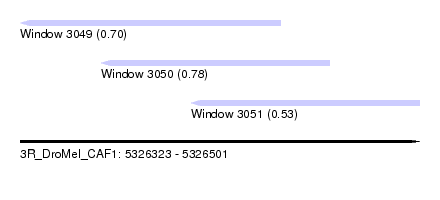
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 5,326,323 – 5,326,501 |
| Length | 178 |
| Max. P | 0.784026 |

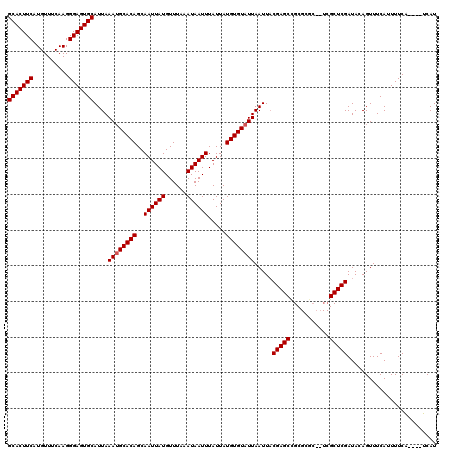
| Location | 5,326,323 – 5,326,439 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.83 |
| Mean single sequence MFE | -24.90 |
| Consensus MFE | -24.10 |
| Energy contribution | -24.43 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.701744 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5326323 116 - 27905053 GCACUUCAUGUUUCAAGGGAGUGCAUUAAAUGCACAGCAAUUAUGUUUAAAUAAUUUAUUAUGUGUAUUAAUUACGAGCCGCGCGCUCUCGCUCGAUACAGUUUCAUUUUCA----UCAU (((((((..........)))))))....((((((((..((((((......)))))).....)))))))).....(((((...........))))).................----.... ( -25.10) >DroSim_CAF1 31899 114 - 1 GCACUUCAUGUUUCAAGGGAGUGCAUUAAAUGCACAGCAAUUAUGUUUAAAUAAUUUAUUAUGUGUAUUAAUUACGAGCUGCGCGC--UCGCUCGAUACAGUUUCAUUUUCA----UCAU (((((((..........)))))))....((((((((..((((((......)))))).....)))))))).....(((((.....))--))).....................----.... ( -25.90) >DroYak_CAF1 31479 112 - 1 GCACUUCAUGUUCCAAGGGAGUGCAUUAAAGGCACAGCAAUUAUGUUUAAAUAAUUUAUUAUGUGUAUUAAUUUCGAGCCGC--------GCUCGAUACAGUUUCAUUUUCCAUUUGCAU (((((((..........)))))))(((((..(((((..((((((......)))))).....))))).))))).((((((...--------))))))........................ ( -23.70) >consensus GCACUUCAUGUUUCAAGGGAGUGCAUUAAAUGCACAGCAAUUAUGUUUAAAUAAUUUAUUAUGUGUAUUAAUUACGAGCCGCGCGC__UCGCUCGAUACAGUUUCAUUUUCA____UCAU (((((((..........)))))))....((((((((..((((((......)))))).....)))))))).....(((((...........)))))......................... (-24.10 = -24.43 + 0.33)


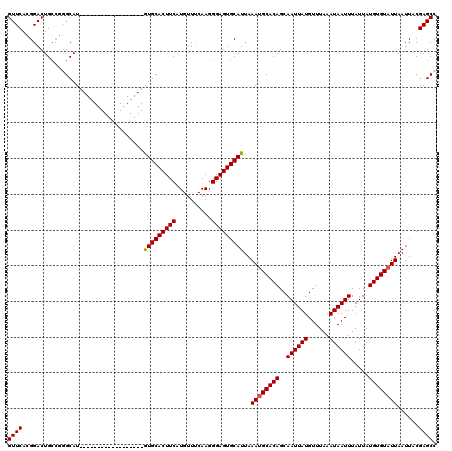
| Location | 5,326,359 – 5,326,461 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.63 |
| Mean single sequence MFE | -27.87 |
| Consensus MFE | -23.29 |
| Energy contribution | -23.40 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.784026 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5326359 102 - 27905053 GUUCACGGACUGCCGGGCAU------------------GUGCACUUCAUGUUUCAAGGGAGUGCAUUAAAUGCACAGCAAUUAUGUUUAAAUAAUUUAUUAUGUGUAUUAAUUACGAGCC .....(((....)))(((..------------------(((((((((..........)))))))))..((((((((..((((((......)))))).....))))))))........))) ( -27.70) >DroSim_CAF1 31933 102 - 1 GUUCACGGACUGGCGGGCAU------------------GUGCACUUCAUGUUUCAAGGGAGUGCAUUAAAUGCACAGCAAUUAUGUUUAAAUAAUUUAUUAUGUGUAUUAAUUACGAGCU ((((................------------------(((((((((..........)))))))))..((((((((..((((((......)))))).....))))))))......)))). ( -24.60) >DroYak_CAF1 31511 120 - 1 GUUCAGGGACUGCAAUGCAUGUACAUGUGCAUAUGCAUAUGCACUUCAUGUUCCAAGGGAGUGCAUUAAAGGCACAGCAAUUAUGUUUAAAUAAUUUAUUAUGUGUAUUAAUUUCGAGCC ((((.(..(.((((((((((......)))))).)))).(((((((((..........))))))))).....(((((..((((((......)))))).....))))).....)..))))). ( -31.30) >consensus GUUCACGGACUGCCGGGCAU__________________GUGCACUUCAUGUUUCAAGGGAGUGCAUUAAAUGCACAGCAAUUAUGUUUAAAUAAUUUAUUAUGUGUAUUAAUUACGAGCC ((((..................................(((((((((..........)))))))))..((((((((..((((((......)))))).....))))))))......)))). (-23.29 = -23.40 + 0.11)


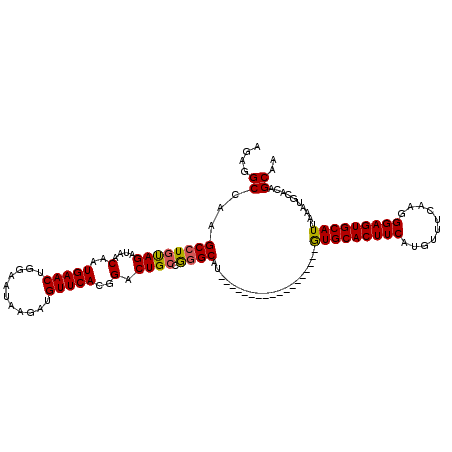
| Location | 5,326,399 – 5,326,501 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.04 |
| Mean single sequence MFE | -30.29 |
| Consensus MFE | -22.99 |
| Energy contribution | -23.00 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.76 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.528477 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5326399 102 - 27905053 AGAGGCCAAGCCUGUAGAUAACAAUGAACUGGAAUAAGUUGUUCACGGACUGCCGGGCAU------------------GUGCACUUCAUGUUUCAAGGGAGUGCAUUAAAUGCACAGCAA ...(((..((.((((..(((((...............)))))..)))).)))))..((((------------------(((((((((..........)))))))))...))))....... ( -27.86) >DroSim_CAF1 31973 102 - 1 AGAGGCCAAGCCUGUAGAUAACAAUGAACUGGAAUAAGAUGUUCACGGACUGGCGGGCAU------------------GUGCACUUCAUGUUUCAAGGGAGUGCAUUAAAUGCACAGCAA ....((((..(((((.....))).(((((...........))))).))..))))..((((------------------(((((((((..........)))))))))...))))....... ( -28.10) >DroYak_CAF1 31551 120 - 1 AGAGGCCAAGCCUGCAGAUAACAAUGAACUGGAGUAAGAUGUUCAGGGACUGCAAUGCAUGUACAUGUGCAUAUGCAUAUGCACUUCAUGUUCCAAGGGAGUGCAUUAAAGGCACAGCAA ....((...((((((((....(..(((((...........)))))..).)))).((((((((.(....).))))))))(((((((((..........)))))))))...))))...)).. ( -34.90) >consensus AGAGGCCAAGCCUGUAGAUAACAAUGAACUGGAAUAAGAUGUUCACGGACUGCCGGGCAU__________________GUGCACUUCAUGUUUCAAGGGAGUGCAUUAAAUGCACAGCAA ....((...((((((((....(..(((((...........)))))..).)))).))))....................(((((((((..........)))))))))..........)).. (-22.99 = -23.00 + 0.01)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:21:11 2006