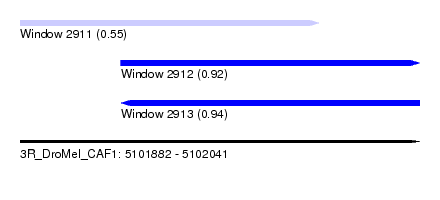

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 5,101,882 – 5,102,041 |
| Length | 159 |
| Max. P | 0.936480 |

| Location | 5,101,882 – 5,102,001 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.89 |
| Mean single sequence MFE | -27.58 |
| Consensus MFE | -19.48 |
| Energy contribution | -20.64 |
| Covariance contribution | 1.16 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.546070 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5101882 119 + 27905053 GCGAAAACAUUGGUAAUAACUUCAUAAAACACGAUAGUCGACUAAUUUGAUUGCGCUUUGCAAUUAAACUUGCCCCA-AAUGGGGGCACAAUCCCAUUUCAGGGUUGGCCAAACUCGUUC ((((.....(((((.................((.....)).....(((((((((.....)))))))))..((((((.-....))))))((((((.......)))))))))))..)))).. ( -31.20) >DroSec_CAF1 8813 110 + 1 GCGAAAACAUUUGUAAUAACUUCAUAAAACACGAUAGUCGACUAAUUUGAUUGCGCUUCGCAAUUAAACUUGCCCCA-AAUGGGGGCACAAUCCCAUUUCAGGGCACCUGA--------- (((((.((....)).................((.(((((((.....))))))))).))))).........(((((.(-(((((((......))))))))..))))).....--------- ( -28.10) >DroSim_CAF1 8847 119 + 1 GCGAAAACAUUUGUAAUAACUUCAUAAAACACGAUAGUCGACUAAUUUGAUUGCGCUUUGCAAUUAAACUUGCCCCA-AAUGGGGGCACAAUCCCAUUUCAGGGUUGGCCAAACUCGUUC (((((....)))))................((((..(((((((..(((((((((.....)))))))))..((((((.-....))))))..............))))))).....)))).. ( -29.30) >DroEre_CAF1 8895 120 + 1 GCGAAAACAUUUGUAAUUACUUCAUAAAACACGGUAGUCCACUAAUUUGAUUGCAUUUUGCAAUUAAACUUGCCCCAAAAUGGUAGCACAACCCCAUUUCAGGGUUGGCCGAACCCGUUC (((((....)))))................((((...((......(((((((((.....)))))))))..((((.......))))((.((((((.......)))))))).))..)))).. ( -26.50) >DroYak_CAF1 9792 120 + 1 GCGAAAACAUUUGUAAUAACUUCAUAAAACACGAUAGUCGACUAAUUUGAUUGCAUUUUGCAAUUAAACUUGCCCCAAAAUGCGAGCACAACCCCAUUUCGGUGUUGGCCAAACUCGUUC (((((....)))))................(((((((....))).((((.(((((((((((((......))))...)))))))))((.((((.((.....)).)))))))))).)))).. ( -22.80) >consensus GCGAAAACAUUUGUAAUAACUUCAUAAAACACGAUAGUCGACUAAUUUGAUUGCGCUUUGCAAUUAAACUUGCCCCA_AAUGGGGGCACAAUCCCAUUUCAGGGUUGGCCAAACUCGUUC (((((....)))))....................(((....))).((((((((((...))))))))))..((((((......))))))((((((.......))))))............. (-19.48 = -20.64 + 1.16)

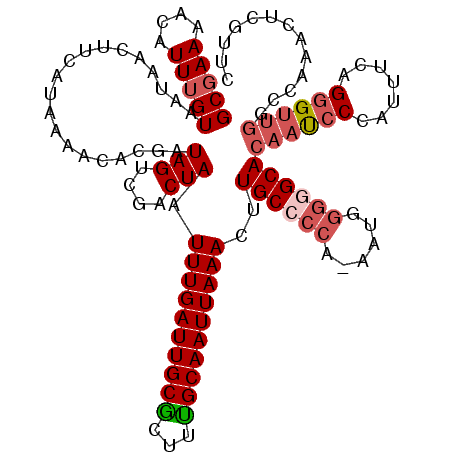
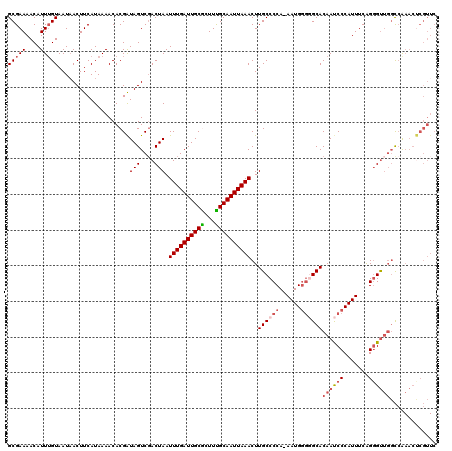
| Location | 5,101,922 – 5,102,041 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.78 |
| Mean single sequence MFE | -29.57 |
| Consensus MFE | -25.68 |
| Energy contribution | -26.24 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.916200 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5101922 119 + 27905053 ACUAAUUUGAUUGCGCUUUGCAAUUAAACUUGCCCCA-AAUGGGGGCACAAUCCCAUUUCAGGGUUGGCCAAACUCGUUCGUGUACCUUAGCCCUUUGCCUAAUUAAACGAAAUUGUUUC .....(((((((((.....)))))))))..((((((.-....))))))((((((.......))))))...((((...(((((.....((((........))))....)))))...)))). ( -32.00) >DroSim_CAF1 8887 119 + 1 ACUAAUUUGAUUGCGCUUUGCAAUUAAACUUGCCCCA-AAUGGGGGCACAAUCCCAUUUCAGGGUUGGCCAAACUCGUUCGUGUACCUUAGCCCUUUGCAUAAUUAAACGAAAUUGUUUC .....(((((((((.....)))))))))..((((((.-....))))))((((((.......))))))...((((...(((((........((.....))........)))))...)))). ( -31.79) >DroEre_CAF1 8935 119 + 1 ACUAAUUUGAUUGCAUUUUGCAAUUAAACUUGCCCCAAAAUGGUAGCACAACCCCAUUUCAGGGUUGGCCGAACCCGUUCGUGUACC-CAGUCCUUUGCCUAAUUAAACGAAAUUGUUUC .....(((((((((.....)))))))))..((((.......))))((.((((((.......)))))))).((((...(((((.((..-(((....)))..)).....)))))...)))). ( -26.40) >DroYak_CAF1 9832 119 + 1 ACUAAUUUGAUUGCAUUUUGCAAUUAAACUUGCCCCAAAAUGCGAGCACAACCCCAUUUCGGUGUUGGCCAAACUCGUUCGUGGGCC-CAGUCCUUUGCCUAAUUAAACGAAAUUGUUUC .....((((.(((((((((((((......))))...)))))))))((.((((.((.....)).)))))))))).(((((..(((((.-.........)))))....)))))......... ( -28.10) >consensus ACUAAUUUGAUUGCACUUUGCAAUUAAACUUGCCCCA_AAUGGGAGCACAACCCCAUUUCAGGGUUGGCCAAACUCGUUCGUGUACC_CAGCCCUUUGCCUAAUUAAACGAAAUUGUUUC .....(((((((((.....)))))))))..((((((......))))))((((((.......))))))...((((...(((((........((.....))........)))))...)))). (-25.68 = -26.24 + 0.56)

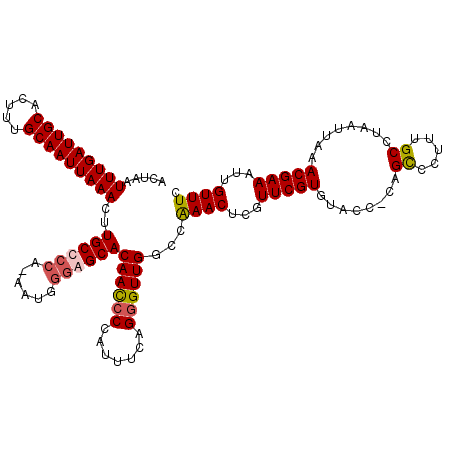
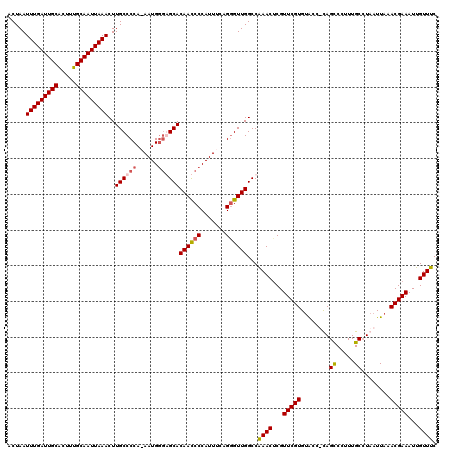
| Location | 5,101,922 – 5,102,041 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.78 |
| Mean single sequence MFE | -35.00 |
| Consensus MFE | -29.36 |
| Energy contribution | -28.99 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.936480 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 5101922 119 - 27905053 GAAACAAUUUCGUUUAAUUAGGCAAAGGGCUAAGGUACACGAACGAGUUUGGCCAACCCUGAAAUGGGAUUGUGCCCCCAUU-UGGGGCAAGUUUAAUUGCAAAGCGCAAUCAAAUUAGU ...(((((..((((...(((((.....(((((((.(.(......)).)))))))...)))))))))..)))))(((((....-.))))).(((((.(((((.....))))).)))))... ( -37.90) >DroSim_CAF1 8887 119 - 1 GAAACAAUUUCGUUUAAUUAUGCAAAGGGCUAAGGUACACGAACGAGUUUGGCCAACCCUGAAAUGGGAUUGUGCCCCCAUU-UGGGGCAAGUUUAAUUGCAAAGCGCAAUCAAAUUAGU ...(((((..(((((..........((((....(((..((......))...)))..)))).)))))..)))))(((((....-.))))).(((((.(((((.....))))).)))))... ( -34.20) >DroEre_CAF1 8935 119 - 1 GAAACAAUUUCGUUUAAUUAGGCAAAGGACUG-GGUACACGAACGGGUUCGGCCAACCCUGAAAUGGGGUUGUGCUACCAUUUUGGGGCAAGUUUAAUUGCAAAAUGCAAUCAAAUUAGU .((((......)))).....(((...(((((.-(.........).))))).)))..(((..((((((((.....)).))))))..)))..(((((.(((((.....))))).)))))... ( -34.90) >DroYak_CAF1 9832 119 - 1 GAAACAAUUUCGUUUAAUUAGGCAAAGGACUG-GGCCCACGAACGAGUUUGGCCAACACCGAAAUGGGGUUGUGCUCGCAUUUUGGGGCAAGUUUAAUUGCAAAAUGCAAUCAAAUUAGU ....(((.(((((((.....(((..(....).-.)))...))))))).)))(((....(((((((((((.....))).))))))))))).(((((.(((((.....))))).)))))... ( -33.00) >consensus GAAACAAUUUCGUUUAAUUAGGCAAAGGACUA_GGUACACGAACGAGUUUGGCCAACCCUGAAAUGGGAUUGUGCCCCCAUU_UGGGGCAAGUUUAAUUGCAAAACGCAAUCAAAUUAGU ...(((((..(((((.....(((...(((((..............))))).))).......)))))..)))))(((((......))))).(((((.(((((.....))))).)))))... (-29.36 = -28.99 + -0.37)

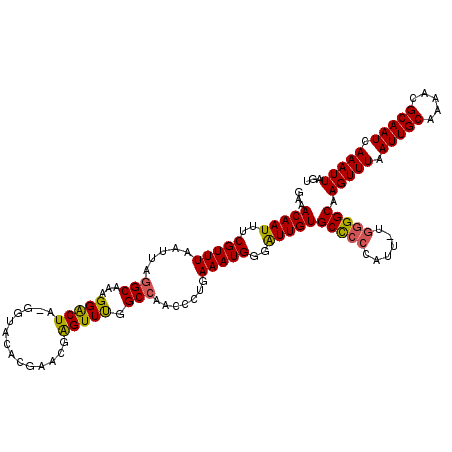

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:19:00 2006