| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,624,208 – 4,624,312 |
| Length | 104 |
| Max. P | 0.555624 |

| Location | 4,624,208 – 4,624,312 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 79.32 |
| Mean single sequence MFE | -25.60 |
| Consensus MFE | -18.24 |
| Energy contribution | -20.14 |
| Covariance contribution | 1.89 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.555624 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
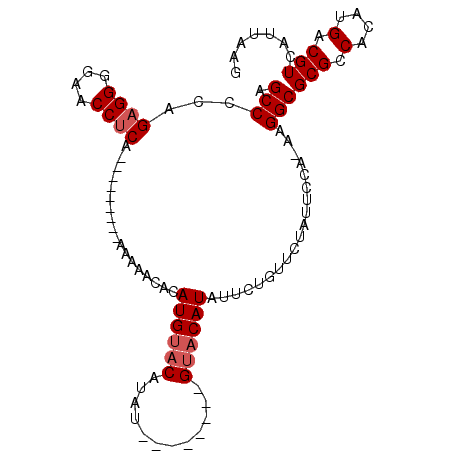
>3R_DroMel_CAF1 4624208 104 + 27905053 CUUAAUGACGUCAUGUGGCGCGCCUUUUGGAAUAGAACAGAAUAUGUACGUACAUUCAUAUGUACAUGUGUUUUUUUUGUUUUGUGAGGUUCCCCACUGGGGCG ........((((..((((.(.(((((.(..(((((((.((((((((((((((......))))))))))).))).)))))))..).))))).).))))...)))) ( -35.00) >DroEre_CAF1 24189 86 + 1 CUUAAUGACGUCAUGAGGCGCGCCUU-UGGAAUAGAACAGAAUAUGUAC--------AUAUGUACAUGUGUUUUU---------UGAGGUUCCCCUCUGGGGCU .........(((..((((.(.(((((-......((((((...(((((((--------....))))))))))))).---------.))))).).))))...))). ( -25.10) >DroYak_CAF1 24053 79 + 1 CUUAAUGACGUCAUGUGGCGCGCCUUGUGGAAUAGAACAGAAUAUG----------------UACAUGUGUUUUU---------UGAGGUUUCCCUCUGGGGCU ......(.((((....)))))(((((............(((((((.----------------.....))))))).---------.((((....)))).))))). ( -16.70) >consensus CUUAAUGACGUCAUGUGGCGCGCCUU_UGGAAUAGAACAGAAUAUGUAC________AUAUGUACAUGUGUUUUU_________UGAGGUUCCCCUCUGGGGCU .........(((..((((.(.(((((.......((((((...((((((((((......))))))))))))))))...........))))).).))))...))). (-18.24 = -20.14 + 1.89)



| Location | 4,624,208 – 4,624,312 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 79.32 |
| Mean single sequence MFE | -18.53 |
| Consensus MFE | -13.22 |
| Energy contribution | -14.30 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.538531 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4624208 104 - 27905053 CGCCCCAGUGGGGAACCUCACAAAACAAAAAAAACACAUGUACAUAUGAAUGUACGUACAUAUUCUGUUCUAUUCCAAAAGGCGCGCCACAUGACGUCAUUAAG (((.((.(((((....)))))................((((((((....)))))))).......................)).))).................. ( -20.50) >DroEre_CAF1 24189 86 - 1 AGCCCCAGAGGGGAACCUCA---------AAAAACACAUGUACAUAU--------GUACAUAUUCUGUUCUAUUCCA-AAGGCGCGCCUCAUGACGUCAUUAAG .(((...((((....)))).---------........((((((....--------))))))................-..)))(((.(....).)))....... ( -19.40) >DroYak_CAF1 24053 79 - 1 AGCCCCAGAGGGAAACCUCA---------AAAAACACAUGUA----------------CAUAUUCUGUUCUAUUCCACAAGGCGCGCCACAUGACGUCAUUAAG .......((((....)))).---------.......(((((.----------------.......(((........))).((....)))))))........... ( -15.70) >consensus AGCCCCAGAGGGGAACCUCA_________AAAAACACAUGUACAUAU________GUACAUAUUCUGUUCUAUUCCA_AAGGCGCGCCACAUGACGUCAUUAAG .(((...((((....))))..................((((((............))))))...................)))(((.(....).)))....... (-13.22 = -14.30 + 1.08)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:14:44 2006