
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,320,045 – 4,320,361 |
| Length | 316 |
| Max. P | 0.964578 |

| Location | 4,320,045 – 4,320,163 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.21 |
| Mean single sequence MFE | -21.93 |
| Consensus MFE | -21.35 |
| Energy contribution | -21.47 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.851958 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4320045 118 + 27905053 UAUAAAAUAUUCAAUGGUCGGUAUAUUUUUUGCUCGGCUUGAUUUUAAUUCCAAAUUCAUUUCGUCUAUAAAUGCAAUUUUCUAAAUUGCAUUGAUUUGUGCGCCCAAAACACACAA--C ..((((((.......((((((((.......))).)))))..)))))).......................(((((((((.....)))))))))...(((((.(.......).)))))--. ( -18.70) >DroSec_CAF1 7412 118 + 1 UAUAAAAUAUUCAAUGGUCGGUAUAUUUUUUGCUCGGCUUGAUUUUAAUUCCAAAUUCAUUUCGUCUAUAAAUGCAAUUUUCUAAAUUGCAUUGAUUUGUGUGCCCAAAACACACAA--C ..((((((.......((((((((.......))).)))))..)))))).......................(((((((((.....)))))))))...(((((((.......)))))))--. ( -22.60) >DroEre_CAF1 7418 120 + 1 UAUAAAAUAUUCAAUGGUCGGUAUAUUUUUUGCUCGGCCUGAUUUUAAUUCCAAAUUCAUUUCGUCUACAAAUGCAAUUUUCUAAAUUGCAUUGAUUUGUGUGCCCAAAACACACAACAC ..((((((.......((((((((.......))).)))))..)))))).......................(((((((((.....)))))))))...(((((((.......)))))))... ( -24.50) >consensus UAUAAAAUAUUCAAUGGUCGGUAUAUUUUUUGCUCGGCUUGAUUUUAAUUCCAAAUUCAUUUCGUCUAUAAAUGCAAUUUUCUAAAUUGCAUUGAUUUGUGUGCCCAAAACACACAA__C ..((((((.......((((((((.......))).)))))..)))))).......................(((((((((.....)))))))))...(((((((.......)))))))... (-21.35 = -21.47 + 0.11)



| Location | 4,320,085 – 4,320,203 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.21 |
| Mean single sequence MFE | -23.80 |
| Consensus MFE | -23.00 |
| Energy contribution | -23.33 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.57 |
| SVM RNA-class probability | 0.964578 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4320085 118 + 27905053 GAUUUUAAUUCCAAAUUCAUUUCGUCUAUAAAUGCAAUUUUCUAAAUUGCAUUGAUUUGUGCGCCCAAAACACACAA--CAGCCAGACGGCAAAGCUGCAAAACCAAUGGCAAAUACCGA ..........(((.........(((((...(((((((((.....)))))))))...(((((.(.......).)))))--.....)))))(((....)))........))).......... ( -21.20) >DroSec_CAF1 7452 118 + 1 GAUUUUAAUUCCAAAUUCAUUUCGUCUAUAAAUGCAAUUUUCUAAAUUGCAUUGAUUUGUGUGCCCAAAACACACAA--CAGCCAGACGGCAAAGCUGCAAAACCAAUGGCAAAUACCAA ..........(((.........(((((...(((((((((.....)))))))))...(((((((.......)))))))--.....)))))(((....)))........))).......... ( -25.10) >DroEre_CAF1 7458 120 + 1 GAUUUUAAUUCCAAAUUCAUUUCGUCUACAAAUGCAAUUUUCUAAAUUGCAUUGAUUUGUGUGCCCAAAACACACAACACAGCCAGACGGCAAAGCUGCAAAACCAAUGGCAAAUACCGA ..........(((.........(((((...(((((((((.....)))))))))...(((((((.......))))))).......)))))(((....)))........))).......... ( -25.10) >consensus GAUUUUAAUUCCAAAUUCAUUUCGUCUAUAAAUGCAAUUUUCUAAAUUGCAUUGAUUUGUGUGCCCAAAACACACAA__CAGCCAGACGGCAAAGCUGCAAAACCAAUGGCAAAUACCGA ..........(((.........(((((...(((((((((.....)))))))))...(((((((.......))))))).......)))))(((....)))........))).......... (-23.00 = -23.33 + 0.33)



| Location | 4,320,085 – 4,320,203 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.21 |
| Mean single sequence MFE | -30.37 |
| Consensus MFE | -30.22 |
| Energy contribution | -30.00 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.60 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.732774 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4320085 118 - 27905053 UCGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCUG--UUGUGUGUUUUGGGCGCACAAAUCAAUGCAAUUUAGAAAAUUGCAUUUAUAGACGAAAUGAAUUUGGAAUUAAAAUC ..(((....))).((((((((((......))((((((....--((((((((.....))))))))...(((((((((.....)))))))))..))))))..........)))))))).... ( -30.00) >DroSec_CAF1 7452 118 - 1 UUGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCUG--UUGUGUGUUUUGGGCACACAAAUCAAUGCAAUUUAGAAAAUUGCAUUUAUAGACGAAAUGAAUUUGGAAUUAAAAUC .((((....))))((((((((((......))((((((....--((((((((.....))))))))...(((((((((.....)))))))))..))))))..........)))))))).... ( -30.80) >DroEre_CAF1 7458 120 - 1 UCGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCUGUGUUGUGUGUUUUGGGCACACAAAUCAAUGCAAUUUAGAAAAUUGCAUUUGUAGACGAAAUGAAUUUGGAAUUAAAAUC ..(((....))).((((((((((......))((((((......((((((((.....))))))))...(((((((((.....)))))))))..))))))..........)))))))).... ( -30.30) >consensus UCGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCUG__UUGUGUGUUUUGGGCACACAAAUCAAUGCAAUUUAGAAAAUUGCAUUUAUAGACGAAAUGAAUUUGGAAUUAAAAUC ..(((....))).((((((((((......))((((((......((((((((.....))))))))...(((((((((.....)))))))))..))))))..........)))))))).... (-30.22 = -30.00 + -0.22)


| Location | 4,320,125 – 4,320,243 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.21 |
| Mean single sequence MFE | -33.27 |
| Consensus MFE | -31.51 |
| Energy contribution | -31.07 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.731152 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
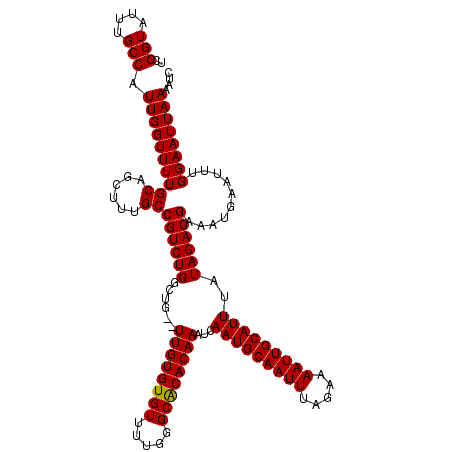
>3R_DroMel_CAF1 4320125 118 - 27905053 GCGCAAUUUUCGAAUUAUUGAUGGCACACAGAAAUCAAUUUCGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCUG--UUGUGUGUUUUGGGCGCACAAAUCAAUGCAAUUUAGA ..((((...(((((..((((((..(.....)..)))))))))))...))))(((((((((.((((((.........)))))--).((((((.....)))))))))))))))......... ( -33.40) >DroSec_CAF1 7492 118 - 1 GCGCAAUUUUCGAAUUAUUGAUGGCACACAGAAAUCAAUUUUGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCUG--UUGUGUGUUUUGGGCACACAAAUCAAUGCAAUUUAGA ..(((..............(((((((....((((((((...((((....)))))))))))).......)))))))(((...--((((((((.....)))))))).))).)))........ ( -32.00) >DroEre_CAF1 7498 120 - 1 ACGCAAUUUUCGAAUUAUUGAUGGCACACAGAAAUCAAUUUCGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCUGUGUUGUGUGUUUUGGGCACACAAAUCAAUGCAAUUUAGA ..((((...(((((..((((((..(.....)..)))))))))))...))))(((((((((.((((((.........))))))...((((((.....)))))))))))))))......... ( -34.40) >consensus GCGCAAUUUUCGAAUUAUUGAUGGCACACAGAAAUCAAUUUCGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCUG__UUGUGUGUUUUGGGCACACAAAUCAAUGCAAUUUAGA ..((((...(((((..((((((..(.....)..)))))))))))...))))(((((((((..(((((.........)))))....((((((.....)))))))))))))))......... (-31.51 = -31.07 + -0.44)


| Location | 4,320,163 – 4,320,283 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -27.07 |
| Consensus MFE | -26.35 |
| Energy contribution | -26.80 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.709014 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
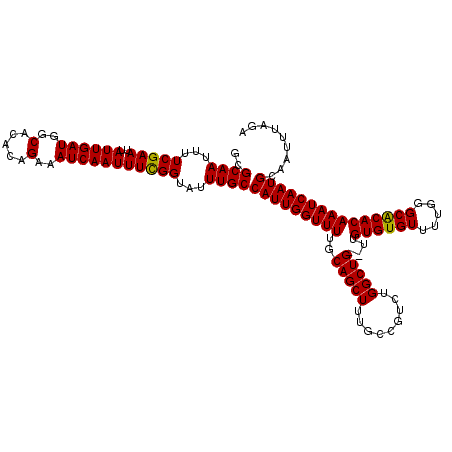
>3R_DroMel_CAF1 4320163 120 - 27905053 GGAAUUCCAAAUUAAAUUCCCUUAAAUUAGCAUUAAAAAUGCGCAAUUUUCGAAUUAUUGAUGGCACACAGAAAUCAAUUUCGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCU ((((((........)))))).........(((..(..((((.((((...(((((..((((((..(.....)..)))))))))))...))))))))..)..)))......(((....))). ( -28.70) >DroSec_CAF1 7530 120 - 1 GGAAUUUCAAAUUAAAUUCCCUUAAAUUAGCAUUAAAAAUGCGCAAUUUUCGAAUUAUUGAUGGCACACAGAAAUCAAUUUUGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCU (((((((......))))))).........(((..(..((((.((((...(((((..((((((..(.....)..)))))))))))...))))))))..)..)))......(((....))). ( -27.50) >DroEre_CAF1 7538 120 - 1 GGAAUUUCAAAUUAAAUUCCCUUAAAUUAGCAUUAAAAAUACGCAAUUUUCGAAUUAUUGAUGGCACACAGAAAUCAAUUUCGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCU (((((((......))))))).........(((..(..(((..((((...(((((..((((((..(.....)..)))))))))))...)))).)))..)..)))......(((....))). ( -25.00) >consensus GGAAUUUCAAAUUAAAUUCCCUUAAAUUAGCAUUAAAAAUGCGCAAUUUUCGAAUUAUUGAUGGCACACAGAAAUCAAUUUCGGUAUUUGCCAUUGGUUUUGCAGCUUUGCCGUCUGGCU (((((((......))))))).........(((..(..((((.((((...(((((..((((((..(.....)..)))))))))))...))))))))..)..)))......(((....))). (-26.35 = -26.80 + 0.45)


| Location | 4,320,243 – 4,320,361 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.02 |
| Mean single sequence MFE | -28.17 |
| Consensus MFE | -23.89 |
| Energy contribution | -24.33 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.85 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
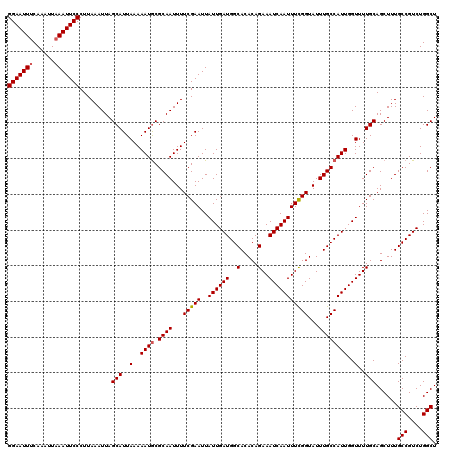
>3R_DroMel_CAF1 4320243 118 - 27905053 UUUAUGCCUAAUCUCCGAUUCCC--CGGGGCUGUUUGCACUGCCCAUUUUAGCGGCAGAAAAUUUAAUAGGCCCGCCGAUGGAAUUCCAAAUUAAAUUCCCUUAAAUUAGCAUUAAAAAU ...((((.((((.......((..--((((.(((((....((((((......).))))).......))))).))))..)).((((((........)))))).....))))))))....... ( -27.60) >DroSec_CAF1 7610 120 - 1 UUUAUGCCUACUCUCCGAUCGCCCGCGGGGAUGUUUGCACUGCCCAUUUCAGCGGCAGAAAAUUUAAUAGGCCCGUCGAUGGAAUUUCAAAUUAAAUUCCCUUAAAUUAGCAUUAAAAAU .....(((((.((((((........))))))........((((((......).))))).........)))))..((..(((((((((......))))))).....))..))......... ( -28.00) >DroEre_CAF1 7618 117 - 1 UUUAUGCCUAAUCUCCGAUC-CC--CGGGGCUGUUUGCACUGCCCAUUUCAGCGGCACAGAAUUUAAUAGGCCCGCUGAUGGAAUUUCAAAUUAAAUUCCCUUAAAUUAGCAUUAAAAAU .....(((((..(((((...-..--)))))...((((..((((........))))..))))......)))))..(((((((((((((......))))))).....))))))......... ( -28.90) >consensus UUUAUGCCUAAUCUCCGAUC_CC__CGGGGCUGUUUGCACUGCCCAUUUCAGCGGCAGAAAAUUUAAUAGGCCCGCCGAUGGAAUUUCAAAUUAAAUUCCCUUAAAUUAGCAUUAAAAAU .....(((((..(((((........))))).........((((((......).))))).........)))))..((.((((((((((......))))))).....))).))......... (-23.89 = -24.33 + 0.45)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:12:25 2006