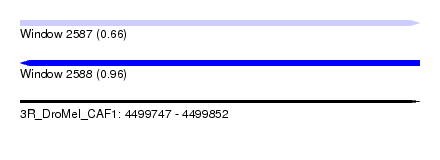

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,499,747 – 4,499,852 |
| Length | 105 |
| Max. P | 0.963295 |

| Location | 4,499,747 – 4,499,852 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 84.68 |
| Mean single sequence MFE | -24.78 |
| Consensus MFE | -17.66 |
| Energy contribution | -18.85 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.657214 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4499747 105 + 27905053 AAA-AAAAAUCGGAGCUUAAAACUAACAAGUUGGCAGUACCCUUCCAAUUAUAAAGGGUUAUCGAUAACUCUGCUGUAGCUAUCGGUGCUGCCAGCUGAUUAUAGU ...-.................((((...(((((((((((((.......(((((.(((((((....)))))))..))))).....))))))))))))).....)))) ( -31.60) >DroSec_CAF1 9378 105 + 1 AAU-AAAUAAAUGAGCUUAAAACUGACAAGUUGGCAGUUCCCUUCCAAUUAGAAAAGGCUAUCGAUAACUCUGCUAUUGCUAUCGGUGCUGCCAGCUGAUUAUAGU ...-.....(((.((((...((((....))))((((((..((((..........))))..(((((((.....((....))))))))))))))))))).)))..... ( -24.10) >DroSim_CAF1 9499 105 + 1 AAU-AAAUAUCAGAGCUUAAAACUGAUAAGUUGGCAGUUCCCUUCCAAUUAGAAAAGGUUAUCGAUAACUCUGCUAUUGCUAUCGGUGCAGCCAGCUGAUUAUAGU ...-...((((((.........))))))(((((((.....((((..........)))).((((((((.....((....))))))))))..)))))))......... ( -23.60) >DroEre_CAF1 9607 102 + 1 ---UAUAGCACUGAGCUUAAAACUGACAAGCUGGCAGUUACCUUCCAAUUAGAAAA-ACUAUCGAUAACUAUGCUAUCGCCAUCGCUGGUGCCAGCUGAUAAUCGU ---...(((.....)))...........(((((((((((...(((......))).)-)))...((((.......))))((((....)))))))))))......... ( -19.80) >consensus AAU_AAAUAACUGAGCUUAAAACUGACAAGUUGGCAGUUCCCUUCCAAUUAGAAAAGGCUAUCGAUAACUCUGCUAUUGCUAUCGGUGCUGCCAGCUGAUUAUAGU .....................((((...((((((((((..((((..........))))..(((((((.....((....))))))))))))))))))).....)))) (-17.66 = -18.85 + 1.19)


| Location | 4,499,747 – 4,499,852 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 84.68 |
| Mean single sequence MFE | -28.43 |
| Consensus MFE | -21.79 |
| Energy contribution | -22.47 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963295 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4499747 105 - 27905053 ACUAUAAUCAGCUGGCAGCACCGAUAGCUACAGCAGAGUUAUCGAUAACCCUUUAUAAUUGGAAGGGUACUGCCAACUUGUUAGUUUUAAGCUCCGAUUUUU-UUU .....(((((((((((((...((((((((.......))))))))...(((((((.......))))))).)))))((((....))))...))))..))))...-... ( -30.40) >DroSec_CAF1 9378 105 - 1 ACUAUAAUCAGCUGGCAGCACCGAUAGCAAUAGCAGAGUUAUCGAUAGCCUUUUCUAAUUGGAAGGGAACUGCCAACUUGUCAGUUUUAAGCUCAUUUAUUU-AUU .....(((.(((((((((...(((((((.........)))))))....(((((((.....)))))))..)))))((((....))))...)))).))).....-... ( -26.30) >DroSim_CAF1 9499 105 - 1 ACUAUAAUCAGCUGGCUGCACCGAUAGCAAUAGCAGAGUUAUCGAUAACCUUUUCUAAUUGGAAGGGAACUGCCAACUUAUCAGUUUUAAGCUCUGAUAUUU-AUU .........((.((((.....(((((((.........)))))))....(((((((.....)))))))....)))).))((((((.........))))))...-... ( -25.20) >DroEre_CAF1 9607 102 - 1 ACGAUUAUCAGCUGGCACCAGCGAUGGCGAUAGCAUAGUUAUCGAUAGU-UUUUCUAAUUGGAAGGUAACUGCCAGCUUGUCAGUUUUAAGCUCAGUGCUAUA--- .((.(((((.((((....)))))))))))((((((((((((((..((((-(.....)))))...)))))))...((((((.......))))))..))))))).--- ( -31.80) >consensus ACUAUAAUCAGCUGGCAGCACCGAUAGCAAUAGCAGAGUUAUCGAUAACCUUUUCUAAUUGGAAGGGAACUGCCAACUUGUCAGUUUUAAGCUCAGAUAUUU_AUU .........(((((((((...(((((((.........)))))))....(((((((.....)))))))..)))))((((....))))...))))............. (-21.79 = -22.47 + 0.69)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:43 2006