
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 5,120,323 – 5,120,442 |
| Length | 119 |
| Max. P | 0.993008 |

| Location | 5,120,323 – 5,120,442 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.55 |
| Mean single sequence MFE | -23.20 |
| Consensus MFE | -22.36 |
| Energy contribution | -22.70 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.97 |
| Structure conservation index | 0.96 |
| SVM decision value | 2.37 |
| SVM RNA-class probability | 0.993008 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5120323 119 + 27905053 UCCAGCAAAUAUUACAAUACCAAAGUGCAAAAUCACAGACCCCAAAAUAAACUCACUGUGACUUUGACAUUUGCGUAAUAUUUGUGGUCCUAUAUGCAAAUCAAAUA-AAAAAAAAACAA .(((.((((((((((.......(((((((((.((((((.................)))))).)))).)))))..)))))))))))))....................-............ ( -23.53) >DroSim_CAF1 22495 113 + 1 UCCAGCAAAUAUUACAGUACCAAAGUGCAAAAUCACAGACCCCAAAAUAAACUCACUGUGACUUUGACAUUUGCGUAAUAUUUGUGGUCCUA--UGCAAAU----UA-AAAAAAAAACAA .(((.((((((((((.(((....((((((((.((((((.................)))))).)))).)))))))))))))))))))).....--.......----..-............ ( -24.63) >DroYak_CAF1 23625 116 + 1 GCCAGCAAAUAUUACAGUUCCAAAGUGCAAACUCACAGACCCCAAAAUAAACUCACUGUGACUUUGACAUUUGCAUAAUAUUUGUGGACCUAUAUGCAAAU----GAUAAAAACAAACAA .(((.(((((((((..((....(((((((((.((((((.................)))))).)))).))))))).))))))))))))..............----............... ( -21.43) >consensus UCCAGCAAAUAUUACAGUACCAAAGUGCAAAAUCACAGACCCCAAAAUAAACUCACUGUGACUUUGACAUUUGCGUAAUAUUUGUGGUCCUAUAUGCAAAU____UA_AAAAAAAAACAA .(((.((((((((((.......(((((((((.((((((.................)))))).)))).)))))..)))))))))))))................................. (-22.36 = -22.70 + 0.33)
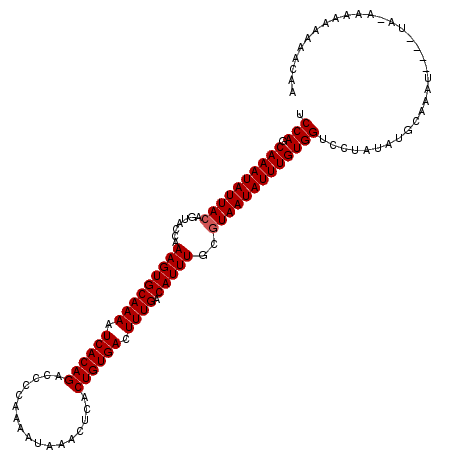


| Location | 5,120,323 – 5,120,442 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.55 |
| Mean single sequence MFE | -27.63 |
| Consensus MFE | -26.75 |
| Energy contribution | -26.86 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938192 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5120323 119 - 27905053 UUGUUUUUUUUU-UAUUUGAUUUGCAUAUAGGACCACAAAUAUUACGCAAAUGUCAAAGUCACAGUGAGUUUAUUUUGGGGUCUGUGAUUUUGCACUUUGGUAUUGUAAUAUUUGCUGGA .........(((-(((.((.....)).))))))((((((((((((((((((.(((((((((((((.................))))))))))).))))))....))))))))))).))). ( -30.03) >DroSim_CAF1 22495 113 - 1 UUGUUUUUUUUU-UA----AUUUGCA--UAGGACCACAAAUAUUACGCAAAUGUCAAAGUCACAGUGAGUUUAUUUUGGGGUCUGUGAUUUUGCACUUUGGUACUGUAAUAUUUGCUGGA ............-..----.......--.....((((((((((((((((((.(((((((((((((.................))))))))))).))))))....))))))))))).))). ( -28.73) >DroYak_CAF1 23625 116 - 1 UUGUUUGUUUUUAUC----AUUUGCAUAUAGGUCCACAAAUAUUAUGCAAAUGUCAAAGUCACAGUGAGUUUAUUUUGGGGUCUGUGAGUUUGCACUUUGGAACUGUAAUAUUUGCUGGC ((((.(((.......----....))).))))..((((((((((((((((((.((((((.((((((.................)))))).)))).))))))....))))))))))).))). ( -24.13) >consensus UUGUUUUUUUUU_UA____AUUUGCAUAUAGGACCACAAAUAUUACGCAAAUGUCAAAGUCACAGUGAGUUUAUUUUGGGGUCUGUGAUUUUGCACUUUGGUACUGUAAUAUUUGCUGGA .................................((((((((((((((((((.(((((((((((((.................))))))))))).))))))....))))))))))).))). (-26.75 = -26.86 + 0.11)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:19:44 2006