
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 5,103,155 – 5,103,289 |
| Length | 134 |
| Max. P | 0.994027 |

| Location | 5,103,155 – 5,103,254 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.33 |
| Mean single sequence MFE | -36.14 |
| Consensus MFE | -28.70 |
| Energy contribution | -29.14 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.59 |
| SVM RNA-class probability | 0.966251 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5103155 99 - 27905053 UGCUGGGCAAAUAUUUACAUGGCGCACUGACAU---UGUUGGCUCACUCGAGUCAACAG------------------CAAAAUGCCAGCGUUGUGGUGCCUCAGGAAUUCAGCCGGAGAC ..(((((((........((..((((........---(((((((((....)))))))))(------------------(.....))..))))..)).)))).)))..........(....) ( -33.20) >DroSec_CAF1 9863 101 - 1 UGCUGGCCAAAUAUUUACAUGGCGCACUGCCAUU-UUGUUGGCUCACUCGAGUCAACAG------------------CAAAAUGCCAGCGUUGUGGUGCCUCAGGAAUUCAGCCGGAGAC .((((((...........(((((.....))))).-((((((((((....))))))))))------------------......))))))..........(((.((.......)).))).. ( -34.20) >DroSim_CAF1 10110 101 - 1 UGCUGGCCAAAUAUUUACAUGGCGCACUGGCAUU-UUGUUGGCUCACUCGAGUCAACAG------------------CAAAAUGCCAGCGUUGUGGUGCCUCAGGAAUUCAGCCGGAGAC ..(((((.....((((....(((((.((((((((-((((((((((....))))))...)------------------))))))))))).......)))))....))))...))))).... ( -40.40) >DroEre_CAF1 10154 120 - 1 UGCUGGCCAAAUAUUUACAUGGCGCACUGACAUUGUUGUUGGCUCACUCGAGUCAACAGCAAAAACAGCAAAAACAGCAAAAUGCCAGCGUUGUGGUGCCCCAGGAAUUCUGCCGGAGAC ..(((((.....((((...(((.(((((....(((((((((((((....))))))))))))).....((((.....((.....)).....))))))))).))).))))...))))).... ( -42.00) >DroYak_CAF1 11056 99 - 1 UGCUGGCCAAAUAUUUACAUGGCGCACUGACAUUG---UUGGUUCACUCGAGUCAACAG------------------CAAAAUGCCAGCGUUGUGGUGCCCCAGGAAUUUUGCCGGAGAC ..(((((.(((.((((...(((.(((((.((.(((---(((..((....))..))))))------------------...((((....))))))))))).))).)))))))))))).... ( -30.90) >consensus UGCUGGCCAAAUAUUUACAUGGCGCACUGACAUU_UUGUUGGCUCACUCGAGUCAACAG__________________CAAAAUGCCAGCGUUGUGGUGCCUCAGGAAUUCAGCCGGAGAC ..(((((.....((((....(((((...........(((((((((....)))))))))...................((.((((....)))).)))))))....))))...))))).... (-28.70 = -29.14 + 0.44)



| Location | 5,103,195 – 5,103,289 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.32 |
| Mean single sequence MFE | -30.50 |
| Consensus MFE | -21.52 |
| Energy contribution | -21.56 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.11 |
| Mean z-score | -3.40 |
| Structure conservation index | 0.71 |
| SVM decision value | 2.44 |
| SVM RNA-class probability | 0.994027 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5103195 94 - 27905053 UGCGACGUUAUUAUCUGGCAAAAUAGCCGA-----AAAAAUGCUGGGCAAAUAUUUACAUGGCGCACUGACAU---UGUUGGCUCACUCGAGUCAACAG------------------CAA ((((.((((((....((((......)))).-----.....(((...))).........)))))))........---(((((((((....))))))))))------------------)). ( -24.70) >DroSec_CAF1 9903 95 - 1 UGCGCCGCUAUUAUCUGGCGAAAUAGCC-A-----AAAAAUGCUGGCCAAAUAUUUACAUGGCGCACUGCCAUU-UUGUUGGCUCACUCGAGUCAACAG------------------CAA (((((((.((((.(.((((......)))-)-----.).)))).))))...........(((((.....))))).-.(((((((((....))))))))))------------------)). ( -32.90) >DroSim_CAF1 10150 95 - 1 UGCGCCGCUAUUAUCUGGCGAAAUAGCC-A-----AAAAAUGCUGGCCAAAUAUUUACAUGGCGCACUGGCAUU-UUGUUGGCUCACUCGAGUCAACAG------------------CAA ((((((((.(((.(.((((......)))-)-----.).)))))..((((..........)))).....)))...-.(((((((((....))))))))))------------------)). ( -32.80) >DroEre_CAF1 10194 119 - 1 UUCGUCGCCAUUAUCUGGCAAAAUAGCC-AACAAAAAAAAUGCUGGCCAAAUAUUUACAUGGCGCACUGACAUUGUUGUUGGCUCACUCGAGUCAACAGCAAAAACAGCAAAAACAGCAA ......((((.....)))).........-...........(((((((((..........))))...(((...(((((((((((((....)))))))))))))...)))......))))). ( -37.50) >DroYak_CAF1 11096 93 - 1 UUCGACGCCAUUAUCUGGCAAAAUAGCC-A-----AAAAAUGCUGGCCAAAUAUUUACAUGGCGCACUGACAUUG---UUGGUUCACUCGAGUCAACAG------------------CAA ...(.(((((((((.((((....((((.-.-----......)))))))).))).....))))))).......(((---(((..((....))..))))))------------------... ( -24.60) >consensus UGCGACGCUAUUAUCUGGCAAAAUAGCC_A_____AAAAAUGCUGGCCAAAUAUUUACAUGGCGCACUGACAUU_UUGUUGGCUCACUCGAGUCAACAG__________________CAA ...(.(((((((((.((((....((((..............)))))))).))).....)))))))...........(((((((((....)))))))))...................... (-21.52 = -21.56 + 0.04)

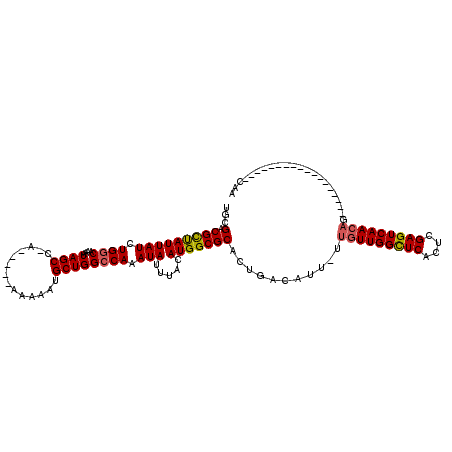

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:19:07 2006