
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,497,242 – 4,497,365 |
| Length | 123 |
| Max. P | 0.824545 |

| Location | 4,497,242 – 4,497,333 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 75.99 |
| Mean single sequence MFE | -15.88 |
| Consensus MFE | -9.29 |
| Energy contribution | -9.85 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.824545 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
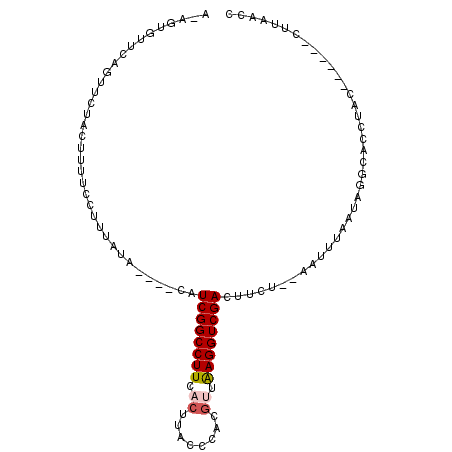
>3R_DroMel_CAF1 4497242 91 + 27905053 UUUAUUAAACAACGUUUUUAACAAGUAUGCACAGUGUUCAGUUCCACUUUUCCUUUGUU----GAUCGGCCUUCACUUACCCACCGUGAGGUCGA .................((((((((.......((((........)))).....))))))----))(((((((.(((.........)))))))))) ( -17.90) >DroSim_CAF1 6987 91 + 1 UUUAUUAAACAAAGUUUUCAACAAUUUUGCAAAGUGUUCAGUUCUACUUUUCCGUUGUU----GAUCGGCCUUCACUUACCCACGAUAAGGUCGA .................((((((((...(.((((((........)))))).).))))))----))((((((((..(........)..)))))))) ( -17.90) >DroEre_CAF1 7168 95 + 1 UUUAUAAAGCAAAGUUUUCCACAAUUUUGUAUAGUUUUCAGUUCUACUUUUCCUUUAUACAUACAUCGGCCUUCACUUACCCACGUUAAGGUCGA ..((((((((((((((......)))))))..(((.........)))......)))))))......((((((((.((........)).)))))))) ( -16.20) >DroYak_CAF1 7063 71 + 1 UUUAUUAAACAAAGUU--------------------UUUAGUUAUACUUUUCCUUUAUA----CAUCGGCCUUCACUUACCUACGUUAAGGUCGA ..........(((((.--------------------.........))))).........----..((((((((.((........)).)))))))) ( -11.50) >consensus UUUAUUAAACAAAGUUUUCAACAAUUUUGCA_AGUGUUCAGUUCUACUUUUCCUUUAUA____CAUCGGCCUUCACUUACCCACGUUAAGGUCGA .................................................................((((((((.((........)).)))))))) ( -9.29 = -9.85 + 0.56)



| Location | 4,497,272 – 4,497,365 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 79.04 |
| Mean single sequence MFE | -16.02 |
| Consensus MFE | -9.79 |
| Energy contribution | -10.35 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.685989 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4497272 93 + 27905053 ACAGUGUUCAGUUCCACUUUUCCUUUGUU----GAUCGGCCUUCACUUACCCACCGUGAGGUCGACUUCUAUAAUUCAUUAGGCACUUAC------CUUAACC ..(((((..(((((.((.........)).----))(((((((.(((.........))))))))))............)))..)))))...------....... ( -17.90) >DroSim_CAF1 7017 97 + 1 AAAGUGUUCAGUUCUACUUUUCCGUUGUU----GAUCGGCCUUCACUUACCCACGAUAAGGUCGACUUCU--AAUUUAUUAGGCACAUACCUCAACCUUAACC ((((((........))))))......(((----((((((((((..(........)..))))))))..(((--((....)))))........)))))....... ( -14.90) >DroEre_CAF1 7198 95 + 1 AUAGUUUUCAGUUCUACUUUUCCUUUAUACAUACAUCGGCCUUCACUUACCCACGUUAAGGUCGACUUUU--AGUUUAAUAGGCACCUGC------CUUAACC ...................................((((((((.((........)).)))))))).....--.(((....((((....))------)).))). ( -17.50) >DroYak_CAF1 7079 85 + 1 ------UUUAGUUAUACUUUUCCUUUAUA----CAUCGGCCUUCACUUACCUACGUUAAGGUCGACUUCU--AGUUUCAUAGGCACCUAC------CUUAACC ------....((((...............----..((((((((.((........)).)))))))).....--.......(((....))).------..)))). ( -13.80) >consensus A_AGUGUUCAGUUCUACUUUUCCUUUAUA____CAUCGGCCUUCACUUACCCACGUUAAGGUCGACUUCU__AAUUUAAUAGGCACCUAC______CUUAACC ...................................((((((((.((........)).))))))))...................................... ( -9.79 = -10.35 + 0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:41 2006