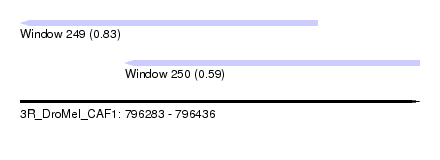
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 796,283 – 796,436 |
| Length | 153 |
| Max. P | 0.831210 |

| Location | 796,283 – 796,397 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 84.68 |
| Mean single sequence MFE | -18.27 |
| Consensus MFE | -12.54 |
| Energy contribution | -12.43 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.51 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.831210 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
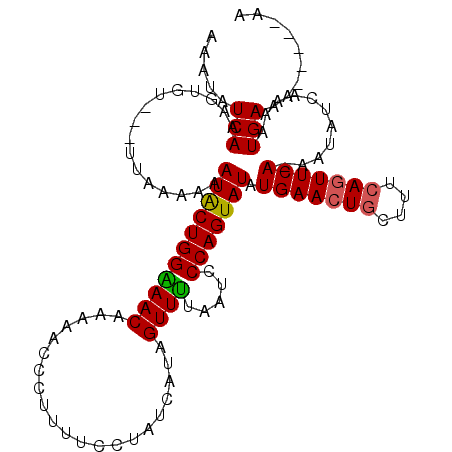
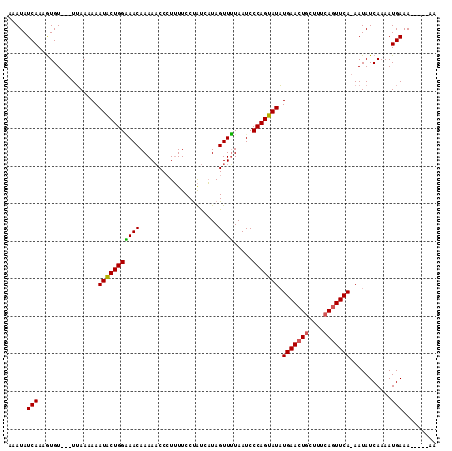
>3R_DroMel_CAF1 796283 114 - 27905053 AAAUAUCAAAAUGUUAUUGAAAAAAUGCUGGUAACAAAACCCGUAUUCCCGCCUUAGUUGUAAUCCCAGUAUUUGAACUGCUUGCAAUUCAAAAUAUCAAAAUGAAAGGGAAAA .....((((.......)))).....(((.(((......))).)))(((((.....((((((((...((((......)))).)))))))).......((.....))..))))).. ( -18.80) >DroSec_CAF1 20209 105 - 1 AAAUAUCAAAGUGU---UUAAAAAAUACUGGGAACAAAAACCCUUUUCCUAUCAUAGUUUUAAUCCCAGUAUAUGAACUUCUUUCAGUUCA-AAUAUCAAAAUGAAA-----AA .....(((......---.......(((((((((...(((((...............)))))..))))))))).((((((......))))))-..........)))..-----.. ( -19.66) >DroSim_CAF1 21084 105 - 1 AAAUAUCAAAGUGU---UUAAAAAAUACUGGAAACAAAAACCCUUUUCCUAUCAUAGUUUUAAUCCCAGUAUAUGAACUGCUUUCAGUUCA-AAUAUCAAAAUGAAA-----AA .....(((......---.......(((((((.....(((((...............)))))....))))))).(((((((....)))))))-..........)))..-----.. ( -16.36) >consensus AAAUAUCAAAGUGU___UUAAAAAAUACUGGAAACAAAAACCCUUUUCCUAUCAUAGUUUUAAUCCCAGUAUAUGAACUGCUUUCAGUUCA_AAUAUCAAAAUGAAA_____AA .....(((................(((((((((((.....................)))).....))))))).(((((((....)))))))...........)))......... (-12.54 = -12.43 + -0.11)

| Location | 796,323 – 796,436 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 71.84 |
| Mean single sequence MFE | -15.99 |
| Consensus MFE | -8.26 |
| Energy contribution | -8.07 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.591869 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
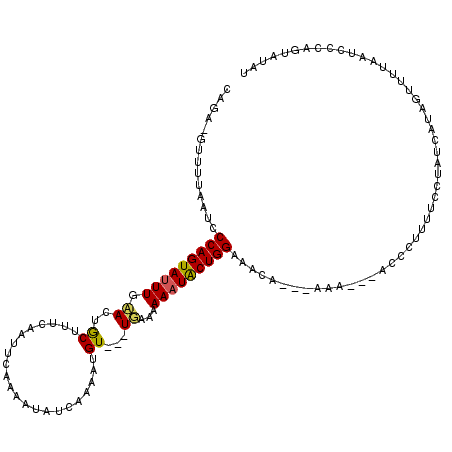
>3R_DroMel_CAF1 796323 113 - 27905053 CCGA-GAUGAAAUCCCAGUAUUUUAACUGCUUUCAAAUCAAAAUAUCAAAAUGUUAUUGAAAAAAUGCUGGUAACA---AAA---CCCGUAUUCCCGCCUUAGUUGUAAUCCCAGUAUUU ..((-..(((((...((((......)))).)))))..))......((((.......))))..(((((((((..(((---(..---...((......)).....))))....))))))))) ( -18.20) >DroSec_CAF1 20243 109 - 1 CAUA-GUUUUAAUCCCAGUAUUUGAACUACUUUCA-UCCCAAAUAUCAAAGUGU---UUAAAAAAUACUGGGAACA---AAA---ACCCUUUUCCUAUCAUAGUUUUAAUCCCAGUAUAU ....-(((((..(((((((((((((((.(((((.(-(.......)).)))))))---)))...))))))))))...---)))---))................................. ( -20.00) >DroSim_CAF1 21118 110 - 1 CAAA-GUUUUAGUCCCAGUAUUUGAACUGCUUUCAAUUCCAAAUAUCAAAGUGU---UUAAAAAAUACUGGAAACA---AAA---ACCCUUUUCCUAUCAUAGUUUUAAUCCCAGUAUAU ....-(((((.((.(((((((((((((.(((((..............)))))))---)))...))))))))..)).---)))---))................................. ( -15.34) >DroEre_CAF1 19282 99 - 1 CAGAAAUUUUAAUCCCAGUACUUGCACUGCUUUCAAUUCAAAAUAUCAAAAUGUUAUUGAAAAAAUGCUGGAAUCAGAAAAAAAAAACAUUUACCUAUC--------------------- ......((((.((.((((((...((...))(((((((....(((((....))))))))))))...)))))).)).))))....................--------------------- ( -10.40) >consensus CAGA_GUUUUAAUCCCAGUAUUUGAACUGCUUUCAAUUCAAAAUAUCAAAAUGU___UGAAAAAAUACUGGAAACA___AAA___ACCCUUUUCCUAUCAUAGUUUUAAUCCCAGUAUAU ..............(((((((((.((..((......................))...))...)))))))))................................................. ( -8.26 = -8.07 + -0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:35:04 2006