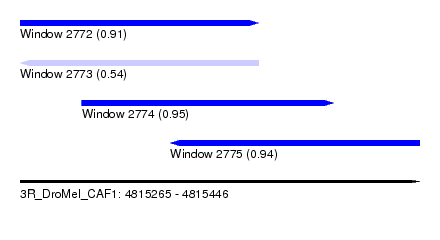

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,815,265 – 4,815,446 |
| Length | 181 |
| Max. P | 0.953130 |

| Location | 4,815,265 – 4,815,373 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 82.47 |
| Mean single sequence MFE | -35.35 |
| Consensus MFE | -29.11 |
| Energy contribution | -29.45 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.82 |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.909392 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4815265 108 + 27905053 GAAACGCGUCGACUGUGGUGCCCAAAAUUAUUGGAAAAACUACUGUGCUUCCGGGAAAACCACUACUACUCCACGCCAAAACAUUAAAACAGAUUGGCGGUAGUAGUG ..............(((((.(((.........((((..((....))..)))))))...)))))(((((((.(.((((((..............))))))).))))))) ( -29.64) >DroPse_CAF1 6957 108 + 1 GAGGGGGACCCACGAUGGUGCCCAAGAUCAUUGGCAAGACGACAGUGCUGUCGGGCAAGCCACUGCUGCUGCAUGCCAAGGGAAUCAAGCAGCUAGGAGGCGGUAGUG ...(((.(((......))).))).....(((((.(....(((((....))))).....(((.((((((((.....((...)).....)))))).))..)))).))))) ( -38.20) >DroSec_CAF1 4777 108 + 1 GAAACGCGUCGACUGUGGUGCCCAAAAUUAUUGGAAAAACUACUGUGCUUCCGGGAAAACCACUACUACUGAACGCCAAGACAUUAAAACAGCUUGGCAGCAGUAGUG ..............(((((.(((.........((((..((....))..)))))))...)))))((((((((...((((((.(.........)))))))..)))))))) ( -33.10) >DroSim_CAF1 6210 108 + 1 GAAACGCGUCGACUGUGGUGCCCAAAAUUAUUGGAAAAACUACUGUGCUUCCGGGAAAACCACUACUACUGCACGCCAAGACAUUAAAACAGCUUGGCAGCAGUAGUG ..............(((((.(((.........((((..((....))..)))))))...)))))(((((((((..((((((.(.........))))))).))))))))) ( -35.50) >DroYak_CAF1 4610 108 + 1 GAAACACAUCGACUGUGGUACCCAAAAUUAUUGGAAAAACUACUGUGCUUCCGGGAAAACCACUACUACUGCACGCCAAAACAUUAAAACAGCUUGGCGGCAGUAGUG ..............(((((.(((.........((((..((....))..)))))))...)))))(((((((((.((((((..............))))))))))))))) ( -34.24) >DroPer_CAF1 6921 108 + 1 GAGGGGGACCCACGAUGGUGCCCAAGAUCAUUGGCAAGACGACAGUGCUGUCGGGCAAGCCACUGCUGCUGCAUGCCAAGGCAAUCAAGCAGCUAGGAGGCGGUAGUG ...(((.(((......))).))).....(((((.(....(((((....))))).....(((.((((((((...(((....)))....)))))).))..)))).))))) ( -41.40) >consensus GAAACGCAUCGACUGUGGUGCCCAAAAUUAUUGGAAAAACUACUGUGCUUCCGGGAAAACCACUACUACUGCACGCCAAGACAUUAAAACAGCUUGGCGGCAGUAGUG ..............(((((.(((((.....))((((..((....))..)))))))...))))).((((((((.(((((((............))))))))))))))). (-29.11 = -29.45 + 0.34)



| Location | 4,815,265 – 4,815,373 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 82.47 |
| Mean single sequence MFE | -34.52 |
| Consensus MFE | -21.42 |
| Energy contribution | -23.70 |
| Covariance contribution | 2.28 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.536928 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4815265 108 - 27905053 CACUACUACCGCCAAUCUGUUUUAAUGUUUUGGCGUGGAGUAGUAGUGGUUUUCCCGGAAGCACAGUAGUUUUUCCAAUAAUUUUGGGCACCACAGUCGACGCGUUUC .((((((.(((((((..(((....)))..)))))).).)))))).(((((...((((((((.((....)).)))))((.....))))).))))).............. ( -29.80) >DroPse_CAF1 6957 108 - 1 CACUACCGCCUCCUAGCUGCUUGAUUCCCUUGGCAUGCAGCAGCAGUGGCUUGCCCGACAGCACUGUCGUCUUGCCAAUGAUCUUGGGCACCAUCGUGGGUCCCCCUC .....((((......((((((..((.((...)).))..)))))).((((..(((((((((....))))(((........)))...))))))))).))))......... ( -32.40) >DroSec_CAF1 4777 108 - 1 CACUACUGCUGCCAAGCUGUUUUAAUGUCUUGGCGUUCAGUAGUAGUGGUUUUCCCGGAAGCACAGUAGUUUUUCCAAUAAUUUUGGGCACCACAGUCGACGCGUUUC .(((((((.((((((((.........).)))))))..))))))).(((((...((((((((.((....)).)))))((.....))))).))))).............. ( -32.40) >DroSim_CAF1 6210 108 - 1 CACUACUGCUGCCAAGCUGUUUUAAUGUCUUGGCGUGCAGUAGUAGUGGUUUUCCCGGAAGCACAGUAGUUUUUCCAAUAAUUUUGGGCACCACAGUCGACGCGUUUC .((((((((((((((((.........).))))))).)))))))).(((((...((((((((.((....)).)))))((.....))))).))))).............. ( -37.10) >DroYak_CAF1 4610 108 - 1 CACUACUGCCGCCAAGCUGUUUUAAUGUUUUGGCGUGCAGUAGUAGUGGUUUUCCCGGAAGCACAGUAGUUUUUCCAAUAAUUUUGGGUACCACAGUCGAUGUGUUUC .((((((((((((((((.........).))))))).)))))))).(((((...((((((((.((....)).)))))((.....))))).))))).............. ( -38.30) >DroPer_CAF1 6921 108 - 1 CACUACCGCCUCCUAGCUGCUUGAUUGCCUUGGCAUGCAGCAGCAGUGGCUUGCCCGACAGCACUGUCGUCUUGCCAAUGAUCUUGGGCACCAUCGUGGGUCCCCCUC .....((((......((((((.(..(((....)))..))))))).((((..(((((((((....))))(((........)))...))))))))).))))......... ( -37.10) >consensus CACUACUGCCGCCAAGCUGUUUUAAUGUCUUGGCGUGCAGUAGUAGUGGUUUUCCCGGAAGCACAGUAGUUUUUCCAAUAAUUUUGGGCACCACAGUCGACGCGUUUC .(((((((((((((((............))))))).)))))))).(((((...((((((((.((....)).)))))((.....))))).))))).............. (-21.42 = -23.70 + 2.28)

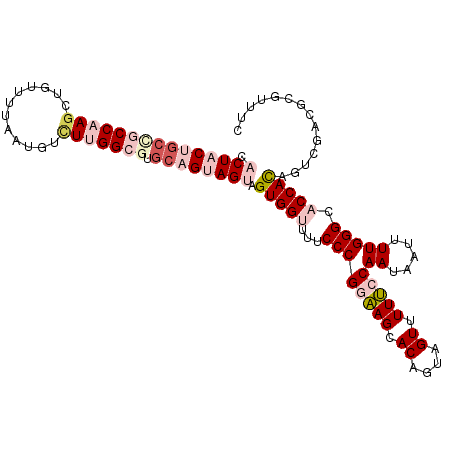

| Location | 4,815,293 – 4,815,407 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 71.40 |
| Mean single sequence MFE | -32.60 |
| Consensus MFE | -16.84 |
| Energy contribution | -16.81 |
| Covariance contribution | -0.02 |
| Combinations/Pair | 1.46 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.52 |
| SVM decision value | 1.43 |
| SVM RNA-class probability | 0.953130 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4815293 114 + 27905053 UAUUGGAAAAACUACUGUGCUUCCGGGAAAACCACUACUACUCCACGCCAAAACAUUAAAACAGAUUGGCGGUAGUAGUGUGCCCACAACCCAGACUACCACAACGACGACAAC ....((((..((....))..))))(((.....(((((((((....((((((..............)))))))))))))))..)))............................. ( -28.74) >DroVir_CAF1 7358 109 + 1 UAUUGGCAAGACAACUGUUGUGCCCGGCAAACCGCUGCUGCUGCAUGCCAAGACUCUUAAAU---UGGGCGGCAAUACAGGAGUCAGUACACAAACACCCACAGCCACAACG-- ...((((..(((..((((..((((.((((..(.((....)).)..))))..(.(((......---.))))))))..))))..))).((......)).......)))).....-- ( -31.00) >DroPse_CAF1 6985 114 + 1 CAUUGGCAAGACGACAGUGCUGUCGGGCAAGCCACUGCUGCUGCAUGCCAAGGGAAUCAAGCAGCUAGGAGGCGGUAGUGUUGUGACCACUCAACCCACAGCUACAACGACAAC ...((((...((((((.(((((((.((....)).((((((((.....((...)).....)))))).))..))))))).))))))((....))........)))).......... ( -34.90) >DroEre_CAF1 4655 114 + 1 UAUUGGAAAAACUACUGUGCUUCCUGGAAAACCACUUCUACUGCACGCCAAAACAUUAAAACAGCUUGGCGGCAGUAGUGUGGCCACCACCCAAACCACUACAACGACAACAAC ....((((..((....))..))))(((....((((..(((((((.((((((..............))))))))))))).))))...)))......................... ( -32.44) >DroAna_CAF1 8049 114 + 1 AAUUGGGAAAACCACAGUGCUCCCGGGAAAACCUCUGCUUUUGAAUGCAAAGACGAUAAAGCAACUGAGCGGCAGCAGUGUGGCCACCACCCAAGUAACCACCGCCACGACCAC ..(((((....(((((.((((.((((.....))...((((.....(((............)))...)))))).)))).)))))......))))).................... ( -30.40) >DroPer_CAF1 6949 114 + 1 CAUUGGCAAGACGACAGUGCUGUCGGGCAAGCCACUGCUGCUGCAUGCCAAGGCAAUCAAGCAGCUAGGAGGCGGUAGUGUUGUGACCACUCAACCCACAGCUACAACGACAAC ...((((...((((((.(((((((.((....)).((((((((...(((....)))....)))))).))..))))))).))))))((....))........)))).......... ( -38.10) >consensus UAUUGGCAAAACGACAGUGCUGCCGGGAAAACCACUGCUGCUGCAUGCCAAGACAAUAAAACAGCUAGGCGGCAGUAGUGUGGCCACCACCCAAACCACCACAACAACGACAAC ....((((..((....))..)))).((....))...((((((((.((((..................))))))))))))((((...............))))............ (-16.84 = -16.81 + -0.02)
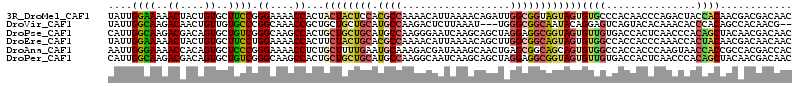



| Location | 4,815,333 – 4,815,446 |
|---|---|
| Length | 113 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 80.37 |
| Mean single sequence MFE | -40.85 |
| Consensus MFE | -29.91 |
| Energy contribution | -30.33 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.36 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.938570 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4815333 113 - 27905053 GCUCCACUGCCGCUUCCCACUGCCGCU-GAUGCAACUGCUGUUGUCGUCGUUGUGGUAGUCUGGGUUGUGGGCACACUACUACCGCCAAUCUGUUUUAAUGUUUUGGCGUGGAG .((((((((((((..((((((((((((-((((((((....)))).)))))..))))))))..)))..)).))))..........(((((..(((....)))..))))))))))) ( -46.50) >DroSec_CAF1 4845 104 - 1 GUUCC---------UCCCACUGCCGCU-GAUGCAACUGCUGUUGUCGUCGUUGUGGUGGUCUGGGUAGUGGCCACACUACUGCUGCCAAGCUGUUUUAAUGUCUUGGCGUUCAG .....---------..((((..(.((.-((((((((....)))).)))))).)..)))).((((((((((....)))))))...(((((((.........).))))))...))) ( -38.50) >DroSim_CAF1 6278 104 - 1 GUUCC---------UCCCACUGCCGCU-GAUGCAACUGCUGUUGUCGUCGUUGUGGUGGUCUGGGUAGUGGCCACACUACUGCUGCCAAGCUGUUUUAAUGUCUUGGCGUGCAG ...((---------..((((..(.((.-((((((((....)))).)))))).)..))))...))((((((....))))))(((((((((((.........).))))))).))). ( -42.00) >DroEre_CAF1 4695 104 - 1 GCUCC---------UCCCACUGCCGCU-GAUGCAACUGCUGUUGUUGUCGUUGUAGUGGUUUGGGUGGUGGCCACACUACUGCCGCCAAGCUGUUUUAAUGUUUUGGCGUGCAG (((((---------.((((..((((((-((((((((.......)))).))))..)))))).)))).)).))).......((((((((((((.........).))))))).)))) ( -41.10) >DroYak_CAF1 4678 104 - 1 GCUCC---------UCCCACUGCCGCU-GAUGCAACUGCUGUCGUCGUCGUUGUAGUGGUCUGGGUAGUGGCCACACUACUGCCGCCAAGCUGUUUUAAUGUUUUGGCGUGCAG ...((---------..(((((((.((.-(((((.((....)).).)))))).)))))))...))((((((....))))))(((((((((((.........).))))))).))). ( -38.90) >DroAna_CAF1 8089 91 - 1 -----------------------CGCUAGACGCCGCUGUUGUGGUCGUGGCGGUGGUUACUUGGGUGGUGGCCACACUGCUGCCGCUCAGUUGCUUUAUCGUCUUUGCAUUCAA -----------------------.((.((((((((((((..(....)..))))))))...((((((((..((......))..))))))))..........))))..))...... ( -38.10) >consensus GCUCC_________UCCCACUGCCGCU_GAUGCAACUGCUGUUGUCGUCGUUGUGGUGGUCUGGGUAGUGGCCACACUACUGCCGCCAAGCUGUUUUAAUGUCUUGGCGUGCAG ..................((((((((..((((((((....)))).))))...))))))))(((((((((((.....))))))))((((((.(((....))).))))))...))) (-29.91 = -30.33 + 0.42)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:16:46 2006