
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 5,297,945 – 5,298,041 |
| Length | 96 |
| Max. P | 0.701522 |

| Location | 5,297,945 – 5,298,041 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 82.08 |
| Mean single sequence MFE | -38.65 |
| Consensus MFE | -24.08 |
| Energy contribution | -23.25 |
| Covariance contribution | -0.83 |
| Combinations/Pair | 1.47 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.701522 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
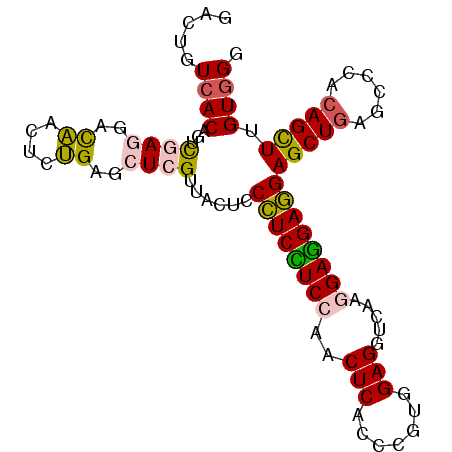
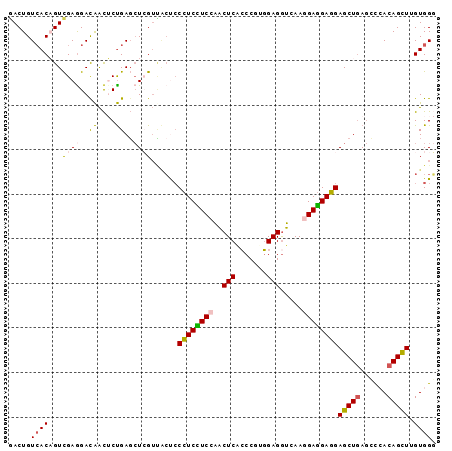
>3R_DroMel_CAF1 5297945 96 + 27905053 CCCACAAGCUGUGAGCUGAGCUCCUCCUCCUUGACCUCCACGGGUGAGUUGGAGGAGGGAGUAACGAGCUCAGAGUUGUCCUCGACUGUGACAGUU ......((((((.(.((((((((...((((((..(((((((......).))))))))))))....))))))))(((((....))))).).)))))) ( -39.90) >DroSim_CAF1 2179 96 + 1 CCCACAAGCUGUGGACUCAGCUCCUCCUCCUUGACCUCCACGGGUGAGUUGGAGGAGGGAGUAACGAGCUCAGAGUUGUCUUCGACUGUGACAGUC .((((.....))))((((..(((((((..(((.((((....)))))))..))))))).)))).....(.(((.(((((....))))).)))).... ( -38.00) >DroEre_CAF1 2416 96 + 1 CCCACCAGCUGUGGGCUCAGCUCCUCCUCCUUGACCUCCACGGGCGAGUUGGAGGAGGGGGUAACGAGCUCAGAGUUGUCCUCGACUGUGACAGUC .....(((((.(((((((....((((((((((.(.((((....).))).).))))))))))....))))))).))))).....(((((...))))) ( -44.50) >DroYak_CAF1 2171 96 + 1 CCCACUAACUGUGGGCUCAGCUCCUCCUCCUUGACCUCCACGGGUGAGUUCGAGGAGGGAGUGACGAGCUCAGAGUUGUCCUCGACUGUGACAGUC .....(((((.(((((((.(((((..((((((((.((((....).))).)))))))))))))...))))))).))))).....(((((...))))) ( -42.70) >DroMoj_CAF1 2509 96 + 1 GCCACCAGCUGUUGGCUCAGCUCAUCGUCUUUGACCUCGUCGGCUGAGCCUGACGAUGUGCCACCCAGCUCGGUGCCAUCCUCGACAGUUACAGUC ......((((((((((((((((.((.(((...)))...)).))))))))).((.((((.((.(((......))))))))).)))))))))...... ( -35.30) >DroAna_CAF1 2674 96 + 1 GCCACAAGCUGUUGGUUCAGCUCCUCCUCCUUCAUCUCCACGGGGGAGUUGGAAGAGGGAGGCACCAUGUCAGUAUUAUCCUCAACGGUGACAGUC (((...(((((......)))))..(((..((((..((((.....))))...))))..))))))....(((((.(.((......)).).)))))... ( -31.50) >consensus CCCACAAGCUGUGGGCUCAGCUCCUCCUCCUUGACCUCCACGGGUGAGUUGGAGGAGGGAGUAACGAGCUCAGAGUUGUCCUCGACUGUGACAGUC .......(((((.((((..((((((((((((((.((.....)).))))...))))))).)))....))))((.(((((....))))).))))))). (-24.08 = -23.25 + -0.83)



| Location | 5,297,945 – 5,298,041 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 82.08 |
| Mean single sequence MFE | -36.48 |
| Consensus MFE | -22.94 |
| Energy contribution | -22.83 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.63 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5297945 96 - 27905053 AACUGUCACAGUCGAGGACAACUCUGAGCUCGUUACUCCCUCCUCCAACUCACCCGUGGAGGUCAAGGAGGAGGAGCUCAGCUCACAGCUUGUGGG ...((((.(......)))))...((((((((.((.((((..((((((.(......)))))))....)))))).))))))))(((((.....))))) ( -37.20) >DroSim_CAF1 2179 96 - 1 GACUGUCACAGUCGAAGACAACUCUGAGCUCGUUACUCCCUCCUCCAACUCACCCGUGGAGGUCAAGGAGGAGGAGCUGAGUCCACAGCUUGUGGG (((((...))))).......((((..(((((.((.((((..((((((.(......)))))))....)))))).)))))))))((((.....)))). ( -36.40) >DroEre_CAF1 2416 96 - 1 GACUGUCACAGUCGAGGACAACUCUGAGCUCGUUACCCCCUCCUCCAACUCGCCCGUGGAGGUCAAGGAGGAGGAGCUGAGCCCACAGCUGGUGGG (((((...)))))(((.....)))...(((((((.((.((((((....(((.......)))....)))))).))))).))))((((.....)))). ( -36.30) >DroYak_CAF1 2171 96 - 1 GACUGUCACAGUCGAGGACAACUCUGAGCUCGUCACUCCCUCCUCGAACUCACCCGUGGAGGUCAAGGAGGAGGAGCUGAGCCCACAGUUAGUGGG (((((...)))))(((.....)))(.(((((.((.((((((((.((........)).)))).....)))))).))))).).(((((.....))))) ( -38.50) >DroMoj_CAF1 2509 96 - 1 GACUGUAACUGUCGAGGAUGGCACCGAGCUGGGUGGCACAUCGUCAGGCUCAGCCGACGAGGUCAAAGACGAUGAGCUGAGCCAACAGCUGGUGGC (((.......)))......(.((((.(((((((((((.(((((((.(((...)))(((...)))...))))))).)))..)))..))))))))).) ( -38.10) >DroAna_CAF1 2674 96 - 1 GACUGUCACCGUUGAGGAUAAUACUGACAUGGUGCCUCCCUCUUCCAACUCCCCCGUGGAGAUGAAGGAGGAGGAGCUGAACCAACAGCUUGUGGC ......(((((((.((.......)).).))))))(((((..((((...((((.....))))..))))..)))))(((((......)))))...... ( -32.40) >consensus GACUGUCACAGUCGAGGACAACUCUGAGCUCGUUACUCCCUCCUCCAACUCACCCGUGGAGGUCAAGGAGGAGGAGCUGAGCCCACAGCUUGUGGG .....((((...((((..((....))..))))......((((((((..(((.......))).....))))))))(((((......))))).)))). (-22.94 = -22.83 + -0.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:20:57 2006